-
PDF
- Split View
-
Views
-
Cite
Cite
Luke A Hoekstra, Mohammad A Siddiq, Kristi L Montooth, Pleiotropic Effects of a Mitochondrial–Nuclear Incompatibility Depend upon the Accelerating Effect of Temperature in Drosophila, Genetics, Volume 195, Issue 3, 1 November 2013, Pages 1129–1139, https://doi.org/10.1534/genetics.113.154914
Close - Share Icon Share
Abstract
Interactions between mitochondrial and nuclear gene products that underlie eukaryotic energy metabolism can cause the fitness effects of mutations in one genome to be conditional on variation in the other genome. In ectotherms, the effects of these interactions are likely to depend upon the thermal environment, because increasing temperature accelerates molecular rates. We find that temperature strongly modifies the pleiotropic phenotypic effects of an incompatible interaction between a Drosophila melanogaster polymorphism in the nuclear-encoded, mitochondrial tyrosyl-transfer (t)RNA synthetase and a D. simulans polymorphism in the mitochondrially encoded tRNATyr. The incompatible mitochondrial–nuclear genotype extends development time, decreases larval survivorship, and reduces pupation height, indicative of decreased energetic performance. These deleterious effects are ameliorated when larvae develop at 16° and exacerbated at warmer temperatures, leading to complete sterility in both sexes at 28°. The incompatible genotype has a normal metabolic rate at 16° but a significantly elevated rate at 25°, consistent with the hypothesis that inefficient energy metabolism extends development in this genotype at warmer temperatures. Furthermore, the incompatibility decreases metabolic plasticity of larvae developed at 16°, indicating that cooler development temperatures do not completely mitigate the deleterious effects of this genetic interaction. Our results suggest that the epistatic fitness effects of metabolic mutations may generally be conditional on the thermal environment. The expression of epistatic interactions in some environments, but not others, weakens the efficacy of selection in removing deleterious epistatic variants from populations and may promote the accumulation of incompatibilities whose fitness effects will depend upon the environment in which hybrids occur.
INTERACTIONS between genes coordinate development and underlie physiological performance to determine fitness within environmental contexts. In eukaryotic cells, interactions between mitochondrial and nuclear gene products provide energy for metabolism, development, and life-history traits. Because of this, the effects of mutations on metabolism in one genome are expected to be conditional on mutations in the other genome (Blier et al. 2001; Rand et al. 2004; Burton and Barreto 2012) and these genetic interactions are expected to affect suites of physiological and behavioral traits that depend on energy acquisition and allocation (Clark et al. 1995a,b; Montooth et al. 2003; Schmidt et al. 2005; Nelson et al. 2007; Kent et al. 2009; Ballard and Melvin 2010). Furthermore, the known thermodynamic effects of temperature on biochemical processes and metabolic rate (Krogh 1916; Clarke and Fraser 2004) suggest that the phenotypic effects of metabolic mutations in ectotherms will depend upon the thermal environment. Thus, understanding the pleiotropic effects of genotype-by-genotype interactions (G × G), genotype-by-environment interactions (G × E), and more complex interactions such as environment-dependent epistasis (G × G × E) is necessary to understand the evolutionary dynamics of metabolism, life history, and disease, as well as genetic differentiation and incompatibility between locally adapted populations (Clark and Fucito 1998; Wade 2000; Moore 2003; Mackay 2004; Bergland et al. 2008; Phillips 2008; Chandler et al. 2013; Flynn et al. 2013).
Metabolic mutations and genetic interactions that affect energy allocation are likely to influence trade-offs among components of fitness (Zera and Harshman 2001; Zera and Zhao 2003; King and Roff 2010; Robinson and Beckerman 2013). Regulation of energy metabolism may also be important for physiological responses that maintain fitness across heterogeneous environments experienced within a lifetime. Induction of environmental responses can be energetically costly (Hoekstra and Montooth 2013), and low resting metabolic rates may be selectively favored to maximize the energy available for growth, reproduction, and physiological responses to the environment (Artacho and Nespolo 2009; Ketola and Kotiaho 2009). Thus, we expect the fate of mutations affecting energy metabolism to be governed by selection via their pleiotropic effects on traits associated with growth, reproduction, and the maintenance of homeostasis across heterogeneous environments.
The machinery of aerobic metabolism requires protein synthesis within the mitochondria that depends upon the aminoacylation of mitochondrial transfer (t)RNAs by nuclear-encoded tRNA synthetases and upon interactions between ribosomal proteins encoded by both genomes. The mitochondrial-encoded proteins then interact with nuclear-encoded components of oxidative phosphorylation (OXPHOS) to generate ATP. This metabolic function is thought to be maintained by the fixation of compensatory and coadapted mutations in nuclear genomes (Blier et al. 2001; Grossman et al. 2004; Rand et al. 2004; Meiklejohn et al. 2007; Dowling et al. 2008; Osada and Akashi 2012; Barreto and Burton 2013) in response to the accumulation of deleterious substitutions and the fixation of favorable mutations in the mitochondrial (mt)DNA (Lynch 1996; Bazin et al. 2006; Meiklejohn et al. 2007; Montooth and Rand 2008; Neiman and Taylor 2009). Coevolved substitutions in mitochondrial and nuclear genomes can generate incompatibilities that decrease fitness in hybrids between divergent populations and closely related species (recently reviewed by Burton and Barreto 2012). As predicted, these cytonuclear incompatibilities often disrupt energetic function (Kenyon and Moraes 1997; Mckenzie et al. 2003; Ellison and Burton 2006; Arnqvist et al. 2010; Chou et al. 2010; Meiklejohn et al. 2013), and their effects on development and metabolic rate can be thermally dependent (Dowling et al. 2007; Arnqvist et al. 2010).
