-
PDF
- Split View
-
Views
-
Cite
Cite
Marcus E Marvin, Craig D Griffin, David E Eyre, David B H Barton, Edward J Louis, In Saccharomyces cerevisiae, yKu and Subtelomeric Core X Sequences Repress Homologous Recombination Near Telomeres as Part of the Same Pathway, Genetics, Volume 183, Issue 2, 1 October 2009, Pages 441–451, https://doi.org/10.1534/genetics.109.106674
Close - Share Icon Share
Abstract
Unlike in meiosis where recombination near telomeres is repressed, subtelomeric regions appear to recombine with each other frequently in vegetative cells with no detrimental consequences. To test whether or not such recombination is prevented in the core of chromosomes for maintenance of genome stability, we measured allelic homologous recombination (HR) along chromosome arms and between different ectopic locations. We found that there is an increase of recombination at telomeres in wild-type cells compared with sequences at proximal subtelomeric and interstitial regions of the genome. We also screened for mutations that result in an increase in HR between a telomeric sequence and a more internal sequence, which normally exhibit very low rates of HR. YKU80 was hit most frequently in our screen, and we show that the yKu heterodimer specifically represses HR in the vicinity of telomeres. This repression of HR is not explained solely by the role of yKu in maintaining telomere length, silencing, or tethering to the nuclear periphery. Analysis of mutant strains harboring deleted core X sequences revealed a role for this subtelomeric element in preventing telomeric recombination. Furthermore, core X bestowed this protection as part of the same pathway as yKu. Our findings implicate a role for both yKu and core X in stabilizing the genome against recombination events involving telomeric sequences.
THE chromosome ends of Saccharomyces cerevisiae consist of terminal TG-rich telomeric repeats flanked by blocks of homologous subtelomeric sequences known as core X and Y′ elements (http://www.nottingham.ac.uk/genetics/people/louis/images/ends-lowres.jpg). The core X sequences are present at all ends, suggesting that they play an important role in the nucleus, whereas Y′ elements are found at 17 of the 32 chromosome ends in either long or short form. The function of both these elements is unknown, but their presence or absence has been found to affect both subtelomeric silencing [telomere position effect (TPE)] and nuclear tethering capabilities (Fourel et al. 1999; Pryde and Louis 1999; Tham et al. 2001; Hediger et al. 2002; Taddei et al. 2004).
The analysis of genetic exchange in S. cerevisiae has shown that there is dynamic recombination between Y′ sequences, whereas the same is not true for the rest of the genome (Louis and Haber 1990; Louis et al. 1994). Sequence analysis of the chromosome ends suggests that the X element may present a barrier against genetic exchange, with elevated recombination between Y′ elements on the telomere-proximal side and limited recombination on the centromere-proximal side (Pryde and Louis 1997). There are also large blocks of homology proximal to the X element that indicate the exchange of sequences between ends, but these blocks are restricted to the subtelomeric regions and are shared by only a few ends (Flint et al. 1997). The formation of a telosome structure that provides protection from DNA degradation may also protect the ends from double-strand-breaks (DSBs) and/or prevent chromosome ends from being recognized as them. This structure may sequester the telomeres from the rest of the genome to avoid unwanted deleterious recombination events and end-to-end fusions. Therefore, telomeric proteins such as the yKu heterodimer may play an important role in stabilizing the genome against such activities (Downs and Jackson 2004).
The yKu protein of S. cerevisiae is encoded by two genes, YKU70 and YKU80, which make up a stable heterodimer that plays many important roles in the nucleus. In yeast, DSBs are mainly repaired by RAD52-dependent HR (reviewed in Krogh and Symington 2004). This is a yKu-independent process, but the protein is involved in additional important DNA-repair mechanisms such as nonhomologous end joining [NHEJ; reviewed in Daley et al. (2005)] and telomere maintenance, including the recruitment and activation of telomerase (reviewed in Downs and Jackson 2004). In addition, yKu is also involved in many other nuclear processes, such as retro-transposition, subtelomeric silencing, TPE, and anchoring to the nuclear envelope (Downs and Jackson 2004). The DNA repair and telomeric functions associated with yKu may work independently from each other, although yKu has been shown to aid the repair of bleomycin-damaged DNA, mainly through its telomere-related functions (Tam et al. 2007). Ku has also been shown to prevent deleterious chromosome end-to-end fusions by repressing NHEJ (Gravel et al. 1998). This paradox may be explained by a protective fold-back structure at the chromosome termini that changes the local architecture to prevent ends from interacting with each other (Pryde and Louis 1999; de Bruin et al. 2000, 2001). This conformation may be driven by telomeric Rap1p-bound Sir proteins (Sir2p–4p) interacting with more internally bound subtelomeric histones to provide a platform for the formation of transcriptional silent heterochromatin that results in TPE. The focal point of these interactions appears to be the subtelomeric core X region (that is found with ∼80% sequence homology at every chromosome end) as maximal silencing has been shown to occur there (Pryde and Louis 1999). Importantly, it has been demonstrated that yKu is required for establishing TPE (Boulton and Jackson 1998) and that the X element does not silence independently of the telomere (Pryde and Louis 1999).
Both mammalian Ku and yKu are known to repress HR at telomeres. TRF2-deficient mouse cells display a Ku-specific repression of telomeric sister-chromatid exchange (Celli et al. 2006), which is independent of NHEJ. While in yeast, yKu is known to repress homologous-recombination-dependent rapid deletion of telomeres (Polotnianka et al. 1998) and subtelomeric Y′ recombination (Fellerhoff et al. 2000) and is required for type I survival (Y′ amplification) in the absence of telomerase (Grandin and Charbonneau 2003).
