-
PDF
- Split View
-
Views
-
Cite
Cite
Christopher H Corzett, Myron F Goodman, Steven E Finkel, Competitive Fitness During Feast and Famine: How SOS DNA Polymerases Influence Physiology and Evolution in Escherichia coli, Genetics, Volume 194, Issue 2, 1 June 2013, Pages 409–420, https://doi.org/10.1534/genetics.113.151837
Close - Share Icon Share
Abstract
Escherichia coli DNA polymerases (Pol) II, IV, and V serve dual roles by facilitating efficient translesion DNA synthesis while simultaneously introducing genetic variation that can promote adaptive evolution. Here we show that these alternative polymerases are induced as cells transition from exponential to long-term stationary-phase growth in the absence of induction of the SOS regulon by external agents that damage DNA. By monitoring the relative fitness of isogenic mutant strains expressing only one alternative polymerase over time, spanning hours to weeks, we establish distinct growth phase-dependent hierarchies of polymerase mutant strain competitiveness. Pol II confers a significant physiological advantage by facilitating efficient replication and creating genetic diversity during periods of rapid growth. Pol IV and Pol V make the largest contributions to evolutionary fitness during long-term stationary phase. Consistent with their roles providing both a physiological and an adaptive advantage during stationary phase, the expression patterns of all three SOS polymerases change during the transition from log phase to long-term stationary phase. Compared to the alternative polymerases, Pol III transcription dominates during mid-exponential phase; however, its abundance decreases to <20% during long-term stationary phase. Pol IV transcription dominates as cells transition out of exponential phase into stationary phase and a burst of Pol V transcription is observed as cells transition from death phase to long-term stationary phase. These changes in alternative DNA polymerase transcription occur in the absence of SOS induction by exogenous agents and indicate that cell populations require appropriate expression of all three alternative DNA polymerases during exponential, stationary, and long-term stationary phases to attain optimal fitness and undergo adaptive evolution.
THE generation of genetic diversity directly impacts the evolutionary fitness of a population, and the molecular mechanisms influencing the formation of allelic variation often dictate success or failure within complex bacterial communities (Chao and Cox 1983; Bjedov et al. 2003; Saint-Ruf and Matic 2006; Pigliucci 2008; Woods et al. 2011). Given that the majority of mutations are introduced during replication, perturbations in the fidelity of replication can have dramatic consequences on the evolutionary trajectory of a population (Yeiser et al. 2002; Kunkel 2004; Foster 2007; Galhardo et al. 2007). Accordingly, characterization of the mechanisms and of the extent to which DNA polymerases introduce genetic variation is critical to understanding the physiology and evolution of bacteria.
Escherichia coli encodes five DNA polymerases (Goodman 2002; Johnson and O’Donnell 2005; Friedberg 2006). High-fidelity DNA polymerase III performs the majority of DNA replication under vegetative conditions, with Pol I contributing principally to maturation of Okazaki fragments (Kornberg and Baker 1992; Friedberg 2006). Three alternative DNA polymerases (Pol II, Pol IV, and Pol V) perform a vital physiological role by mediating translesion synthesis, enabling efficient replication past DNA damage that would otherwise halt replication, albeit with significantly reduced fidelity (Goodman 2002; Fuchs et al. 2004; Tippin et al. 2004; Nohmi 2006; Bichara et al. 2011). These error-prone DNA polymerases can be induced under a variety of environmental stresses (Taddei et al. 1995; Yeiser et al. 2002; Layton and Foster 2003, 2005; Stumpf and Foster 2005; MacPhee and Ambrose 2010) and have been characterized most extensively following induction of the SOS regulon in response to DNA damage, leading them to be referred to as SOS-induced polymerases (Courcelle et al. 2001; Goodman 2002; Friedberg 2006; Nohmi 2006; Yang and Woodgate 2007). The SOS regulon, which contains >40 genes, was first discovered as a response to DNA damage caused by exposure to UV irradiation or mutagenic chemicals (Witkin 1976, Walker 1984). DNA polymerase II (encoded by polB) is a B-family polymerase (Rangarajan et al. 1999, 2002; Pham et al. 2001; Banach-Orlowska et al. 2005) capable of 3′-exonuclease proofreading, enabling it to replicate undamaged DNA with considerable accuracy (Cai et al. 1995). DNA polymerases IV (dinB) and V (umuDC) are Y-family polymerases (Ohmori et al. 2001; Goodman 2002; Nohmi 2006) that replicate DNA with relatively lower fidelity (Tang et al. 2000; Fuchs et al. 2004; Jarosz et al. 2007). Homologs of these alternative polymerases are found in all three domains of life (Ohmori et al. 2001) and have been implicated in a variety of human diseases (Robbins et al. 1974, 1975; Stallons and McGregor 2010; Xie et al. 2010; Lange et al. 2011).
As a consequence of their physiological roles and comparatively low fidelity, considerable interest has focused on characterizing the impact of each alternative polymerase on the formation of genetic diversity, their impact on cell survival during stress, and the dynamics of polymerase competition for access to the replication fork (Yeiser et al. 2002; Delmas and Matic 2006; Nohmi 2006; Petrosino et al. 2009; Hastings et al. 2010; Shee et al. 2011). While most in vivo studies have focused on the roles of Pols II, IV, and V either in the survival of rapidly dividing cells in the presence of artificial DNA damage sufficient to activate the E. coli SOS response or in a plate-based assay system studying the process of “adaptive mutation” (Foster 2005, 2007; Roth et al. 2006; Galhardo et al. 2007), recent findings have also suggested that growth conditions can influence which polymerases play greater roles in replication and alter the spectrum of mutations generated (Nowosielska et al. 2004).
Given the well-characterized roles of the alternative polymerases in the generation of adaptive mutations within these assays, numerous authors have speculated that the alternative polymerases might serve to mediate stress-induced mutagenesis and influence the survival and evolution of bacterial populations. We have previously found (Yeiser et al. 2002) that, in the absence of exogenous DNA damage, Pols II, IV, and V provide a substantial fitness advantage to wild-type cells during long-term stationary phase competition, including expression of the growth advantage in stationary phase (GASP) phenotype where mutants of increased fitness are isolated after long-term incubation in stationary phase (Zambrano et al. 1993; Finkel 2006). However, Yeiser et al. (2002) used strains with only single DNA polymerase gene deletions, making it impossible to assign specific activities to each enzyme.