Phenotypes that depend upon mitochondrial–nuclear interactions may be particularly thermally dependent, not only due to the temperature sensitivity of molecular interactions, but also by the potential of temperature to increase demands on ATP pools by accelerating energy usage and development in ectotherms (Krogh 1916; Hochachka and Somero 2002; Clarke and Fraser 2004; Ghosh et al. 2013). While many mutations in core metabolic processes will disrupt energy metabolism such that the phenotypic effects are manifest under all conditions, those mutations segregating in natural populations are more likely to cause inefficiencies in metabolism that are revealed only when temperature accelerates biological rates and increases energetic demands (Gibson and Dworkin 2004; Hermisson and Wagner 2004). The ubiquitous effects of temperature on biological rates in ectotherms and of energy metabolism on fitness traits warrant quantification and mechanistic dissection of how temperatures well within critical thermal limits modify the pleiotropic effects of mitochondrial–nuclear interactions on suites of organismal traits.
Here we report that development temperature strongly modifies an incompatibility between Drosophila simulans and D. melanogaster polymorphisms in the mt-tRNATyr and the nuclear-encoded tyrosyl-tRNA synthetase (mt-TyrRS) (Montooth et al. 2010; Meiklejohn et al. 2013), such that the deleterious effects of this mitochondrial–nuclear interaction are mitigated by cooler temperatures and magnified at warmer temperatures. Larvae of the incompatible genotype developed at warmer temperatures have less efficient metabolism and experience a trade-off such that the allocation of resources to growth comes at a cost to other performance traits. While these effects are largely alleviated at cooler temperatures, effects on metabolic plasticity persist. We discuss the implications of these results in the context of theory on the evolutionary dynamics of mitochondrial–nuclear interactions and epistasis (Whitlock et al. 1995; Phillips and Johnson 1998; Rand et al. 2001; Phillips 2008; Lachance et al. 2011), as well as interactions with the environment that reduce the efficacy of selection in purging deleterious and fixing advantageous alleles in populations (Whitlock 1996; Turelli and Barton 2004; Van Dyken and Wade 2010).
Materials and Methods
Drosophila genotypes and maintenance
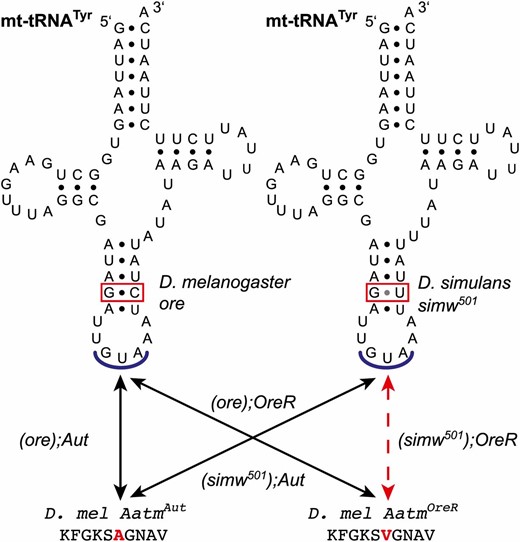
The four mitochondrial–nuclear genotypes used in this experiment are a subset of the (mtDNA);nuclear genotypes generated by Montooth et al. (2010) that exhibit a synergistic epistatic interaction between a SNP in the D. simulans mt-tRNATyr and a SNP in the D. melanogaster nuclear-encoded mt-TyrRS (Meiklejohn et al. 2013). A G:C to G:U mutation in the anticodon stem of the D. simulans simw501 tRNATyr is incompatible with an amino acid substitution (A275V) in the D. melanogaster OreR allele of Aatm, which is the nuclear-encoded tRNA synthetase that aminoacylates the mt-tRNATyr (Meiklejohn et al. 2013) (Figure 1). When developed at 25°, the genotype (simw501);OreR has severely extended development time, decreased OXPHOS activity, reduced female fecundity, and shortened mechanosensory bristles, while the other three mitochondrial–nuclear combinations—(ore);OreR, (simw501);Aut, and (ore);Aut—are predominantly wild type (Meiklejohn et al. 2013). These genotypes allowed us to quantify the main effects of mtDNA and nuclear genome, as well as the mitochondrial–nuclear interaction.
Four mitochondrial–nuclear genotypes combine two mtDNA alleles with two nuclear alleles underlying an epistatic interaction. A polymorphism in the anticodon stem of the mt-tRNATyr interacts epistatically with an amino acid polymorphism in the nuclear-encoded mt-TyrRS that aminoacylates this mitochondrial tRNA. The particular combination of the D. simulans simw501 mtDNA with the D. melanogaster OreR allele of Aatm is incompatible (dashed line). (simw501);OreR individuals have low OXPHOS activity, compromised development, and decreased fecundity, while the other three (mtDNA);Nuclear genotypes have predominantly similar and wild-type phenotypes (Meiklejohn et al. 2013).
Each of these genotypes was maintained at three temperatures—16°, 22°, and 25°—on Bloomington Drosophila Stock Center media with a 12-hr:12-hr light cycle. Genotypes were acclimated to their environment for at least three generations prior to phenotypic measurements to control for trans-generational effects. We were not able to maintain cultures of the incompatible genotype (simw501);OreR at 28° due to temperature-dependent sterility (see results below). However, we measured development time at 28°, using embryos from parents reared at 25°.
Development time and pupation height
We quantified the extent to which development temperature modifies the mitochondrial–nuclear interaction effect on development time by scoring the time from egg to adult eclosion for each genotype developed at 16°, 22°, 25°, and 28°. Replicate pools of 50 0- to 12-hr-old eggs were collected and placed in a vial to avoid known effects of larval density on development time and pupation height (e.g., Mueller and Sweet 1986). For each experiment, ∼20 vials containing 50 eggs for each genotype at each temperature were scored for pupation and eclosion twice a day at 12-hr intervals.