Recently, several differentiation of function mutants of YKU70 and YKU80 have been characterized that shed light onto each subunit's contribution to the role of yKu in the cell (Bertuch and Lundblad 2003; Stellwagen et al. 2003; Palmbos et al. 2005; Ribes-Zamora et al. 2007). Two surface α-helices have been identified in the von Willebrand A domain of each corresponding protein subunit that each confer a specific role (Ribes-Zamora et al. 2007). The α-helix 5 of yKu70p is essential for NHEJ and, when bound to telomeric DNA, is proposed to form part of the surface of yKu that faces toward the terminal nucleotides. On the other hand, the α-helix 5 of yKu80p is responsible mainly for the telomeric maintenance functions of yKu and has been proposed to face inward toward the centromere (Ribes-Zamora et al. 2007). This is consistent with nearly all mutations characterized in YKU80 that affect telomeric functions and TPE, while those in YKU70 affect mainly NHEJ (Bertuch and Lundblad 2003; Stellwagen et al. 2003; Palmbos et al. 2005; Ribes-Zamora et al. 2007).
In this study, we have analyzed the properties of the yKu heterodimer with respect to preventing HR near telomeres. We initially isolated YKU80 in a screen designed to identify genes that maintain nuclear architecture by preventing recombination between telomeric and interstitial sequences. Further analysis showed that yKu helps to prevent mitotic recombination between telomeric sequences and plays a role in repressing recombination between telomeric and interstitial sequences to stabilize the genome.
MATERIALS AND METHODS
Yeast strains and growth:
Strain yAU15sir3Δ (MATα; hoΔpstI; ura3-N; leu2ΔX-R; trp1-B; spo13Δ; sir3Δ∷KanMX4; VIR∷URA3∷lys2-NdeI-1; lys2-NdeI-2), used to screen for mutations in genes that promote the subtelomeric-interstitial recombination barrier, is an isogenic derivative of Y55 (McCusker and Haber 1988). This strain is haploid, but both meiotic and mitotic recombination rates can be analyzed. This is achieved by the introduction of a sir3-null allele to abolish silencing of the HM loci and express both MATa and MATα to allow meiosis to occur (Rine and Herskowitz 1987). In addition, a spo13-null allele was introduced into yAU15sir3Δ to prevent the problem of dead spores through haploid meiosis, as these mutants do not undergo meiosis I (Rutkowski and Esposito 2000). To monitor ectopic recombination, yAU15sir3Δ contains lys2 heteroalleles (lys2-Nde1-1 situated at the telomere of chromosome VIR and lys2-Nde1-2 at the endogenous interstitial locus on chromosome II). The heteroalleles were produced by partial digestion of a cloned wild-type LYS2 gene using NdeI and filled in using Klenow polymerase to produce blunt ends. These were subsequently religated and introduced back into the strain. The first heteroallele (lys2-Nde1-2) is at its native location and was generated by “pop-in/pop-out” using a URA3-containing vector (Rothstein 1983) while the second telomeric allele was created using a telomeric repeat and a URA3-containing vector (Louis and Borts 1995). Strain yACt2 (MATa; hoΔpst; ura3-N; leu2ΔX-R; ade1; canR; VIR∷URA3∷lys2-NdeI-1; lys2-NdeI-2) was used for crosses with yAU15sir3Δ for genetic analysis. These crosses were possible because in Y55 strains a sir3 mutation does not completely derepress HMR to allow some residual mating. This is not seen in other strain backgrounds or in sir2 and sir4 mutations in this strain (data not shown). Isogenic strains to yAU15sir3Δ containing sir2- or sir4-null alleles instead of a sir3-null allele were also constructed (yAU15sir2Δ and yAU15sir4Δ). In addition, strains yAU15 and yAU36 were wild type for SIR, with the latter carrying a copy of MATα integrated at the TRP1 locus to express both mating types without affecting silencing.
Additional S. cerevisiae strains used for analyzing recombination rates were derivatives of YP1 (Louis and Haber 1989) and are shown in the supporting information, Table S1. Each strain carried lys2-801 and lys2-r2 heteroalleles (Louis and Haber 1989) at the endogenous interstitial site on chromosome IIR, as an internal allelic recombination control, and leu2 heteroalleles at the various sites shown in Figure 1A. The leu2-B heteroallele contains a 2-bp frameshift mutation introduced into the BstXI site of LEU2 by site-directed mutagenesis to produce a StuI site. The leu2-C heteroallele has a 2-bp insertion frameshift at the ClaI site of LEU2 that was produced by full digestion, filling in the 5′ overhang with Klenow fragment and subsequent blunt-end ligation. Gene deletions and insertions were introduced into these strains using PCR-generated disruption cassettes (Longtine et al. 1998) with the primers listed in Table S2 and the lithium acetate transformation method (Gietz et al. 1992). The heteroalleles were adjacent to a copy of the Kluyveromyces lactis URA3 gene. This was used as the selectable marker growing strains at 30° on the appropriate growth medium (Sherman et al. 1986) and checked using pulsed-field gel electrophoresis (Liti and Louis 2003) and Southern blot hybridization analysis to determine the location of genomic inserts. DNA was digested with NotI and hybridized using the K. lactis URA3 sequence as a probe and the Gene Images CDP-Star detection module from Amersham Pharmacia Biotech.