Here we investigate the roles and relative contributions of Pols II, IV, and V to cell growth and evolutionary fitness using a series of isogenic mutant strains lacking each alternative polymerase in all possible combinations: as single null mutants, as combinations of double null mutants that express only one alternative polymerase (referred to here as Pol II+-only, Pol IV+-only, and Pol V+-only), and as a triple null mutant. This study provides a comprehensive analysis of the expression patterns and the roles of Pols II, IV, and V in the absence of exogenous DNA damage. We determine each polymerase’s contribution toward cell survival and evolutionary fitness during periods of feast—in exponential batch culture growth where nutrients are abundant or in a chemostat where nutrients are continually being replenished—and during periods of famine as cells enter stationary phase and transition into long-term stationary phase where nutrients are being depleted. These data demonstrate significant evolutionary consequences and establish individual roles for each polymerase under conditions where different strains must compete to survive. We show that, while any one of the polymerases can randomly generate a mutation that enables a cell to survive, there are nevertheless conditions during batch culture where the activity of each individual enzyme is either dominant or codominant, enabling the prediction of evolutionary outcomes. While previous studies on the regulation of the alternative polymerases have focused on increased transcription in response to DNA damage from exogenous agents, here we observe major changes in the transcription pattern of each alternative polymerase throughout the five phases of the bacterial life cycle (Finkel 2006) in the absence of induction by exogenous stressors, demonstrating a remarkable shift toward error-prone polymerase expression under stationary-phase conditions.
Materials and Methods
Strains used and mutant construction
All strains (Table 1) are derived from E. coli K-12 strain ZK126 (W3110 ΔlacU169 tna-2), including nalidixic acid-resistant parental strain ZK1142 (Zambrano et al. 1993). DNA polymerase single, double, and triple mutants were constructed by bacteriophage P1 transduction into ZK126 using the following donor strains: for Pol II−, SH2101 (polB::Spc) (Bonner et al. 1992); for Pol IV−, RW626 (dinB::Kan); and for Pol V−, RW82 (umuDC::Cam) (both RW626 and RW82 were generous gifts from Roger Woodgate, National Institutes of Health, Bethesda, MD). Genetic elements conferring antibiotic resistance are effectively neutral in the absence of drug selection (Yeiser et al. 2002; Kraigsley and Finkel 2009). Strains lacking a single alternative polymerase are designated with a superscript “minus” sign (Pol II−, Pol IV−, and Pol V−), whereas double-mutant strains capable of expressing only one alternative polymerase are designated with a superscript “plus” sign coupled with “-only” (Pol II+-only, Pol IV+-only, and Pol V+-only).
Strains used in this study
| Strain . | Relevent genotype/phenotype . | Nomenclature . | Pol II . | Pol IV . | Pol V . | Reference . |
|---|---|---|---|---|---|---|
| ZK126 | W3110 ΔlacU169 tna-2 | Wild type | + | + | + | Zambrano et al. (1993) |
| ZK1142 | ZK126 NalR | Wild type | + | + | + | Zambrano et al. (1993) |
| SF2003 | ZK126 polB::SpcR | Pol II− | — | + | + | Yeiser et al. (2002) |
| SF2006 | ZK126 dinB::KanR | Pol IV− | + | — | + | Yeiser et al. (2002) |
| SF2009 | ZK126 umuDC::CamR | Pol V− | + | + | — | Yeiser et al. (2002) |
| SF2012 | ZK126 polB::SpcR dinB::KanR | Pol V+-only | — | — | + | This study |
| SF2014 | ZK126 polB::SpcR umuDC::CamR | Pol IV+-only | — | + | — | This study |
| SF2016 | ZK126 dinB::KanR umuDC::CamR | Pol II+-only | + | — | — | This study |
| SF2018 | ZK126 polB::SpcR dinB::KanR umuDC::CamR | Triple mutant | — | — | — | Yamanaka et al. (2011) |
| Strain . | Relevent genotype/phenotype . | Nomenclature . | Pol II . | Pol IV . | Pol V . | Reference . |
|---|---|---|---|---|---|---|
| ZK126 | W3110 ΔlacU169 tna-2 | Wild type | + | + | + | Zambrano et al. (1993) |
| ZK1142 | ZK126 NalR | Wild type | + | + | + | Zambrano et al. (1993) |
| SF2003 | ZK126 polB::SpcR | Pol II− | — | + | + | Yeiser et al. (2002) |
| SF2006 | ZK126 dinB::KanR | Pol IV− | + | — | + | Yeiser et al. (2002) |
| SF2009 | ZK126 umuDC::CamR | Pol V− | + | + | — | Yeiser et al. (2002) |
| SF2012 | ZK126 polB::SpcR dinB::KanR | Pol V+-only | — | — | + | This study |
| SF2014 | ZK126 polB::SpcR umuDC::CamR | Pol IV+-only | — | + | — | This study |
| SF2016 | ZK126 dinB::KanR umuDC::CamR | Pol II+-only | + | — | — | This study |
| SF2018 | ZK126 polB::SpcR dinB::KanR umuDC::CamR | Triple mutant | — | — | — | Yamanaka et al. (2011) |
| Strain . | Relevent genotype/phenotype . | Nomenclature . | Pol II . | Pol IV . | Pol V . | Reference . |
|---|---|---|---|---|---|---|
| ZK126 | W3110 ΔlacU169 tna-2 | Wild type | + | + | + | Zambrano et al. (1993) |
| ZK1142 | ZK126 NalR | Wild type | + | + | + | Zambrano et al. (1993) |
| SF2003 | ZK126 polB::SpcR | Pol II− | — | + | + | Yeiser et al. (2002) |
| SF2006 | ZK126 dinB::KanR | Pol IV− | + | — | + | Yeiser et al. (2002) |
| SF2009 | ZK126 umuDC::CamR | Pol V− | + | + | — | Yeiser et al. (2002) |
| SF2012 | ZK126 polB::SpcR dinB::KanR | Pol V+-only | — | — | + | This study |
| SF2014 | ZK126 polB::SpcR umuDC::CamR | Pol IV+-only | — | + | — | This study |
| SF2016 | ZK126 dinB::KanR umuDC::CamR | Pol II+-only | + | — | — | This study |
| SF2018 | ZK126 polB::SpcR dinB::KanR umuDC::CamR | Triple mutant | — | — | — | Yamanaka et al. (2011) |
| Strain . | Relevent genotype/phenotype . | Nomenclature . | Pol II . | Pol IV . | Pol V . | Reference . |