Pupation height was scored using a subset of the vials used to measure development time. Each vial was divided into four, 2-cm-length quadrants, except for quadrant 1, which contained the length of the fly media (∼3.5 cm in length). Each pupa was assigned a quadrant score based on the quadrant in which it pupated, with all pupae on the food assigned a value of 1. The average height of quadrant 2 was 4.5 cm (1 cm above media level), that of quadrant 3 was 6.5 cm (3 cm above media level), and that of quadrant 4 was 8.5 cm (5 cm above media level). We then calculated mean pupation height for 10 replicate vials of each genotype from each of the three developmental temperatures.
Temperature-dependent sterility
We attempted to rear each genotype at an additional temperature of 28°. However, we could not maintain cultures of the incompatible (simw501);OreR genotype at this temperature. To determine whether this was due to sterility of males or females, we set up reciprocal crosses between virgin (simw501);OreR and virgin (ore);OreR females and males, as (ore);OreR males and females were fertile when developed at 28°. We then scored whether vials produced offspring. We also shifted a subset of mating pairs back to 25° to determine whether the temperature-dependent effects on fertility were reversible in adults.
Larval metabolic rate
We measured larval CO2 production (microliters per hour), using flow-through respirometry as a measure of metabolic rate as in Hoekstra and Montooth (2013). We measured larval metabolic rate for all genotypes developed at 16° and 25° and measured at 16° and 25°, to generate the thermal reaction norm for larval metabolic rate for each genotype. Every respirometry experiment (i.e., a single run of the respirometer) measured pools of five larvae from each genotype aged either 10 days (at 16°) or 6 days (at 25°). At this age, the larvae at each developmental temperature were at the third-instar stage but had not begun to wander. We randomly picked five, prewandering larvae of each genotype and placed them in the cap of a 1.7-ml microcentrifuge tube that contained 0.5 ml of fly media and had been tared on the balance. We then massed the larvae to the nearest 0.1 mg. Genotypes from each development temperature were randomized across respirometry chambers. All respirometry experiments were conducted during the hours of 8:00 am to 8:00 pm to minimize known effects of circadian rhythm on metabolism. Larvae were acclimated in the respirometer for 1–2 hr before recordings were taken.
CO2 production was measured using a Sable Systems flow-through respirometry system (Sable Systems International, Las Vegas), using air that had been scrubbed of CO2 and then rehydrated. The air stream passed through the chamber containing larvae, was scrubbed for water, and then passed into a CO2 analyzer (Li-Cor 7000 CO2/H2O Analyzer; LI-COR, Lincoln, NE). In each respirometry experiment, CO2 was sampled from seven chambers, with an empty chamber that served as the baseline. CO2 values for each sample of larvae were drift corrected in ExpeData version 1.1.15 (Sable Systems International), using baseline CO2 data before and after each sample and employing the two-endpoint automatic method. These drift-corrected values were exported for the remaining analyses, which were done using the R statistical package version 2.15.1 (R Development Core Team 2011). Drift-corrected CO2 values were converted from parts per million to microliters per hour, using the mass flow rate of 80 ml⋅min−1, and then log-transformed to improve normality and homoscedasticity. For all subsequent analyses, our measure of metabolic rate [the volume of CO2 (VCO2) produced by five larvae in microliters per hour] was calculated as the mean of these drift-corrected values from 10-min recordings of CO2 production. We measured 20 replicate pools of five larvae from each genotype in each temperature treatment. This experimental design with high replication allowed us to test for complex interactions between mtDNA, nuclear genome, development temperature (TDEV), and measurement temperature (TMEASURE).
Statistical analyses
We used analysis of variance (ANOVA) to test for the fixed effects of mtDNA, nuclear genome, temperature, and their interactions on development time, survival, pupation height, and larval mass, including the replicate vial as a random effect in a mixed-model ANOVA where appropriate. All statistical analyses used the statistical package R version 2.15.1 (R Development Core Team 2011). Full results from ANOVA models are provided in Supporting Information, Table S1, Table S2, Table S3, Table S4, Table S5, and Table S6.
To analyze effects of mitochondrial–nuclear genotype and temperature on metabolic rate, we used type II model regression, to account for the relationship between mass and metabolic rate, as well as error in larval body mass. We used standard major axis regression to test for the heterogeneity of slopes for the bivariate relationship of log-transformed body mass and VCO2, using SMATR, version 3.2.6 (Warton et al. 2006). When justified, common slopes were fitted to all genotypes and we tested for shifts in elevation on the y-axis (i.e., genotype differences in metabolic rate) and for shifts along the common slope (i.e., genotype differences in body mass) (Table S5). Within development-by-measurement temperature treatments, common slopes could be fitted to all genotypes. This allowed us to calculate mass-corrected metabolic rates by taking the residuals of a regression of metabolic rate on body mass and adding the grand mean of all fitted values to provide meaningful scale. The genotype means of the mass-corrected metabolic rates were used to compare metabolic rates across measurement temperatures; the slope of this reaction norm is a measure of the thermal plasticity of metabolic rate. The Q10 was calculated from the genotype mean mass-corrected routine metabolic rates (RMRs) as (Table S7).