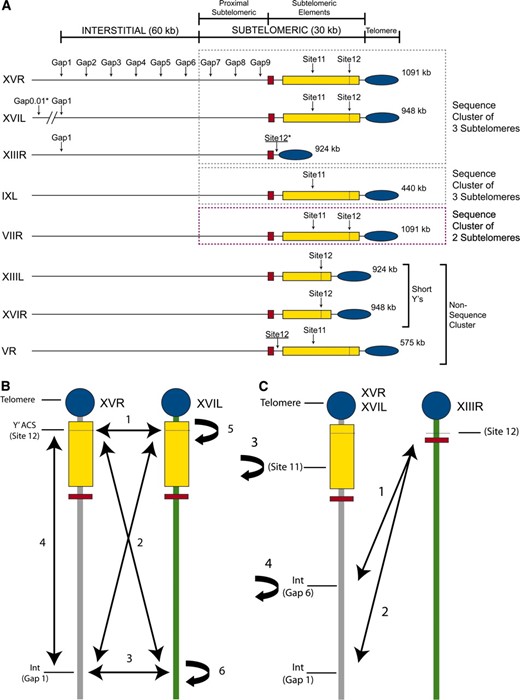
Genomic locations marked with KlURA3∷leu2C and KlURA3∷leu2B. (A) Cassettes are represented by arrows and were inserted into eight S. cerevisiae chromosome arms (see Table S1 for strains). Y′ elements (yellow boxes) are all long unless stated otherwise, and all possess a replication origin consensus sequence (Y′ ACS, gray dashed line) situated ∼250 bp from the telomeric TG1-3 repeats (blue ellipse). Physical map sizes are stated at the right end of each chromosome arm, and all are aligned by core X (red box). On chromosome XVR, gaps 1–9 are situated at subtelomeric intergenic regions spaced 10 kb apart, while gap 9, site 11, and site 12 are spaced at 5-kb intervals (not to scale). Three locations are in conserved subtelomeric elements: site 12 is located in the STRs; site 11 is in the middle of Y′ elements; site 12 is close to the Y′ ACS. Subdivisions according to sequence homology for each chromosome arm are shown on the right. An asterisk indicates a potentially unreliable location. (B) Arrows show the different combinations of ectopic and allelic recombination measured between heteroalleles placed at gap 1 and site 12 of chromosome XVR and XVIL: (1) ectopic recombination between site 12 (XVR) and site 12 (XVIL); (2) ectopic recombination between site 12 (XVR) and gap 1 (XVIL) or between site 12 (XVIL) and gap 1 (XVR); (3) ectopic recombination between gap 1 (XVR) and gap 1 (XVIL); (4) ectopic recombination between site 12 (XVR) and gap 1 (XVR) or between site 12 (XVIL) and gap 1 (XVIL); (5) allelic recombination between site 12 (XVR or XVIL); (6) allelic recombination between gap 1 sites (XVR or XVIL). (C) Arrows show the different combinations of ectopic and allelic recombination measured between heteroalleles placed at gap 1, gap 6, site 11, and site 12 of chromosomes XVR, XVIL, and XIIIR: (1) ectopic recombination between site 12 (XIIIR) and gap 6 (XVR); (2) ectopic recombination between site 12 (XIIIR) and gap 1 (XVIL); (3) allelic recombination between site 11 sites (XVR); (4) allelic recombination between gap 6 sites (XVR).
Genetic screen for hyper-recombination:
Strain yAU15sir3Δ was transformed with a mini-Tn3∷LEU2 transposon-tagged genomic library (Burns et al. 1994) and the resulting colonies were selected on leucine drop-out medium containing ethanol and glycerol. We originally found that 90% of the initial transformants were petite and unable to sporulate, hence the selection on a nonfermentable carbon source. Approximately 25,000 transformed colonies were isolated and transferred to fresh leucine drop-out medium in patches and allowed to grow. The patched colonies were then transferred sequentially onto YEPD and lysine drop-out medium to analyze mitotic recombination or onto YEPD, sporulation media, and lysine drop-out medium to analyze meiotic recombination. Patches containing cells that displayed an increased number of papillae on lysine drop-out medium were rechecked and chosen for further study. Each colony was then subjected to CHEF gel electrophoresis to check for gross chromosomal rearrangements. Telomere length was also analyzed by Southern hybridization analysis using a short TG1-3 probe (∼200 bp) that had been PCR amplified using plasmid pKK38b as a template (Kramer and Haber 1993). Colonies from the screen were then crossed with yACT2, and the resulting diploid was sporulated and tetrad analysis was performed. The Y55 background exhibits some mating in a MATα haploid carrying a sir3-null allele (data not shown) allowing segregation analysis. The resulting spores were checked to confirm that the hyper-recombination phenotype cosegregated with leucine prototrophy. Most of the transposon-tagged locations were identified by vectorette PCR using the Botstein laboratory protocol (http://www.genomics.princeton.edu/botstein/Protocols/vectorette.html) and by WU-BLAST2 searching each sequence obtained on the Saccharomyces Genome Database (SGD; http://www.yeastgenome.org/).
Role of yKu in mitotic homologous recombination:
A subset of the strains marked with leu2 heteroalleles (Table S1) had null alleles of yku70 and yku80 introduced by PCR-targeted mutagenesis (Longtine et al. 1998) and were used in various combinations to test the effects of yKu on interstitial, telomeric, and non-Y′ telomeric mitotic recombination.
Statistical analysis of mitotic recombination and gene ontology annotations:
Recombination assays were performed using a CAS 1200 liquid-handling robot (Corbett Life Science). Recombination data were analyzed with t-tests using 0.05 as the α-value. Mitotic recombination was determined by obtaining a median recombinant count for at least eight independent diploid colonies. Each was picked from YEPD plates and suspended in 1 ml of sterile distilled water, and either 4- or 10-fold serial dilutions were made. These were then transferred in 40-μl aliquots onto complete synthetic medium and synthetic medium without leucine or lysine. After 48 hr growth, the number of papillae formed in each patch of cells was counted. Only dilutions with patches that contained >1 or <50 papillae were routinely used for data. The mitotic recombination rate was calculated using the method of the median first described by Lea and Coulsen (1949) as used to analyze mutation rates in msh1 and msh2 mutants (Reenan and Kolodner 1992). Efficiencies of ectopic recombination were estimated using the method described by Goldman and Lichten (1996). The chi-square distribution was used to determine whether or not recombination rates were significantly different between given strains, using a nonparametric rank-order test. A description of the statistical analysis of gene ontology (GO) annotations associated with genes isolated from the genetic hyper-recombination can be found in File S2.
RESULTS
Mitotic allelic recombination is elevated in subtelomeric regions:
Mitotic recombination rates in S. cerevisiae between heteroalleles on homologous chromosomes has been studied mainly on single loci in interstitial regions and has been found to be very low (reviewed in Paques and Haber 1999). Allelic recombination was not altered significantly when leu2 heteroalleles were placed at proximal subtelomeric and interstitial loci (Lichten and Haber 1989) and between direct repeats placed at subtelomeric and interstitial loci (Stavenhagen and Zakian 1998). We have proposed a model where a recombination barrier is set up between subtelomeric and interstitial sequences via nuclear tethering based on sequence analysis and preliminary recombination studies (Pryde and Louis 1997). To investigate this potential barrier, we assayed allelic recombination between leu2 heteroalleles inserted at 21 genomic positions (see materials and methods). These represented both interstitial and subtelomeric locations from different groups of chromosomes, specifically X-only ends, short Y′ and long Y′ ends, and those that can be subdivided by sequence homology (Figure 1A and Table S1).