|---|---|---|---|---|---|---|
| ZK126 | W3110 ΔlacU169 tna-2 | Wild type | + | + | + | Zambrano et al. (1993) |
| ZK1142 | ZK126 NalR | Wild type | + | + | + | Zambrano et al. (1993) |
| SF2003 | ZK126 polB::SpcR | Pol II− | — | + | + | Yeiser et al. (2002) |
| SF2006 | ZK126 dinB::KanR | Pol IV− | + | — | + | Yeiser et al. (2002) |
| SF2009 | ZK126 umuDC::CamR | Pol V− | + | + | — | Yeiser et al. (2002) |
| SF2012 | ZK126 polB::SpcR dinB::KanR | Pol V+-only | — | — | + | This study |
| SF2014 | ZK126 polB::SpcR umuDC::CamR | Pol IV+-only | — | + | — | This study |
| SF2016 | ZK126 dinB::KanR umuDC::CamR | Pol II+-only | + | — | — | This study |
| SF2018 | ZK126 polB::SpcR dinB::KanR umuDC::CamR | Triple mutant | — | — | — | Yamanaka et al. (2011) |
Culture conditions, media, and titering assays
Strains were cultured in 5.0 ml LB broth, Lennox (Difco-BD), and incubated at 37° with aeration in a TC-7 test tube roller (New Brunswick Scientific), unless otherwise specified. Experiments were initiated from overnight cultures inoculated from frozen LB–glycerol stocks. Viable counts were determined by serial dilution of cells periodically removed from cultures and plating on selective medium containing the appropriate antibiotics: nalidixic acid (Nal) (20 μg/ml); streptomycin (Str) (25 μg/ml); spectinomycin (Spc) (100 μg/ml); kanamycin (Kan) (50 μg/ml); chloramphenicol (Cam) (30 μg/ml); and rifampicin (Rif) (100 μg/ml). This method of titering is accurate within plus-or-minus threefold (Kraigsley and Finkel 2009) and has a limit of detection of 1000 colony-forming units (CFU)/ml in this study.
Batch culture long-term competition assays
Two types of batch culture competitions, distinguished by their initial cell densities, were performed: type 1, where both strains are mixed at a low initial density (∼106 CFU/ml) and co-outgrown, and type 2, where high-density stationary-phase cultures are mixed (∼109 CFU/ml). In type 1 experiments, competitions were initiated by inoculating 5 μl of each competing strain (1:1000 dilution, vol:vol) into the same 5.0-ml LB culture. In type 2 experiments, competitions were initiated by combining 2.5 ml of overnight stationary-phase cultures of each strain (1:1 mix, vol:vol). Viable counts were determined as described above using the appropriate combinations of antibiotics (Kan/Cam for Pol II+-only, Spc/Cam for Pol IV+-only, and Spc/Kan for Pol V+-only). At the conclusion of each 14-day competition, strains showing a 10-fold greater relative population density were scored as winners.
Serial passage aging regimen
To observe changes in relative fitness following repeated outgrowth, strains were serially passaged in 5.0-ml LB cultures. Independent cultures of each strain were incubated for 24 hr, sampled, and diluted 1:1000 (vol:vol) into fresh medium. Cultures were passaged daily for 5 days, and frozen LB–glycerol stocks were prepared following each passage.
Selection of GASP mutants and GASP competitions
To select for GASP mutants, strains were inoculated into independent 5.0-ml LB cultures and incubated for 10 days, and 150 μl was frozen as LB–glycerol stocks. To initiate GASP competitions, an overnight culture of each aged population was grown and introduced as a minority at a 1:1000 (vol/vol) dilution into a 5.0-ml culture of an unaged population of wild-type cells (ZK1142) or polymerase mutant strains (Zambrano et al. 1993). Population densities for each strain were determined by titering, as described above.
Chemostat competitions
To assess the relative fitness of polymerase-deficient strains under conditions promoting rapid growth, competitions were performed under continuous culture conditions in chemostats (Harder and Kuenen 1977; Chao and Cox 1983). For each competition, 250 μl (1:3000, vol:vol) of each double-mutant strain (Pol II+, IV+, and V+) was inoculated into 750 ml LB in a BioFlo 110 bioreactor (New Brunswick Scientific). The chemostat was run for up to 2 hr under batch conditions to obtain the desired density before initiating flow of fresh medium into the growth chamber. Average dilution rates varied from 1 to 4 vol/hr, and chemostats were run between ∼6 to 10 hr. The chemostat culture was regularly sampled to monitor optical density and determine viable counts.
Mutation frequency assay
The frequency of spontaneous rifampicin resistance (RifR) was determined in wild-type and all seven polymerase-deficient strains. For each strain, 159 independent 5.0-ml overnight cultures were grown, and 100 μl of each was spread onto plates containing rifampicin. Total cell counts were determined by plating an appropriate dilution of each culture on LB agar. The number of RifR colonies was determined after 48 hr of incubation at 37°. The frequency of spontaneous rifampicin resistance was calculated by dividing the number of RifR CFU/ml by the total CFU/ml. The distributions of observed RifR frequencies were compared using the two-sample Kolmogorov–Smirnov (K–S) test (P < 0.05) (http://www.physics.csbsju.edu/stats/KS-test.html). This nonparametric test can be applied to compare the cumulative distribution function of two empirical distributions of continuous data to determine the likelihood that both sets were obtained from the same distribution. Importantly, this test statistic has the advantage that it is not contingent upon assumptions of normalcy in the data and is capable of assessing the cumulative distribution of mutation frequencies rather than relying strictly upon mean or median values that might not accurately reflect alterations across the cumulative distribution of the data. Differences in average RifR frequencies were assessed using SEM.