Results
We measured development time, survival, pupation height, and larval metabolic rates across a range of nonstressful development temperatures (16°–28°) in Drosophila mitochondrial–nuclear genotypes that exhibit synergistic epistasis. A single-nucleotide polymorphism in the anticodon stem of the D. simulans simw501 mt-tRNATyr interacts epistatically with an amino acid polymorphism in the D. melanogaster OreR allele of the nuclear-encoded gene Aatm that encodes the mt-TyrRS that aminoacylates the mt-tRNATyr (Figure 1) (Meiklejohn et al. 2013). On their own these mutations have little phenotypic effect, but together they significantly delay larval development, decrease OXPHOS activity, compromise sensory bristle formation during metamorphosis, and reduce adult fecundity (Meiklejohn et al. 2013). Here we report measures from the following (mtDNA);nuclear genotypes that combine two wild-type D. melanogaster nuclear genomes (OreR and Aut) with D. melanogaster ore and D. simulans simw501 mtDNAs—(ore);OreR, (ore);Aut, (simw501);Aut, and (simw501);OreR—of which only (simw501);OreR is an incompatible combination of mitochondrial and nuclear genotypes.
Temperature modifies mitochondrial–nuclear effects on development
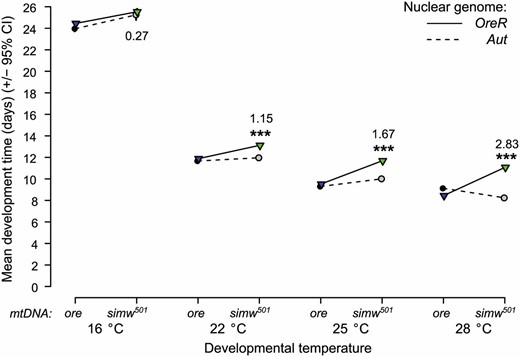
Cooler temperatures greatly extended embryo-to-adult development time of all genotypes (Figure 2), as expected for Drosophila and for ectotherms in general (Powsner 1935; Johnston 1990; Gillooly et al. 2002). Temperature strongly modified the effects of the mitochondrial–nuclear genetic interaction on development time, resulting in a significant epistasis–environment interaction (i.e., G × G × E) (mtDNA × nuclear × TDEV: F3, 309 = 64.0, P < 0.0001) (Figure 2). At 28°, the mitochondrial–nuclear incompatibility extended development time of (simw501);OreR individuals by almost 3 days, relative to the other genotypes (Figure 2). The magnitude and significance of this genetic interaction decreased as temperature decreased (Figure 2 and Table S1). When larvae developed at 16°, the mitochondrial–nuclear interaction did not significantly affect development time (mtDNA × nuclear: F1, 80 = 0.38, P = 0.5413). At 16°, both the mtDNA and the nuclear genome have small but significant main effects on development (Table S1). However, these effects are much smaller than the developmental delay caused by the mitochondrial–nuclear interaction at warmer temperatures, particularly when the delay is considered as a fraction of the overall development time. At 28°, the 2.83-day developmental delay of (simw501);OreR individuals relative to (simw501);Aut individuals represents 26% of their total egg-to-adult development time. This delay decreases to 14%, 9%, and 1% of their total development time at 25°, 22°, and 16°, respectively.
The developmental delay caused by the mitochondrial–nuclear interaction is magnified as temperature increases. The mean egg-to-adult development time for (simw501);OreR individuals is nearly 3 days longer during development at 28°, relative to that of other genotypes. This delay becomes smaller and less statistically significant when flies are reared under increasingly cooler conditions (Table S1). At 16°, there is no significant mitochondrial–nuclear interaction and the development time of (simw501);OreR individuals is nearly identical to that of (simw501);Aut individuals. Numbers represent the difference in mean development time between (simw501);OreR and (simw501);Aut individuals. ***PmtDNA×nuclear < 0.0001.
When developed at 28°, (simw501);OreR individuals did not produce offspring, while other genotypes maintain high culture productivity at this temperature. This heat-induced sterility was observed in both sexes; when developed at 28°, neither male nor female (simw501);OreR adults produced offspring when mated to fertile (ore);OreR individuals of the opposite sex. This effect was reversible; fertility was restored when (simw501);OreR adults developed at 28° were returned to 25° for 3–5 days, indicating that the sterility was not due to a failure to develop gonads. Because of this heat-induced sterility, the developmental time data at 28° (Figure 2) were collected from eggs laid by mothers at 25° and then placed at 28° to develop. For all remaining phenotypes, flies were measured at 16°–25° such that parents and offspring shared developmental thermal environments.
Temperature modifies mitochondrial–nuclear effects on larval performance
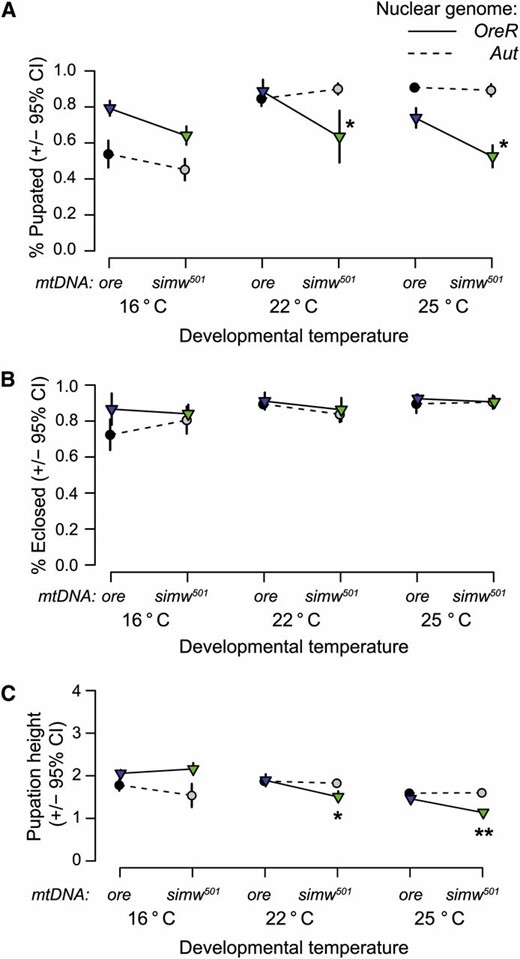
We quantified viability effects of the mitochondrial–nuclear interaction by scoring survival to pupation and survival through metamorphosis. The mitochondrial–nuclear interaction effect on survival to pupation was temperature dependent (mtDNA × nuclear × TDEV: F2, 212 = 3.63, P = 0.028). The interaction significantly decreased survival to pupation at 22° and 25° but not at 16° (Figure 3A and Table S2). Relative to the compatible genotypes, (simw501);OreR experienced a 28% and a 38% decrease in survival to pupation at 22° and 25°, respectively. However, if larvae did survive to pupation, there was no further effect of the mitochondrial–nuclear interaction on survival through metamorphosis (Figure 3B and Table S2). Survival of pupae to adult eclosion was high, even though metamorphosis time of the incompatible genotype is extended at 25° (Meiklejohn et al. 2013).