Mitotic recombination between the leu2 heteroalleles was assayed, and allelic recombination rates were measured using the method of the median (Table 1). For an internal control, rates were also observed between lys2 heteroalleles positioned at the endogenous LYS2 genomic locus. For a global view, we combined the data for heteroalleles that had been inserted at similar locations (not shown). This revealed a similar allelic recombination rate across most of the genome (1.3–1.6 × 10−7 events/mitosis), including interstitial and proximal subtelomeric regions. When heteroalleles were placed in Y′ elements, they exhibited elevated rates of allelic recombination. Five of these inserts were at the Y′ ARS consensus site (ACS, site 12), which is ∼250 bp away from the telomeric TG1-3 repeats, and the sixth was in the middle of the Y′ element (site 11) on chromosome IXL. Interestingly, inserts in the subtelomeric repeats (STRs) of chromosome XIIIR (site 12) gave an allelic recombination rate that was similar to the general allelic recombination rate for the genome, even though of all the inserts, these were closest to the TG1-3 repeats. This may suggest that the STRs possess some kind of anti-recombination properties and that it is not simply the close proximity of repeat sequences such as Y′ elements to telomeres that results in elevated recombination rates.
Fluctuation analysis was used to measure mitotic allelic recombination rates per 108 cells
. | leu2 mitotic recombination rate . | lys2 mitotic recombination rate . | . | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| . | Individual assay . | . | . | Individual assay . | . | . | . | ||||
| Locations . | 1 . | 2 . | 3 . | Mean . | SD . | 1 . | 2 . | 3 . | Mean . | SD . | Cluster . |
| XVIL gap 1 | 13.3 | 11.9 | 13.1 | 12.7 | 0.8 | 55.8 | 57.8 | 52.6 | 55.4 | 2.6 | NA |
| XVR gap 1 | 5.9 | 8.6 | 11.5 | 8.6 | 2.8 | 69.4 | 61.0 | 52.1 | 60.8 | 8.6 | NA |
| XVR gap 3 | 17.4 | 64.1 | NA | ||||||||
| XVR gap 5 | 16.7 | 65.1 | NA | ||||||||
| XVR gap 6 | 17.7 | 68.9 | NA | ||||||||
| XIIIR gap 1 | 15.4 | 56.2 | NA | ||||||||
| XVR gap 7 | 12.1 | 56.4 | + | ||||||||
| XVR gap 8 | 20.5 | 70.8 | |||||||||
| XVR gap 9 | 14.9 | 64.7 | + | ||||||||
| XIIIR site 12 | 7.3 | 56.0 | + | ||||||||
| VR site 12 | 23.1 | 79.4 | – | ||||||||
| XVIL site 11 | 8.2 | 50.8 | + | ||||||||
| XVR site 11 | 17.7 | 70.0 | + | ||||||||
| IXL site 11 | 72.9 | 87.0 | 84.0 | 81.3 | 7.4 | 69.5 | 88.6 | 87.2 | 81.8 | 10.6 | + |
| VIIR site 11 | 22.3 | 72.3 | + | ||||||||
| VR site 11 | 22.7 | 75.9 | − | ||||||||
| XVIR site 12 | 62.8 | 71.4 | − | ||||||||
| XVIL site 12 | 327.7 | 212.4 | 238.4 | 259.5 | 60.5 | 72.0 | 48.5 | 55.6 | 58.7 | 12.0 | + |
| XVR site 12 | 70.9 | 76.6 | 37.7 | 61.7 | 21.0 | 76.2 | 60.1 | 63.8 | 66.7 | 8.4 | + |
| XIIIL site 12 | 61.4 | 66.4 | − | ||||||||
| VIIR site 12 | 75.8 | 71.6 | + | ||||||||
. | leu2 mitotic recombination rate . | lys2 mitotic recombination rate . | . | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| . | Individual assay . | . | . | Individual assay . | . | . | . | ||||
| Locations . | 1 . | 2 . | 3 . | Mean . | SD . | 1 . | 2 . | 3 . | Mean . | SD . | Cluster . |
| XVIL gap 1 | 13.3 | 11.9 | 13.1 | 12.7 | 0.8 | 55.8 | 57.8 | 52.6 | 55.4 | 2.6 | NA |
| XVR gap 1 | 5.9 | 8.6 | 11.5 | 8.6 | 2.8 | 69.4 | 61.0 | 52.1 | 60.8 | 8.6 | NA |
| XVR gap 3 | 17.4 | 64.1 | NA | ||||||||
| XVR gap 5 | 16.7 | 65.1 | NA | ||||||||
| XVR gap 6 | 17.7 | 68.9 | NA | ||||||||
| XIIIR gap 1 | 15.4 | 56.2 | NA | ||||||||
| XVR gap 7 | 12.1 | 56.4 | + | ||||||||
| XVR gap 8 | 20.5 | 70.8 | |||||||||
| XVR gap 9 | 14.9 | 64.7 | + | ||||||||
| XIIIR site 12 | 7.3 | 56.0 | + | ||||||||
| VR site 12 | 23.1 | 79.4 | – | ||||||||
| XVIL site 11 | 8.2 | 50.8 | + | ||||||||
| XVR site 11 | 17.7 | 70.0 | + | ||||||||
| IXL site 11 | 72.9 | 87.0 | 84.0 | 81.3 | 7.4 | 69.5 | 88.6 | 87.2 | 81.8 | 10.6 | + |
| VIIR site 11 | 22.3 | 72.3 | + | ||||||||
| VR site 11 | 22.7 | 75.9 | − | ||||||||
| XVIR site 12 | 62.8 | 71.4 | − | ||||||||
| XVIL site 12 | 327.7 | 212.4 | 238.4 | 259.5 | 60.5 | 72.0 | 48.5 | 55.6 | 58.7 | 12.0 | + |
| XVR site 12 | 70.9 | 76.6 | 37.7 | 61.7 | 21.0 | 76.2 | 60.1 | 63.8 | 66.7 | 8.4 | + |
| XIIIL site 12 | 61.4 | 66.4 | − | ||||||||
| VIIR site 12 | 75.8 | 71.6 | + | ||||||||
Both experimental (leu2) and control (lys2) heteroallele data are shown. The location of the leu2 heteroalleles in each assay is shown on the left (see Figure 1A). All lys2 heteroalleles were located at the endogenous LYS2 locus on the right arm of chromosome II, 353 kb from the telomere. Individual rates were obtained from independent assays and, when more than one assay was carried out, the mean and standard deviation (SD) are shown. Heteroalleles of leu2 that are in subtelomeres that can be aligned into sequence clusters are indicated by a plus sign (+) and those that are not by a minus sign (−).