RifR mutant sequencing
The sequences of RifR mutants were determined using cells obtained from the mutation frequency assay described above. The RifR colony nearest the center of each plate was resuspended in 20 μl LB. Two microliters was used as template for PCR amplification of the rpoB gene using primer 1 (5′-AATGTCAAATCCGTGGCGTG-3′) and primer 2 (5′-TTCACCCGGATACATCTCGTC-3′) with the remaining sample frozen in LB–glycerol. Amplified products were sequenced at High-Throughput Sequencing Solutions (Seattle) using primer 1 or primer 3 (5′-CGTCGTATCCGTTCCGTTGG-3′) specific for cluster I and cluster II, respectively (Garibyan et al. 2003).
Quantitative RT-PCR
Real-time RT-PCR was used to determine the expression patterns of each alternative polymerase gene, as well as induction of the SOS response. LB cultures inoculated with a 1:1000 dilution of an overnight wild-type population were incubated at 37° and periodically sampled for RNA extraction. Samples were treated with RNAlater (Qiagen), and total RNA was isolated using the RNeasy Mini Kit (Qiagen). Quantitative RT-PCR (qRT-PCR) reactions with 100 ng template RNA were performed using the one-step RT-PCR kit (Qiagen) with SYBR Green (Molecular Probes) and amplified on an Opticon-2 real-time PCR Cycler (MJ Research). Primers used for amplification are provided in Table S4. Relative transcript abundance and changes in gene expression were determined using the 2−ΔCt method (Livak and Schmittgen 2001). In control experiments with artificial SOS induction, mitomycin C (Sigma) was added to 1 μg/ml final concentration after 2 hr of incubation at an OD600 of ∼0.1, and RNA was sampled hourly.
Results
Pol IV and Pol V confer greater relative fitness than Pol II during long-term stationary phase
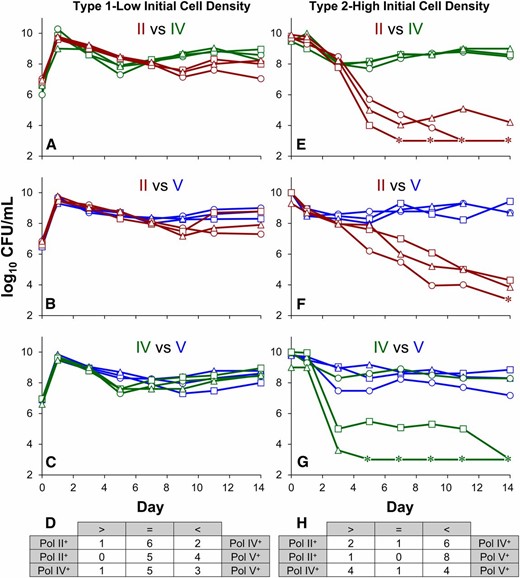
When cultured individually in rich medium, log-phase growth and stationary-phase survival of all seven polymerase-deficient strains is indistinguishable from wild type (Supporting Information, Figure S1). To determine whether strains capable of expressing only one alternative polymerase display altered fitness, double mutants were competed against one another in type 1, initial low cell density (Figure 1, A–C), and in type 2, initial high cell density (Figure 1, E–G), long-term competition experiments. In type 1 experiments, both strains are co-inoculated at low initial population density (∼106 CFU/ml), passing through lag and exponential phase, and cocultured for 2 weeks. In type 2 experiments, dense stationary-phase cultures of each strain are mixed at high initial density (∼109 CFU/ml), after which strains are co-incubated for 2 weeks.
Type 1 and type 2 competition experiments between double-mutant strains. Representative type 1 low initial cell density (A–C) and type 2 high initial cell density (E–G) competitions between unaged polymerase-deficient strains: red lines, Pol II+-only; green lines, Pol IV+-only; blue lines, Pol V+-only. Color-coded Roman numerals in each panel refer to the polymerases expressed in each pair of mutant strain competitions. Three representative competitions are shown where squares, circles, and triangles indicate competition pairs. Competition data, summarized in D and H, reflect the results of nine competition experiments for each of the three pairings of mutant strains. The “>” sign indicates that the strain listed on the left was the “winner”; the “<” sign on the right indicates that the strain listed on the right was the “winner.” One strain outcompeting the other is defined by a >10-fold difference in cell yield on day 14. When final yields are within 10-fold, no winners were declared as reflected by the “=” sign. Asterisks indicate that cell titers were below the limit of detection (<1000 CFU/ml.)
In type 1 competitions, initiated at low population density, most competitions (59%) ended as ties, with both strains reaching final densities within 10-fold of each other (Figure 1D). Among those competitions where one strain clearly dominates, the Pol V+-only strain has an advantage over both the Pol IV+-only and the Pol II+-only strains. However, in type 2 competitions, initiated with stationary-phase populations at high cell density, there was a decisive winner in nearly all (93%) competitions (Figure 1H). Strains expressing only Pol II were significantly less fit than strains expressing only Pol IV (Figure 1E) or Pol V (Figure 1F), losing in 78% of competitions. In competitions between Pol IV+-only and Pol V+-only, both strains performed equally well, with Pol IV+-only winning as many competitions as Pol V+-only (Figure 1G).
Pol II enables cells to express the GASP phenotype faster than Pol IV or Pol V
To assess the capacity of each alternative polymerase to generate beneficial alleles, each double-mutant strain was subjected to GASP competition assays. In a typical GASP assay, strains are aged for 10 days in monoculture to allow spontaneous random mutants to take over the population. Cells from these cultures are then introduced as a minority to a dense culture of unaged wild type cells (Zambrano et al. 1993). After incubating in monoculture for 10 days, all three aged mutant strains were able to outcompete the unaged wild-type population when introduced as a minority (Figure 2, A–C), indicating the presence of a beneficial GASP allele. However, the aged Pol II+-only (Figure 2A) and Pol V+-only (Figure 2C) populations drove unaged wild-type populations to extinction faster than aged Pol IV+-only (Figure 2B).