Temperature modifies the mitochondrial–nuclear interaction effect on larval, but not pupal survival. (A) There is a significant mitochondrial–nuclear–temperature interaction effect on survival from egg to pupation (% Pupated). (simw501);OreR individuals have the lowest survivorship at warmer temperatures and relatively high survivorship at 16°. The mitochondrial–nuclear interaction is significant at 22° and 25°, but not at 16° (Table S2). (B) Among larvae that do pupate, all genotypes have high survivorship during metamorphosis (% Eclosed), with >80% eclosing as adults. (C) Pupation height, a measure of larval energetic performance, is decreased in (simw501);OreR larvae reared at warmer temperatures, relative to all other genotypes. When reared at 16° there is a nearly significant mitochondrial–nuclear interaction (Table S3), but with (simw501);OreR larvae climbing as high as the highest pupating genotype. See Materials and Methods for description of y-axis. *PmtDNA×nuclear < 0.01; **PmtDNA×nuclear < 0.001.
After reaching critical weight, D. melanogaster larvae enter a wandering stage where they climb away from the food source to pupate. The height at which larvae pupate varies among species and genotypes, and pupation height is used as a correlate of energetic performance due to the high energetic cost of larval locomotion in Diptera (Sokolowski and Hansell 1983; Mueller and Sweet 1986; Berrigan and Lighton 1993). The effect of the mitochondrial–nuclear interaction on pupation height was also temperature dependent (mtDNA × nuclear × TDEV: F2, 108 = 8.96, P = 0.0003) (Figure 3C and Table S3). At 22° and 25°, (simw501);OreR larvae crawled to lower pupation heights than any other genotype (Figure 3C), suggesting compromised energetic capacity. The mitochondrial–nuclear interaction effect was nearly significant at 16° (mtDNA × nuclear: F1, 36 = 3.57, P = 0.0668). However, at this cooler temperature the sign of the epistatic effect was reversed; (simw501);OreR larvae pupated among the highest heights of any genotype (Figure 3C), consistent with the lack of deleterious phenotypic effects of this genetic interaction when larvae were reared at cooler temperatures.
Metabolic effects underlie the temperature-dependent mitochondrial–nuclear interaction
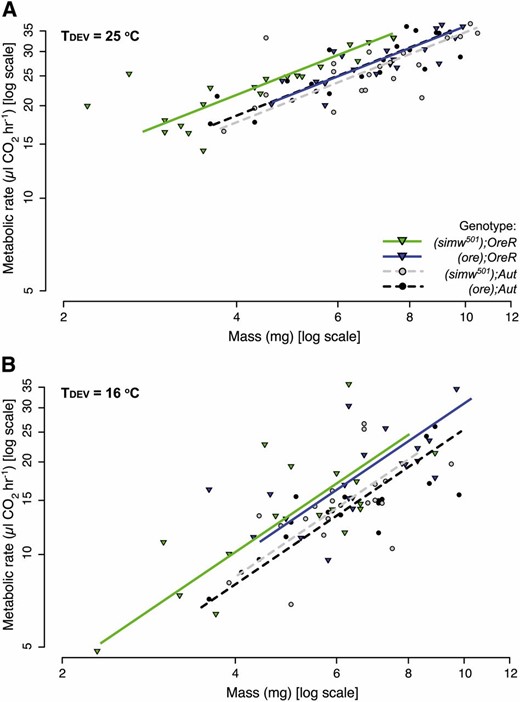
Given the decreased OXPHOS capacity associated with this mitochondrial–nuclear interaction (Meiklejohn et al. 2013) and the effects of temperature on metabolic processes (Krogh 1916; Clarke and Fraser 2004), we tested whether developmental temperature modified mitochondrial–nuclear effects on larval metabolic rates. Larval routine metabolic rates for each genotype were measured as the volume of CO2 produced by pools of five third-instar larvae while they were feeding and hydrated on a small food pellet, using flow-through respirometry at either 16° or 25°. Because ATP production via aerobic respiration is maintained by a stoichiometric equation—C6H12O6 + 6O2 → 6CO2 + 6H2O + 32ATP—the rate of CO2 production is a proxy for the rate of ATP production (Lighton 2008). Metabolic rate scales positively with mass; to account for this relationship and for variation in both traits we used standard major axis regression to test for genetic effects on metabolic rate.