Fluctuation analysis was used to measure mitotic allelic recombination rates per 108 cells
. | leu2 mitotic recombination rate . | lys2 mitotic recombination rate . | . | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| . | Individual assay . | . | . | Individual assay . | . | . | . | ||||
| Locations . | 1 . | 2 . | 3 . | Mean . | SD . | 1 . | 2 . | 3 . | Mean . | SD . | Cluster . |
| XVIL gap 1 | 13.3 | 11.9 | 13.1 | 12.7 | 0.8 | 55.8 | 57.8 | 52.6 | 55.4 | 2.6 | NA |
| XVR gap 1 | 5.9 | 8.6 | 11.5 | 8.6 | 2.8 | 69.4 | 61.0 | 52.1 | 60.8 | 8.6 | NA |
| XVR gap 3 | 17.4 | 64.1 | NA | ||||||||
| XVR gap 5 | 16.7 | 65.1 | NA | ||||||||
| XVR gap 6 | 17.7 | 68.9 | NA | ||||||||
| XIIIR gap 1 | 15.4 | 56.2 | NA | ||||||||
| XVR gap 7 | 12.1 | 56.4 | + | ||||||||
| XVR gap 8 | 20.5 | 70.8 | |||||||||
| XVR gap 9 | 14.9 | 64.7 | + | ||||||||
| XIIIR site 12 | 7.3 | 56.0 | + | ||||||||
| VR site 12 | 23.1 | 79.4 | – | ||||||||
| XVIL site 11 | 8.2 | 50.8 | + | ||||||||
| XVR site 11 | 17.7 | 70.0 | + | ||||||||
| IXL site 11 | 72.9 | 87.0 | 84.0 | 81.3 | 7.4 | 69.5 | 88.6 | 87.2 | 81.8 | 10.6 | + |
| VIIR site 11 | 22.3 | 72.3 | + | ||||||||
| VR site 11 | 22.7 | 75.9 | − | ||||||||
| XVIR site 12 | 62.8 | 71.4 | − | ||||||||
| XVIL site 12 | 327.7 | 212.4 | 238.4 | 259.5 | 60.5 | 72.0 | 48.5 | 55.6 | 58.7 | 12.0 | + |
| XVR site 12 | 70.9 | 76.6 | 37.7 | 61.7 | 21.0 | 76.2 | 60.1 | 63.8 | 66.7 | 8.4 | + |
| XIIIL site 12 | 61.4 | 66.4 | − | ||||||||
| VIIR site 12 | 75.8 | 71.6 | + | ||||||||
. | leu2 mitotic recombination rate . | lys2 mitotic recombination rate . | . | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| . | Individual assay . | . | . | Individual assay . | . | . | . | ||||
| Locations . | 1 . | 2 . | 3 . | Mean . | SD . | 1 . | 2 . | 3 . | Mean . | SD . | Cluster . |
| XVIL gap 1 | 13.3 | 11.9 | 13.1 | 12.7 | 0.8 | 55.8 | 57.8 | 52.6 | 55.4 | 2.6 | NA |
| XVR gap 1 | 5.9 | 8.6 | 11.5 | 8.6 | 2.8 | 69.4 | 61.0 | 52.1 | 60.8 | 8.6 | NA |
| XVR gap 3 | 17.4 | 64.1 | NA | ||||||||
| XVR gap 5 | 16.7 | 65.1 | NA | ||||||||
| XVR gap 6 | 17.7 | 68.9 | NA | ||||||||
| XIIIR gap 1 | 15.4 | 56.2 | NA | ||||||||
| XVR gap 7 | 12.1 | 56.4 | + | ||||||||
| XVR gap 8 | 20.5 | 70.8 | |||||||||
| XVR gap 9 | 14.9 | 64.7 | + | ||||||||
| XIIIR site 12 | 7.3 | 56.0 | + | ||||||||
| VR site 12 | 23.1 | 79.4 | – | ||||||||
| XVIL site 11 | 8.2 | 50.8 | + | ||||||||
| XVR site 11 | 17.7 | 70.0 | + | ||||||||
| IXL site 11 | 72.9 | 87.0 | 84.0 | 81.3 | 7.4 | 69.5 | 88.6 | 87.2 | 81.8 | 10.6 | + |
| VIIR site 11 | 22.3 | 72.3 | + | ||||||||
| VR site 11 | 22.7 | 75.9 | − | ||||||||
| XVIR site 12 | 62.8 | 71.4 | − | ||||||||
| XVIL site 12 | 327.7 | 212.4 | 238.4 | 259.5 | 60.5 | 72.0 | 48.5 | 55.6 | 58.7 | 12.0 | + |
| XVR site 12 | 70.9 | 76.6 | 37.7 | 61.7 | 21.0 | 76.2 | 60.1 | 63.8 | 66.7 | 8.4 | + |
| XIIIL site 12 | 61.4 | 66.4 | − | ||||||||
| VIIR site 12 | 75.8 | 71.6 | + | ||||||||
Both experimental (leu2) and control (lys2) heteroallele data are shown. The location of the leu2 heteroalleles in each assay is shown on the left (see Figure 1A). All lys2 heteroalleles were located at the endogenous LYS2 locus on the right arm of chromosome II, 353 kb from the telomere. Individual rates were obtained from independent assays and, when more than one assay was carried out, the mean and standard deviation (SD) are shown. Heteroalleles of leu2 that are in subtelomeres that can be aligned into sequence clusters are indicated by a plus sign (+) and those that are not by a minus sign (−).