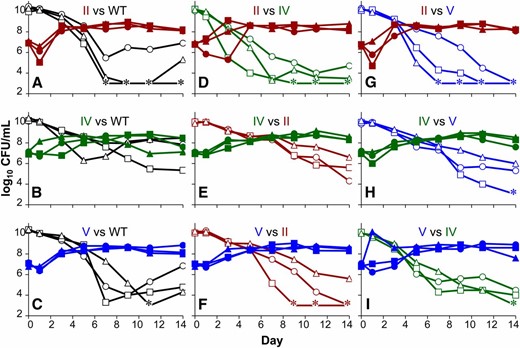
GASP competitions between wild-type and double-mutant strains. Polymerase double-mutant strains were aged for 10 days and competed to determine their GASP phenotypes against unaged wild-type strains (A–C) or each unaged polymerase mutant strain (D–I) in all possible combinations. Solid symbols correspond to strains aged for 10 days; open symbols correspond to unaged strains. Strains are indicated by line color: wild type, black; Pol II+-only, red; Pol IV+-only, green; Pol V+-only, blue. Unaged wild-type cells were competed against 10-day-old (A) Pol II+-only, (B) Pol IV+-only, or (C) Pol V+-only. Aged Pol II+-only strains were competed against unaged (D) Pol IV+-only or (G) Pol V+-only; aged Pol IV+-only strains were competed against unaged (E) Pol II+-only or (H) Pol V+-only; and aged Pol V+-only strains were competed against unaged (F) Pol II+-only or (I) Pol IV+-only. Three representative competitions are shown for each pair where squares, circles, and triangles indicate competition pairs. Asterisks indicate that titers were below the limit of detection (<1000 CFU/ml.) Color-coded Roman numerals in each panel refer to the polymerases expressed in each pair of mutant strain competitions.
We also determined the GASP phenotype of each polymerase-deficient strain with respect to one another, rather than the wild type. Again, every aged population expressed the GASP phenotype over unaged populations (Figure 2, D–I); however, the strength of the GASP phenotype differed among strains as determined by the time it took for the minority population to take over the culture. For each competition, the day on which the minority became the majority was determined, and the average day of takeover was calculated. The Pol II+-only strain consistently exhibited the GASP phenotype fastest, with an average time to takeover of 3.5 days, compared to 4.5 or 5.8 days for the strains capable of expressing only Pol V or Pol IV, respectively.
Pol II contributes to relative fitness more than Pol IV and Pol V during serial passage
Although the Pol II+-only strain was significantly less fit than Pol IV+-only and Pol V+-only in type 2 stationary-phase competitions (Figure 1, E and F) in type 1 competitions initiated at low cell densities the Pol II+-only strain shows less of a competitive disadvantage (Figure 1, A and B). Pol II+-only cell yields are routinely four to five orders-of-magnitude lower than their competitors in type 2 competitions, but only 10- to 100-fold lower in type 1 experiments. The critical difference between these experiments is that in type 1 competitions cells experience an additional outgrowth, providing more opportunity to replicate and generate genotypic diversity under nutrient-rich conditions. Since this additional outgrowth on the first day of competition profoundly influenced the fitness of the Pol II+-only strain weeks later, we determined whether subjecting each strain to five serial passages prior to competition would amplify this effect.
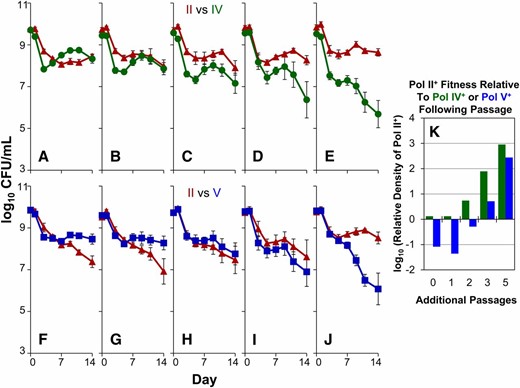
Following serial passage, Pol II+-only populations were competed against unpassaged Pol IV+-only (Figure 3, A–E) or Pol V+-only (Figure 3, F–J) to assess changes in fitness. Prior to the passaging regimen, the unpassaged Pol II+-only populations never outcompete Pol IV+-only or Pol V+-only strains in stationary-phase competitions (Figure 3, A and F). However, following each additional passage the Pol II+-only populations displayed an increase in relative fitness to the extent that Pol II+-only populations passaged five times consistently outcompeted both Pol IV+-only and Pol V+-only strains (Figure 3, E and J). Furthermore, the Pol II+-only strain passaged five times also performed better against similarly passaged Pol IV+-only and Pol V+-only strains, compared to its performance prior to passaging (data not shown).
Increased relative fitness of Pol II+-only mutants following serial passage. Populations of unaged Pol IV+-only (green lines, A–E) or unaged Pol V+-only (blue lines, F–J) were each competed against the Pol II+-only strain (red lines) after one to five serial passages. (A and F) Competition between the Pol II+-only strain with no additional passage. (B and G) Competitions after one additional passage of Pol II+-only. (C and H) Two additional passages of Pol II+-only. (D and I) Three additional passages of Pol II+-only. (E and J) Five additional passages of Pol II+-only. The average relative log10 ratio of final cell densities between Pol II+-only vs. Pol IV+-only (green bars) or Pol V+-only (blue bars) for all six conditions is plotted in K.
Pol II confers greater relative fitness than Pol IV or Pol V during continuous culture
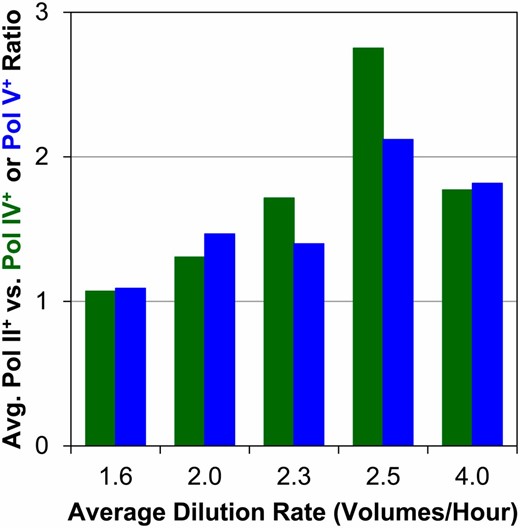
To determine the relative physiological contributions of each polymerase under conditions promoting high rates of cell division, a series of competitions were performed under continuous culture conditions in a chemostat. The three strains encoding a single alternative polymerase were cocultured in chemostats at increasing flow rates, corresponding to faster growth rates. In every competition Pol II+-only outperformed both Pol IV+-only and Pol V+-only (Figure 4), with average cell yields ∼10% more at lower flow rates and as much as 250% more at higher flow rates.