There were significant effects of mtDNA, nuclear genome, and their interaction on larval mass, but these effects were the same across developmental temperatures (mtDNA × nuclear × TDEV: F1, 314 = 0.02, P = 0.8831) (Table S4). When developed at either 16° or 25°, (simw501);OreR larvae were smaller than other genotypes (Figure 4). However, for their mass, the metabolic rate of (simw501);OreR larvae was significantly higher than that of all other genotypes when developed and measured at 25° (Figure 4A), but not when developed and measured at 16° (Figure 4B and Table S5). This result suggests that (simw501);OreR larvae metabolize energy stores at a higher rate than other genotypes when developed at warmer, but not cooler temperatures. Combined with the extended development time at 25°, this indicates that energy metabolism in (simw501);OreR is less efficient, with larvae burning more energy and requiring longer development time to attain critical weight and commit to metamorphosis, while development is proceeding rapidly in other genotypes (Figure 2).
The effects of the mitochondrial–nuclear interaction on metabolic rate depend upon developmental temperature. Plots show metabolic rate as the volume of CO2 produced by pools of five larvae as a function of their mass on a log-log scale. (A) When (simw501);OreR larvae develop at 25° and are measured at 25°, they have a significantly elevated metabolic rate relative to all other genotypes (Table S5), consistent with their extended development time at this temperature (Figure 2). (B) In contrast, when larvae develop at 16° and are measured at 16°, (simw501);OreR larvae have the same metabolic rate as (ore);OreR larvae (Table S5). Similarly, at 16° there is no significant mitochondrial–nuclear effect on development time (Figure 2).
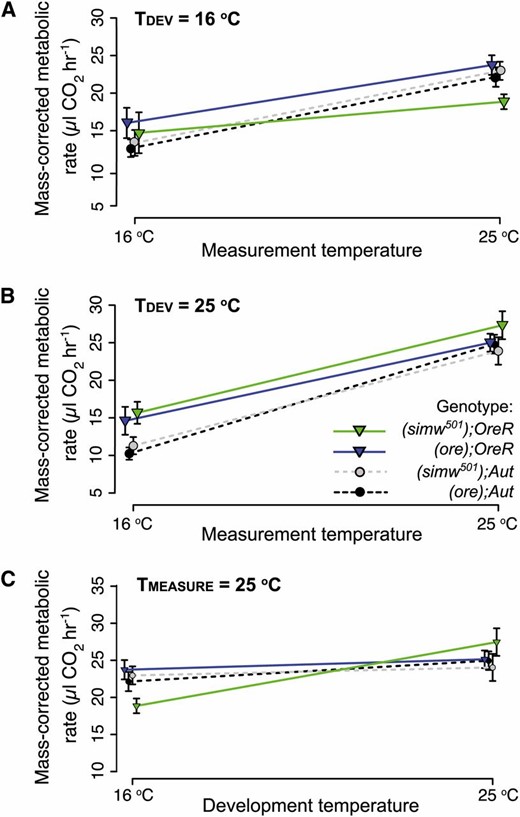
While (simw501);OreR larvae have a normal metabolic rate and development time at 16°, we observed decreased metabolic plasticity in this genotype when developed at 16°. Metabolic rate in ectotherms increases with increasing temperature, an effect known as the Q10, which can be characterized by the slope of the thermal reaction norm of metabolic rate for each genotype measured at two different temperatures (Figure 5, A and B). We measured metabolic rate at 16° and 25° for all four genotypes developed at either 16° or 25°. When developed at 16°, (simw501);OreR larvae had a lower Q10 than larvae of the other three genotypes (Figure 5A and Table S7); when developed at 25°, the Q10 of (simw501);OreR larvae was similar to that of the compatible (ore);OreR genotype (Figure 5B and Table S7). As a consequence, the metabolic rate of incompatible larvae developed at 16° was depressed relative to that of all other genotypes when measured at 25° (Figure 5C). This resulted in a significant four-way interaction between mtDNA, nuclear genome, development temperature, and measurement temperature (i.e., G × G × E × E) (mtDNA × nuclear × TDEV × TMEASURE: F1, 306 = 4.939, P = 0.027) (Table S6). Thus, the effect of the mitochondrial–nuclear interaction was revealed as a lower degree of metabolic rate plasticity but only when larvae developed at 16°. This result indicates that the capacity of developmental plasticity to compensate metabolic performance in this mitochondrial–nuclear incompatible genotype was limited.
The mitochondrial–nuclear interaction affects metabolic plasticity. Plots of the mean mass-corrected metabolic rate (±1 SEM) of each genotype reared at two development temperatures and measured at both temperatures show the effect of measurement temperature on metabolic rate (i.e., the thermal reaction norm or Q10). There is a significant four-way interaction between mtDNA, nuclear genome, development temperature, and measurement temperature (Table S6). (A) When developed at 16°, (simw501);OreR larvae have a more shallow thermal reaction norm or a lower Q10 for metabolic rate. These larvae do not elevate metabolic rate as a function of increasing temperature to the same extent as other genotypes. (B) When developed at 25°, there is an effect of the nuclear genome on the Q10, but no significant effect of the mitochondrial–nuclear interaction. (simw501);OreR larvae reared at 25° have a Q10 similar to that of (ore);OreR larvae. (C) Plotting only metabolic rates measured at 25°, (simw501);OreR larvae developed at 16° have a lower metabolic rate than all other genotypes, suggesting that development at a cooler temperature does not mitigate all of the effects of the mitochondrial–nuclear incompatibility.