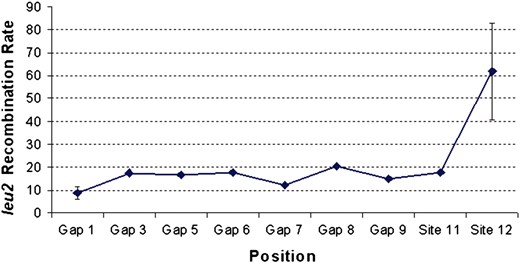
Although little variation in recombination rate was observed between interstitial, proximal subtelomeric, and STR sequences, Y′ and Y′ ACS sequences showed variation in rate. Of the 21 allelic recombination rates tested, 9 were for sites along chromosome XVR (Figure 2). Most of the recombination rates along this arm were very similar, with only the last 5 kb showing a significant increase, particularly at site 12 over the Y′ ACS. Mitotic recombination rates between the control lys2 heteroalleles were very similar in all 21 strains tested (Table 2), showing that the global recombination rate in each strain had not been altered by our leu2 heteroallele insertions. These data show that there is a fundamental difference between allelic recombination rates at sequences in the subtelomeric elements compared with sequences at proximal subtelomeric and interstitial regions of the genome.
Wild-type allelic mitotic recombination rates along chromosome XVR. Fluctuation analysis was used to analyze recombination rates at nine locations (Figure 1A). The data represent experimental (leu2) heteroallele recombination rates per 108 cells. Gap 1 and site 12 rates were calculated from three independent assays, and the mean and standard deviation (error bars) are shown. All other rates are from a single assay. Recombination rates for locations on the additional chromosome arms and for lys2 heteroalleles can be seen in Table 1.
Mitotic ectopic recombination rates between different locations in yku70Δ/yku70Δ strains
Loci marked . | Mitotic leu2 recombination rate . | Fold increase (yku70Δ/YKU70) . | . | Mitotic lys2 control recombination rate . | . | . | ||||
|---|---|---|---|---|---|---|---|---|---|---|
| leu2-C . | leu2-B . | YKU70a . | yku70Δb . | Individual . | Mean . | Two-tailed P test . | YKU70a . | yku70Δb . | Context . | Location . |
| XVR site 12 | XVIL site 12 | 54.1 | 551.2 | 10 | 9 | 0.056 | 74.5 | 85.2 | Y′ ACS–Y′ ACS | XVR–XVIL |
| XVIL site 12 | XVR site 12 | 162.0 | 1158.7 | 7 | — | 0.018 | 84.5 | 83.2 | ||
| XVR gap 1 | XVIL site 12 | 9.7 | 31.8 | 3 | 3 | 0.109 | 92.1 | 83.5 | Int–Y′ ACS | |
| XVIL site 12 | XVR gap 1 | 8.6 | 22.4 | 3 | — | 0.003 | 75.1 | 81.8 | ||
| XVIL gap 1 | XVR site 12 | 14.4 | 89.5 | 6 | 5 | 0.047 | 79.6 | 83.7 | Y′ ACS–Int | |
| XVR site 12 | XVIL gap 1 | 10.1 | 45.4 | 5 | — | 0.109 | 73.6 | 88.9 | ||
| XVR gap 1 | XVIL gap 1 | 5.4 | 5.7 | 1 | 1 | 0.668 | 77.8 | 73.7 | Int–Int | |
| XVIL gap 1 | XVR gap 1 | 3.8 | 3.3 | 1 | — | 0.773 | 79.1 | 78.6 | ||
| XVR gap 1 | XVR site 12 | 12.9 | 46.1 | 4 | 3 | 0.045 | 67.4 | 70.7 | Int-Y′ ACS | XVR–XVR |
| XVR site 12 | XVR gap 1 | 4.6 | 14.3 | 3 | — | 0.025 | 105.9 | 86.0 | ||
| XVIL gap 1 | XVIL site 12 | 12.8 | 45.4 | 4 | 3 | 0.045 | 156.3 | 107.4 | XVIL–XVIL | |
| XVIL site 12 | XVIL gap 1 | 20.7 | 65.9 | 3 | — | 0.002 | 62.6 | 76.3 | ||
| XVR site 12 | XVR site 12 | 61.7 | 970.5 | 16 | 10 | 0.027 | 67.7 | 68.2 | Allelic | |
| XVIL site 12 | XVIL site 12 | 259.5 | 975.0 | 4 | — | 0.105 | 58.7 | 64.6 | ||
| XVR gap 1 | XVR gap 1 | 8.6 | 10.4 | 1 | 1 | 0.435 | 60.8 | 78.0 | ||
| XVIL gap 1 | XVIL gap 1 | 12.7 | 14.5 | 1 | — | 0.472 | 55.4 | 77.3 | ||
Loci marked . | Mitotic leu2 recombination rate . | Fold increase (yku70Δ/YKU70) . | . | Mitotic lys2 control recombination rate . | . | . | ||||
|---|---|---|---|---|---|---|---|---|---|---|
| leu2-C . | leu2-B . | YKU70a . | yku70Δb . | Individual . | Mean . | Two-tailed P test . | YKU70a . | yku70Δb . | Context . | Location . |
| XVR site 12 | XVIL site 12 | 54.1 | 551.2 | 10 | 9 | 0.056 | 74.5 | 85.2 | Y′ ACS–Y′ ACS | XVR–XVIL |
| XVIL site 12 | XVR site 12 | 162.0 | 1158.7 | 7 | — | 0.018 | 84.5 | 83.2 | ||
| XVR gap 1 | XVIL site 12 | 9.7 | 31.8 | 3 | 3 | 0.109 | 92.1 | 83.5 | Int–Y′ ACS | |
| XVIL site 12 | XVR gap 1 | 8.6 | 22.4 | 3 | — | 0.003 | 75.1 | 81.8 | ||
| XVIL gap 1 | XVR site 12 | 14.4 | 89.5 | 6 | 5 | 0.047 | 79.6 | 83.7 | Y′ ACS–Int | |
| XVR site 12 | XVIL gap 1 | 10.1 | 45.4 | 5 | — | 0.109 | 73.6 | 88.9 | ||
| XVR gap 1 | XVIL gap 1 | 5.4 | 5.7 | 1 | 1 | 0.668 | 77.8 | 73.7 | Int–Int | |
| XVIL gap 1 | XVR gap 1 | 3.8 | 3.3 | 1 | — | 0.773 | 79.1 | 78.6 | ||
| XVR gap 1 | XVR site 12 | 12.9 | 46.1 | 4 | 3 | 0.045 | 67.4 | 70.7 | Int-Y′ ACS | XVR–XVR |
| XVR site 12 | XVR gap 1 | 4.6 | 14.3 | 3 | — | 0.025 | 105.9 | 86.0 | ||