Relative fitness during chemostat competitions. The relative fitness of Pol II+-only compared to Pol IV+-only (in green) and Pol V+-only (in blue) during continuous culture competitions is shown for chemostats run with different dilution rates (volumes per hour).
Alternative polymerases affect mutation frequency and spectrum
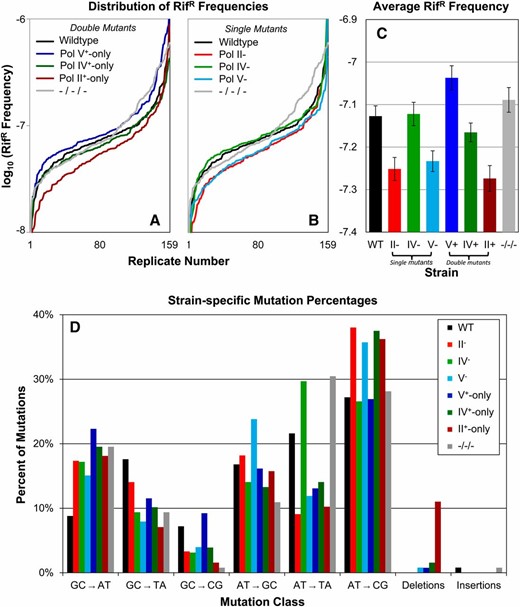
To elucidate the molecular basis of the differences in adaptive potential observed during long-term competition, we characterized the frequency and spectrum of spontaneous mutations in rpoB, encoding the β-subunit of RNA polymerase, conferring RifR. Mutations known to confer drug resistance include all six classes of missense mutations, as well as in-frame amplifications and deletions (Jin and Gross 1988; Singer et al. 1993; Severinov et al. 1994; Reynolds 2000; Garibyan et al. 2003; Wolff et al. 2004; Wrande et al. 2008; Makiela-Dzbenska et al. 2011). A total of 1272 independent cultures were assayed: 159 cultures each for the wild type, all three single-mutant, all three double-mutant, and the triple-mutant strains; significant strain-specific differences in mutation frequencies were observed (Figures 5, A–C). Among double-mutant strains, Pol V+-only had the greatest mutation frequency, significantly higher than Pol IV+-only, which in turn was significantly higher than that of Pol II+-only. Significant differences in mutation frequency distributions were determined using the K–S test (Figure 5, A and B), and average RifR values also reflect these shifts in mutation frequency (Figure 5C). Among single-mutant strains, Pol IV− was similar to wild type while Pol II− and Pol V− showed significantly lower mutation frequencies (Figure 5, B and C). The triple-mutant strain showed an overall mutation frequency similar to the wild type.
Strain-specific RifR mutation frequency and spectrum. The frequency of spontaneous rifampicin resistance was determined for 159 independent replicates of the wild-type and each polymerase mutant strain. Mutation frequencies are presented in ascending order. (A) Mutation frequencies for the wild-type (black), all three double-mutant strains (Pol II+-only, red; Pol IV+-only, green; Pol V+-only, blue), and the triple mutant (gray). (B) Mutation frequencies for the wild-type (black), all three single-mutant strains (Pol II−, red; Pol IV−, green; Pol V−, blue), and the triple mutant (gray). (C) Average RifR frequencies for the wild type, all three single-mutant strains, all three single- and double-mutant strains, and the triple mutant. Error bars denote ±SEM. (D) For all eight strains, each class of mutation is presented as a percentage of all mutations observed. Detailed mutation data are presented in Table S1, Table S2, and Table S3.
We also assessed the spectrum of mutations generated in each strain by sequencing rpoB in individual RifR clones (Figure 5D). Of 1009 sequenced isolates, we identified 85 alleles across 54 nucleotide positions (Table S1 and Table S2), with several alleles that appear to be previously unreported (indicated in Table S1). Strain-specific mutation spectrum differences were observed. The wild-type and Pol V+-only strains produced the greatest numbers of different RifR alleles across the greatest number of nucleotide positions, while the strain deficient in all three alternative polymerases generated the fewest alleles at the fewest positions. Strain-specific mutation spectra are summarized in Figure 5D, with source data presented in Table S3. Wild-type cells generated about half as many GC→AT mutations compared to any other strain, but had more than twice as many GC→CG mutations than all other strains except Pol V+-only. Pol II+-only generated more deletions than all other strains combined.
Alternative polymerase and other SOS genes are induced during stationary phase
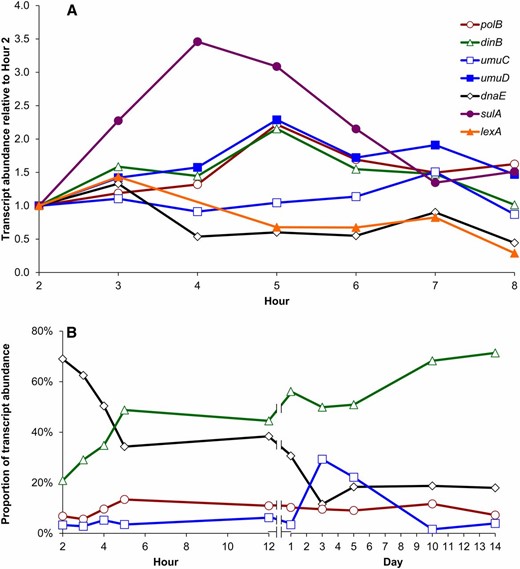
Given their observed physiological and evolutionary impacts during periods of feast and famine in the absence of endogenous stressors, we determined the expression patterns for each alternative DNA polymerase during long-term incubation. Wild-type cultures were sampled hourly for the first 12 hr and then every 24–48 hr over 14 days of long-term stationary-phase incubation. Quantitative RT-PCR on total cellular RNA throughout long-term incubation (Figure 6A) revealed an induction of transcripts encoding all three alternative polymerases (polB, dinB, umuDC), as well as the SOS response gene sulA. Relative to their abundance after 2 hr of exponential-phase growth, by 5 hr, as the cells transition into stationary phase, the average abundance of transcripts encoding Pol II and Pol IV more than doubled (dinB, +2.2-fold; polB, +2.2-fold) and the catalytic subunit of Pol V was induced 50% by hour 7 (umuC, +1.5-fold). Meanwhile, after a modest initial increase, transcript levels for the dnaE-encoded catalytic subunit of Pol III decreased by >40% within 4 hr and the SOS response repressor lexA was reduced >30% within 5 hr. For comparison, expression levels of the early exponential phase gene fis (Ball et al. 1992) peak at ∼2 hr and decrease significantly from mid-logarithmic phase through stationary phase while expression of the stationary-phase-specific dps gene (Martinez and Kolter 1997) increases from late log through stationary phase (data not shown).