Discussion
Temperature-dependent epistatic effects of a tRNA–protein interaction
We have shown that the pleiotropic effects of an epistatic interaction between mutations in a mitochondrial tRNA and its nuclear-encoded tRNA synthetase depend strongly on the environment. Nuclear-encoded mitochondrial aminoacyl-tRNA synthetases are composed of a tRNA anticodon-binding domain and a catalytic domain that aminoacylates the cognate tRNA in the mitochondrion (Bonnefond et al. 2005, 2007). The incompatible alanine-to-valine substitution in the AatmOre allele is adjacent to the conserved “KMSKS” sequence in a loop that connects the binding and catalytic domains. Alanine and valine are highly similar nonpolar amino acids, and the presence of the valine allele at 13% in a natural population of D. melanogaster suggests that this amino acid is not highly deleterious on its own (Meiklejohn et al. 2013). Nevertheless, when paired with the G:C to G:U change in the simw501 mt-tRNATyr, the valine substitution decreases OXPHOS activity (Meiklejohn et al. 2013), consistent with decreased mitochondrial protein synthesis via less efficient mt-tRNATyr aminoacylation (Jacobs 2003; Moreno-Loshuertos et al. 2011).
Conditional on the mt-tRNATyr genotype, the functional effects of the alanine-to-valine change in Aatm on development time, larval survival, and energetic performance were magnified at warmer and mitigated at cooler development temperatures. Minor changes in protein structure caused by single-amino-acid changes can have large effects on the enzyme kinetics of substrate binding that are thermally sensitive (Fields et al. 2006). Alanine-to-valine amino acid changes in phosphoglucomutase also affect enzyme function and vary in frequency across thermal gradients in natural D. melanogaster populations (Verrelli and Eanes 2001a,b). Thus, a proximate explanation for the temperature-dependent effects of this incompatibility may be that the molecular interaction itself is temperature sensitive, such that aminoacylation of the mt-tRNATyr and consequently mitochondrial protein synthesis and OXPHOS are more severely disrupted at warmer temperatures. Warmer temperatures also accelerate energy usage and development in ectotherms, increasing demand upon the ATP generated by mitochondrial–nuclear interactions (Krogh 1916; Hochachka and Somero 2002; Clarke and Fraser 2004; Ghosh et al. 2013). Thus, an alternative explanation for the observed temperature-dependent effects is that the incompatibility disrupts mt-tRNATyr aminoacylation to the same degree at all temperatures, but the energetic demands of development reveal phenotypic effects of this disruption only at warmer temperatures. These are not mutually exclusive mechanistic explanations; increased energetic demands may magnify underlying temperature-sensitive effects of the tRNA–protein interaction on aminoacylation, together revealing inefficient mitochondrial function in incompatible mitochondrial–nuclear genotypes at warmer temperatures.
The temperature-dependent effects of the mitochondrial–nuclear genotype on development are correlated with effects on metabolic rate. At 16° the incompatible (simw501);OreR genotype has a metabolic rate and development time similar to those of other genotypes. In contrast, when developed at 25°, (simw501);OreR larvae have a significantly elevated metabolic rate and a significantly lower growth rate, relative to compatible genotypes. Metabolic models of growth generally describe growth rate as a function of the assimilation of energy resources minus energy loss, largely through respiration (Winberg 1956; Thompson and Bayne 1974; Koehn and Bayne 1989). The elevated respiration of (simw501);OreR larvae at 25° may reduce growth efficiency (i.e., growth per assimilated calorie) via inefficiencies in OXPHOS that require incompatible larvae to use more energy stores at 25° to maintain ATP pools, leaving less energy to be allocated to accumulating the mass required to commit to metamorphosis, and extending development time. Compatible mitochondrial–nuclear genotypes can take advantage of accelerated growth at warm temperatures because their intact respiratory system more efficiently converts energy stores to ATP, allowing excess energy to be allocated to the rapid accumulation of larval mass that occurs in ectotherms at warm temperatures (Powsner 1935; Church and Robertson 1966).
However, thermal reaction norms for metabolic rate (Figure 5) reveal that development at cooler temperatures only partially rescues energetic performance. When developed at 16°, incompatible larvae have decreased metabolic rate plasticity (i.e., a lower Q10). As a consequence, their metabolic rate is depressed relative to that of other genotypes when measured at 25°. This clearly demonstrates that plasticity for metabolic rate has a genetic basis and, for this ectotherm, depends upon efficient mitochondrial OXPHOS function. A possible explanation for the mitochondrial–nuclear effect on metabolic rate plasticity is that mitochondrial function of (simw501);OreR larvae is sufficient for growth rates during development at 16°, but there is no excess mitochondrial capacity to dynamically elevate metabolic rate when temperature increases. This leads to the prediction that, in fluctuating environments, genotypes with energetic inefficiencies will be unable to take advantage of the accelerated growth that normally accompanies thermodynamically favorable periods of warmth during development (Behrens et al. 1983; Ruel and Ayres 1999).
Energetic limitations on the development of adult fertility
Even at 25°, where (simw501);OreR survival rates to pupation are decreased, larvae that do commit to pupation have normal rates of survival to adult eclosion, indicating that some incompatible larvae can accumulate sufficient energy stores to complete metamorphosis. However, their metamorphosis proceeds at a slower rate and the resulting adults have reduced fecundity (Meiklejohn et al. 2013), consistent with reduced allocation of energy to reproduction. The 28°-induced sterility of the mitochondrial–nuclear incompatible genotype is reversible in the adult life stage, indicating a cessation of gametogenesis rather than a failure to develop gonads. In this way, the (simw501);OreR incompatibility phenocopies temperature-sensitive Minute mutants (Sinclair et al. 1981), many of which disrupt protein synthesis and have greater phenotypic effects at warmer temperatures (Lambertsson 1998). These observations suggest that the mitochondrial inefficiencies in (simw501);Ore energetically limit the intensive protein synthesis that is needed both for developmental periods of rapid cell division and for gametogenesis (Britton and Edgar 1998; Lambertsson 1998; Audibert et al. 2005; Mandal et al. 2005; Sugiyama et al. 2006; Lee et al. 2009). Female Drosophila can reversibly arrest oogenesis in response to nutrient availability (Drummond-Barbosa and Spradling 2001; McCall 2004; Terashima and Bownes 2004). One possibility is that oogenesis and spermatogenesis both have checkpoints that are sensitive to genetic and environmental factors that compromise metabolic capacity for protein synthesis.