| XVIL gap 1 | XVIL site 12 | 12.8 | 45.4 | 4 | 3 | 0.045 | 156.3 | 107.4 | XVIL–XVIL | |
| XVIL site 12 | XVIL gap 1 | 20.7 | 65.9 | 3 | — | 0.002 | 62.6 | 76.3 | ||
| XVR site 12 | XVR site 12 | 61.7 | 970.5 | 16 | 10 | 0.027 | 67.7 | 68.2 | Allelic | |
| XVIL site 12 | XVIL site 12 | 259.5 | 975.0 | 4 | — | 0.105 | 58.7 | 64.6 | ||
| XVR gap 1 | XVR gap 1 | 8.6 | 10.4 | 1 | 1 | 0.435 | 60.8 | 78.0 | ||
| XVIL gap 1 | XVIL gap 1 | 12.7 | 14.5 | 1 | — | 0.472 | 55.4 | 77.3 | ||
The recombination rates given are per 108 cells, and all data shown are derived from mean calculations from at least three independent assays (a minimum of 24 colonies). Recombination was assayed between all combinations of inserts located at gap 1 and site 12 on chromosomes XVR and XVIL (see Figure 1A). Gap 1 is at an interstitial (Int) location ∼90 kb from respective telomeres, and site 12 is close to the replication origin consensus sequence located in Y′ elements (Y′ ACS). Both mutant and wild-type data have been analyzed in two ways: fold increase in the mutant (yku70Δ/YKU70) and two-tailed t-tests (column 7) to assign P values (with no adjustment for multiple comparisons). Results for control allelic lys2 alleles are also shown.
Fluctuation analysis was used to determine mitotic recombination rates between leu2 heteroalleles in parental isogenic wild-type strains.
Fluctuation analysis was used to determine mitotic recombination rates between leu2 heteroalleles in parental isogenic yku70Δ/yku70Δ2 strains.
Mitotic ectopic recombination rates between different locations in yku70Δ/yku70Δ strains
Loci marked . | Mitotic leu2 recombination rate . | Fold increase (yku70Δ/YKU70) . | . | Mitotic lys2 control recombination rate . | . | . | ||||
|---|---|---|---|---|---|---|---|---|---|---|
| leu2-C . | leu2-B . | YKU70a . | yku70Δb . | Individual . | Mean . | Two-tailed P test . | YKU70a . | yku70Δb . | Context . | Location . |
| XVR site 12 | XVIL site 12 | 54.1 | 551.2 | 10 | 9 | 0.056 | 74.5 | 85.2 | Y′ ACS–Y′ ACS | XVR–XVIL |
| XVIL site 12 | XVR site 12 | 162.0 | 1158.7 | 7 | — | 0.018 | 84.5 | 83.2 | ||
| XVR gap 1 | XVIL site 12 | 9.7 | 31.8 | 3 | 3 | 0.109 | 92.1 | 83.5 | Int–Y′ ACS | |
| XVIL site 12 | XVR gap 1 | 8.6 | 22.4 | 3 | — | 0.003 | 75.1 | 81.8 | ||
| XVIL gap 1 | XVR site 12 | 14.4 | 89.5 | 6 | 5 | 0.047 | 79.6 | 83.7 | Y′ ACS–Int | |
| XVR site 12 | XVIL gap 1 | 10.1 | 45.4 | 5 | — | 0.109 | 73.6 | 88.9 | ||
| XVR gap 1 | XVIL gap 1 | 5.4 | 5.7 | 1 | 1 | 0.668 | 77.8 | 73.7 | Int–Int | |
| XVIL gap 1 | XVR gap 1 | 3.8 | 3.3 | 1 | — | 0.773 | 79.1 | 78.6 | ||
| XVR gap 1 | XVR site 12 | 12.9 | 46.1 | 4 | 3 | 0.045 | 67.4 | 70.7 | Int-Y′ ACS | XVR–XVR |
| XVR site 12 | XVR gap 1 | 4.6 | 14.3 | 3 | — | 0.025 | 105.9 | 86.0 | ||
| XVIL gap 1 | XVIL site 12 | 12.8 | 45.4 | 4 | 3 | 0.045 | 156.3 | 107.4 | XVIL–XVIL | |
| XVIL site 12 | XVIL gap 1 | 20.7 | 65.9 | 3 | — | 0.002 | 62.6 | 76.3 | ||
| XVR site 12 | XVR site 12 | 61.7 | 970.5 | 16 | 10 | 0.027 | 67.7 | 68.2 | Allelic | |
| XVIL site 12 | XVIL site 12 | 259.5 | 975.0 | 4 | — | 0.105 | 58.7 | 64.6 | ||
| XVR gap 1 | XVR gap 1 | 8.6 | 10.4 | 1 | 1 | 0.435 | 60.8 | 78.0 | ||
| XVIL gap 1 | XVIL gap 1 | 12.7 | 14.5 | 1 | — | 0.472 | 55.4 | 77.3 | ||
Loci marked . | Mitotic leu2 recombination rate . | Fold increase (yku70Δ/YKU70) . | . | Mitotic lys2 control recombination rate . | . | . | ||||
|---|---|---|---|---|---|---|---|---|---|---|
| leu2-C . | leu2-B . | YKU70a . | yku70Δb . | Individual . | Mean . | Two-tailed P test . | YKU70a . | yku70Δb . | Context . | Location . |
| XVR site 12 | XVIL site 12 | 54.1 | 551.2 | 10 | 9 | 0.056 | 74.5 | 85.2 | Y′ ACS–Y′ ACS | XVR–XVIL |
| XVIL site 12 | XVR site 12 | 162.0 | 1158.7 | 7 | — | 0.018 | 84.5 | 83.2 | ||
| XVR gap 1 | XVIL site 12 | 9.7 | 31.8 | 3 | 3 | 0.109 | 92.1 | 83.5 | Int–Y′ ACS | |
| XVIL site 12 | XVR gap 1 | 8.6 | 22.4 | 3 | — | 0.003 | 75.1 | 81.8 | ||
| XVIL gap 1 | XVR site 12 | 14.4 | 89.5 | 6 | 5 | 0.047 | 79.6 | 83.7 | Y′ ACS–Int | |
| XVR site 12 | XVIL gap 1 | 10.1 | 45.4 | 5 | — | 0.109 | 73.6 | 88.9 | ||
| XVR gap 1 | XVIL gap 1 | 5.4 | 5.7 | 1 | 1 | 0.668 | 77.8 | 73.7 | Int–Int | |
| XVIL gap 1 | XVR gap 1 | 3.8 | 3.3 | 1 | — | 0.773 | 79.1 | 78.6 | ||
| XVR gap 1 | XVR site 12 | 12.9 | 46.1 | 4 | 3 | 0.045 | 67.4 | 70.7 | Int-Y′ ACS | XVR–XVR |