Alternative polymerase transcript abundance changes over the cell cycle. Messenger RNA transcript abundance, in the absence of exogenous SOS inducers, was determined by qRT-PCR. (A) Transcript abundance for each gene relative to its concentration at 2 hr of incubation. Genes are identified as the following: polB, red, open circles; dinB, green, open triangles; umuC, blue, open squares; umuD, blue, solid squares; dnaE, black diamonds; sulA, purple, solid circles; and lexA, orange, solid triangles. (B) The proportion of polymerase transcripts over time, expressed as a percentage of total transcript abundance, is shown for four representative transcripts: Pol III, dnaE, black diamonds; Pol IV, dinB, green triangles; Pol II, polB, red circles; Pol V, umuC, blue squares.
When comparing the relative abundance of dnaE, polB, dinB, and umuC transcripts (Figure 6B), there is a dramatic decline in the average proportion of dnaE transcript from ∼69% during logarithmic growth to 34% upon entry into stationary phase. By 24 hr the dinB transcript is most abundant (56%), significantly exceeding that of dnaE (31%). For most time points, there was a consistent hierarchy of transcript abundance among alternative polymerases, with dinB > polB > umuC. The induction of alternative polymerases corresponded with induction of sulA, an SOS gene, indicating that stationary-phase conditions induce the SOS response in the absence of exogenous stressors. For comparison, induction of sulA during the SOS response after addition of the DNA-damaging agent mitomycin C was approximately fourfold higher than the induction observed as cells transition into stationary phase in the absence of drug (data not shown).
Discussion
The long-term viability and evolutionary success of bacterial populations requires a balance between maintaining the capacity to replicate efficiently with high fidelity while simultaneously generating sufficient genetic diversity to facilitate adaptation and survival in changing environments. Our previous study, which showed that alternative DNA polymerases play a role in the evolutionary fitness of bacterial populations (Yeiser et al. 2002), left open the key issue of each enzyme’s physiological and mutagenic contribution. Here we show specific roles for each alternative DNA polymerase. During periods of rapid cell division, Pol II contributes significantly to relative fitness by facilitating faster growth and the generation of genetic diversity, whereas Pol IV and Pol V introduce greater genetic variation, conferring increased relative fitness under the more stressful conditions of long-term stationary phase. Furthermore, differences in the frequency and spectrum of allelic variation attributable to each polymerase suggest a competitive hierarchy for access to replicate DNA in vivo.
Inherent differences in expression patterns and polymerase fidelity influence the relative fitness of the various mutant strains, allowing predictable outcomes of experiments where the strains are engaged in head-to-head competition. Given that success during long-term stationary phase is influenced by the appearance of advantageous mutations, type I and II competition experiments illustrate a race to generate beneficial alleles quickly (Figure 1). The observation that the double-mutant strains expressing only Pol IV or Pol V perform better than the Pol II+-only strain in both type 1 and type 2 competitions suggests that these polymerases play a greater role in creating mutations, including beneficial alleles, during long-term stationary phase. Consistent with the fact that Pol IV and Pol V have been shown to have inherently lower deoxynucleotide insertion fidelity and lack proofreading, we show that these enzymes produce genetic diversity more quickly than Pol II in slowly dividing, stationary-phase populations. Since previous work has shown that many different types of mutations can confer the GASP phenotype (Zambrano et al. 1993; Farrell and Finkel 2003; Finkel 2006), the appearance of novel mutations conferring GASP serves as a proxy for the extent of mutational change that has occurred in the population.
Following a 10-day aging regimen where beneficial alleles are generated and selected, the Pol II+-only strain expressed the strongest GASP phenotype (Figure 2). This finding suggests that, after providing ample time for beneficial mutations to appear, Pol II provides a greater physiological contribution to overall fitness via increased replicative efficiency during growth than either Pol IV or Pol V, enabling cells to capitalize on beneficial GASP alleles more quickly. This role for Pol II is bolstered by our observation that the Pol II+ strain performed significantly better than the Pol IV+-only and Pol V+-only strains when experiments were initiated at low density (allowing additional outgrowth) and within chemostat competitions allowing continuous growth. The fact that serially passaged Pol II+ populations exhibit greater competitive fitness against passaged Pol IV+-only or Pol V+-only strains leads to the conclusion that Pol II not only plays an important physiological role with respect to the efficiency of DNA replication, but also contributes considerable genetic diversity during periods of rapid growth.
Previous work identified differences in the frequency and spectrum of mutations generated by alternative polymerases; however, many of these studies analyzed either a limited subset of mutation classes, evaluated reporter genes on extrachromosomal elements, looked at mutations generated under artificial expression conditions, followed treatment with exogenous stressors, or used strains with altered repair systems (Hersh et al. 2004; Wolff et al. 2004; Foster 2005, 2007; Galhardo et al. 2007; Curti et al. 2009). Here we looked at the distribution of mutation frequencies under long-term laboratory growth conditions with endogenous expression in the absence of exogenous stressors in wild-type and isogenic mutant strains (Figure 5). The distribution of mutation frequencies reveals significant differences in the in vivo fidelity of each polymerase. Use of the double-mutant strains enables the analysis of each polymerase’s contribution to overall mutation frequency. The Pol V+-only strain was responsible for generating the greatest mutation frequency, followed by Pol IV+-only and Pol II+-only. The hierarchy of fidelity that we observe here is consistent with previously published in vitro and in vivo studies (Cai et al. 1995; Maor-Shoshani et al. 2000; Tang et al. 2000; Kobayashi et al. 2002).