Drosophila males experience heat-induced sterility at sharply defined critical temperatures that leave females fertile (Rohmer et al. 2004), and closely related species have diverged in this critical temperature (Parsons 1973; Rohmer et al. 2004; David et al. 2005). The sterility of (simw501);OreR males at 28° occurs more than a full degree centigrade below that required to induce sterility in heat-sensitive temperate D. melanogaster and matches the critical temperature for heat-induced male sterility in D. simulans (Chakir et al. 2002; David et al. 2005). Heat-induced male sterility in Drosophila is the consequence of defects in spermatid elongation that are reversible (Rohmer et al. 2004; David et al. 2005), similar to the reversibility in male sterility that we observed. It is thus possible that there is an energetic component to heat-induced male sterility or that the energetic defect in (simw501);OreR acts synergistically with the mechanisms of heat-induced male sterility to decrease the critical temperature for gametogenesis to proceed.
Evolutionary implications
The strong G × G × E and G × G × E × E interactions that we observed clearly demonstrate that the fitness effects of metabolic mutations can be highly conditional on both internal genetic and external nongenetic environments. The dependence of mitochondrial function on RNA–protein and protein–protein interactions provides a large target for mutations with epistatic effects on metabolism (Burton and Barreto 2012). Furthermore, the ubiquitous effect of temperature on molecular, metabolic, and developmental processes suggests that these epistatic effects on energy metabolism in ectotherms should be broadly temperature dependent. Because their fitness effects are realized only in a fraction of genetic backgrounds and environments, interacting mutations can segregate at higher frequencies relative to the expectation for alleles with unconditional deleterious effects. While the conditions for either cytonuclear interactions or environment dependence of mutational effects to maintain polymorphisms in populations are restricted (Clark 1984; Gregorius and Ross 1984; Rand et al. 2001; Turelli and Barton 2004), both epistasis and genotype-by-environment interactions affect the efficacy by which selection fixes beneficial and removes deleterious mutations in populations (Whitlock 1996; Phillips and Johnson 1998; Van Dyken and Wade 2010). Alleles with synthetic deleterious effects, such as those that we have characterized, are expected to segregate at higher frequencies under mutation–selection balance (Phillips and Johnson 1998; Lachance et al. 2011), and populations may harbor these types of synthetic deleterious alleles as cryptic variants when the permissive genetic background is fixed (Lachance et al. 2011).
Genotype-by-environment interactions are also predicted to facilitate the accumulation of genetic incompatibilities among reproductively isolated and locally adapted populations (Wade 2000; Bordenstein and Drapeau 2001). We have shown that the effects of these types of interacting mutations can be revealed when temperature increases the rates of molecular and metabolic processes. If cold-adapted populations generally harbor higher levels of synthetically deleterious alleles that are masked at cold temperatures and revealed at warm temperatures, this may explain the commonly observed asymmetric patterns of fitness effects between cold- and warm-adapted populations in common-garden experiments. Thus, the observation that “hotter is better (or broader)” (Frazier et al. 2006; Knies et al. 2009) could result from a greater load of segregating synthetically deleterious alleles in cold-adapted populations rather than from an unconditional fitness advantage conferred across a range of environments by hot-adapted alleles.
Finally, we posit that the accelerating effects of increasing temperature may generally worsen molecular incompatibilities and metabolic inefficiencies that underlie organismal fitness in ectotherms. In fact, at least 15 studies have identified temperature-dependent postzygotic hybrid incompatibilities (Bordenstein and Drapeau 2001; Willett and Burton 2003; Bomblies et al. 2007). Increasing temperature can magnify deleterious effects in hybrids (e.g., Hutter and Ashburner 1987; Wade et al. 1999), but some hybrid incompatibilities are masked by warmer temperatures (Bomblies et al. 2007) or have a more complex relationship with temperature (Willett and Burton 2003; Matute et al. 2009). This suggests that whether temperature magnifies or mitigates epistatic genetic effects may depend upon the particular molecular interaction or its physiological consequences. Nevertheless, the prevalence of genotype-by-environment interactions underlying fitness effects in hybrids warrants continued investigation of how the abiotic environment may shape geographic patterns of hybrid incompatibilities (e.g., Yukilevich 2013).
Acknowledgments
We thank Sable Systems International for technical support and N. Johnson, C. Meiklejohn, and M. Wade for their constructive feedback. This research was supported by a National Science Foundation Integrative Graduate Education and Research Traineeship (NSF-IGERT) 206251A-A03 and a Society for Comparative and Integrative Biology grant-in-aid of research (to L.A.H.), research grants from the Indiana University Hutton Honors College (to M.A.S.), and funding from Indiana University and NSF CAREER award IOS-1149178 (to K.L.M.).
Footnotes
Communicating editor: S. F. Chenoweth
Literature Cited
Johnston, I. A., 1990 Cold adaptation in marine organisms. Philos. Trans. R Soc. Lond. B Biol. Sci. 326: 655–666, discussion 666–657.
Winberg, G. G., 1956 Rates of Metabolism and Food Requirements of Fishes (Fisheries Research Board of Canada. Translational Series 194). BeloRussian State University, Minsk, Belarus.
Author notes
Supporting information is available online at http://www.genetics.org/lookup/suppl/doi:10.1534/genetics.113.154914/-/DC1.
Present address: Department of Ecology and Evolution, University of Chicago, Chicago, IL 60637.