It is also important to compare the mutation frequency of the single-mutant strains where two polymerases can compete for access to replicate DNA (Figure 5), enabling additional inferences regarding their relative contributions when acting in concert. Here the Pol IV− strain exhibited a greater mutation frequency than either Pol II− or Pol V−, which were approximately equal. Given that the relative abundance of each competing polymerase can influence which polymerase is most likely to gain access to the replication fork and that the Pol IV transcript was the most abundant under the conditions of our experiments, these findings suggest that the presence or absence of Pol IV is the greatest overall determinant of mutation frequency. Pol IV confers an intermediate mutation frequency among strains capable of expressing only one alternative polymerase. When Pol II is absent, Pol IV appears to mask the lower fidelity of Pol V; when Pol V is absent, Pol IV may outcompete the higher fidelity polymerase Pol II.
In the strain lacking only Pol IV, leaving Pol II and Pol V to compete, the mutation frequency was similar to the strain expressing only Pol V, suggesting that Pol V might outcompete Pol II to replicate during long-term batch culture. Together, these findings are consistent with a competitive hierarchy for access to replicate DNA in vivo of Pol IV > Pol V > Pol II when cells are incubating in long-term stationary phase. Therefore, even though Pol V is inherently more mutagenic, its impact on mutation frequency is modulated by Pol IV. Pol IV’s higher levels of expression enable it to outcompete the other polymerases and give it more opportunity to introduce genetic variation, consistent with its significant role in generating mutations in the lac system (Foster 2000; McKenzie et al. 2001). Surprisingly, the mutation frequency of the triple-mutant strain lacking all three SOS polymerases is similar to the wild-type strain. This possibly can be explained if mutated cells that cannot achieve translesion synthesis are lost from the population and the surviving mutants that we detect are attributable to the basal error rate of the replicative Pol III itself. This may also account for the more restricted spectrum of mutations observed in the triple-mutant strain.
Among >1000 spontaneous mutants resistant to rifampicin (see Table S1), we identified 85 alleles encompassing all six classes of missense mutations, as well as in-frame insertions and deletions. We found that the wild-type and Pol V+-only strains generated the greatest numbers of different alleles across the greatest number of nucleotide positions in rpoB. The triple-mutant strain, completely deficient in alternative polymerase expression, introduced the least variety of alleles at the fewest positions, demonstrating the importance of these enzymes in creating genetic diversity.
Previous reports have identified “fingerprints” or mutational hotspots left by specific polymerases in various genetic backgrounds (Wolff et al. 2004; Curti et al. 2009). We also identified strain-specific fluctuations in the frequency of specific alleles and mutation classes. For example, nearly half as many GC→AT mutations were identified in the wild-type strain compared to any other polymerase-deficient strain. Twice as many GC→CG mutations were observed in wild-type relative to all strains except Pol V+-only, suggesting that Pol V plays an important role in generating these transversions. The number of deletions detected in the Pol II+-only strain was greater than all other strains combined. Since this strain is incapable of expressing Pol IV or Pol V, it suggests that Pol II plays a significant role in generating deletions, as has been reported (Rangarajan et al. 1997; Koskiniemi and Andersson 2009; Wang and Yang 2009; Hastings et al. 2010). While the alterations in mutation spectrum observed differ from polymerase fingerprints reported elsewhere, these discrepancies may reflect strain and environmental differences.
The fact that the alternative polymerases, members of the SOS regulon, are expressed in the absence of endogenous stressors during long-term stationary phase further illustrates their impact on the survival and evolution of bacterial populations. When replacing Pol III, these polymerases each play a central role in providing a balance that achieves rapid growth while generating sufficient mutations to ensure genetic diversity. The observation that each alternative polymerase contributes differently to the variety and quantity of mutations demonstrates the importance of competitive interactions among the polymerases. One explanation for the apparent increase in alternative polymerase expression during long-term stationary phase may be a positive selection on bacterial populations that increase genetic diversity during times of stress. Indeed, the simultaneous downregulation of dnaE and induction of alternative polymerase expression observed here could produce a shift toward stress-induced mutagenesis by shifting polymerase competition in favor of error-prone replication as has been proposed elsewhere (Foster 2007; Curti et al. 2009; Hastings et al. 2010; Sutton 2010).
It has long been appreciated that the alternative DNA polymerases can be induced following stress and influence the formation of adaptive mutations, yet the extent of their contributions to long-term survival and evolution under conditions akin to those found in nature has remained primarily speculative. Our findings demonstrate specific physiological roles and evolutionary implications of each alternative DNA polymerase under conditions of both feast and famine. Given the ubiquitous nature of alternative DNA polymerases and their impact on cell fitness and survival, a deeper understanding of factors affecting their expression, competitive interactions, mutation preferences, and other cellular functions will yield valuable insights toward understanding the physiological responses and evolutionary trajectories of bacterial populations.
Acknowledgments
We thank Phuong Pham, Roger Woodgate, and members of the Finkel Lab for helpful discussions and Ken Nealson and Andrea Cheung for assistance with chemostat experiments. This work was supported in part by a grant from the U.S. Air Force Office of Scientific Research (FA-9550-06-1-0292), National Science Foundation CAREER Award MCB0237975 (to S.E.F.), and National Institutes of Health (NIH) grants GM21422 and ES012259 (to M.F.G.). C.H.C. received support from the NIH Cellular, Biochemical, and Molecular Sciences Training Program (T32GM67587).
Footnotes
Communicating editor: S. Sandler
Literature Cited
Friedberg, E. C., 2006 Mutagenesis and translesion synthesis in prokaryotes, pp. 407–522 in DNA Repair and Mutagenesis. ASM Press, Washington, DC.
Author notes
Supporting information is available online at http://www.genetics.org/lookup/suppl/doi:10.1534/genetics.113.151837/-/DC1.