| XVR site 12 | XVR gap 1 | 4.6 | 14.3 | 3 | — | 0.025 | 105.9 | 86.0 | ||
| XVIL gap 1 | XVIL site 12 | 12.8 | 45.4 | 4 | 3 | 0.045 | 156.3 | 107.4 | XVIL–XVIL | |
| XVIL site 12 | XVIL gap 1 | 20.7 | 65.9 | 3 | — | 0.002 | 62.6 | 76.3 | ||
| XVR site 12 | XVR site 12 | 61.7 | 970.5 | 16 | 10 | 0.027 | 67.7 | 68.2 | Allelic | |
| XVIL site 12 | XVIL site 12 | 259.5 | 975.0 | 4 | — | 0.105 | 58.7 | 64.6 | ||
| XVR gap 1 | XVR gap 1 | 8.6 | 10.4 | 1 | 1 | 0.435 | 60.8 | 78.0 | ||
| XVIL gap 1 | XVIL gap 1 | 12.7 | 14.5 | 1 | — | 0.472 | 55.4 | 77.3 | ||
The recombination rates given are per 108 cells, and all data shown are derived from mean calculations from at least three independent assays (a minimum of 24 colonies). Recombination was assayed between all combinations of inserts located at gap 1 and site 12 on chromosomes XVR and XVIL (see Figure 1A). Gap 1 is at an interstitial (Int) location ∼90 kb from respective telomeres, and site 12 is close to the replication origin consensus sequence located in Y′ elements (Y′ ACS). Both mutant and wild-type data have been analyzed in two ways: fold increase in the mutant (yku70Δ/YKU70) and two-tailed t-tests (column 7) to assign P values (with no adjustment for multiple comparisons). Results for control allelic lys2 alleles are also shown.
Fluctuation analysis was used to determine mitotic recombination rates between leu2 heteroalleles in parental isogenic wild-type strains.
Fluctuation analysis was used to determine mitotic recombination rates between leu2 heteroalleles in parental isogenic yku70Δ/yku70Δ2 strains.
Mitotic ectopic recombination is elevated when one partner is near a telomere:
To ascertain whether or not a recombination barrier exists between subtelomeric and interstitial sequences in S. cerevisiae, we assayed ectopic recombination rates between leu2 heteroalleles placed in the subtelomeric and interstitial regions of chromosome XVR and XVIL (Figure 1A). First we tested whether or not the distance between partners, relative to their respective telomeres, affected the rate of recombination. To achieve this, we measured recombination rates in diploids by crossing haploid strains that harbored heteroalleles that were sequentially moving away from a fixed point relative to their telomeres. We first utilized diploid strains with one heteroallele fixed at the most internal point, gap 1 (Figure 1A) and the second heteroallele sequentially moving away toward the telomere of the other chromosome (gap 1, gap 2, gap 3, and so on through to site 12). We then performed the same set of experiments using the most telomeric point, site 12, as the fixed point, moving the second heteroallele sequentially toward the centromere. Our measurements of recombination rates from these experiments (data not shown) showed that the further one heteroallele was moved away from its partner, the less likely it was able to recombine.
Recombination rates were then measured in all four combinations of heteroalleles located at gap 1 and site 12, which represented three different types of recombination (Figure 1B): between interstitial sequences (Int–Int); between subtelomeric sequences (Y′ ACS × Y′ ACS); and between interstitial and subtelomeric sequences (Int × Y′ ACS). When both heteroalleles were located at site 12, the recombination rate was always higher than when one heteroallele was placed at gap 1. In addition, when both heteroalleles were placed at gap 1, the rate was always lower than when one was placed at site 12 (Table 2). The control allelic lys2 recombination rates were similar in all the strains tested. The initial concept of a barrier (Pryde and Louis 1997) requires modification to incorporate a gradient of decrease in HR as one or both partners increase the distance from their telomeres. Instead, we observed an elevated recombination rate close to the Y′ ACS (site 12) when one or two of the heteroalleles were placed there. By comparing the individual recombination rates using a t-test, we identified significant differences (P < 0.05) between each of them.
YKU80 represses telomeric by interstitial HR:
To isolate genes that maintain a genetic barrier to recombination between subtelomeric and interstitial sequences, we performed the genetic screen described in materials and methods. A total of 102 genes or open reading frames were identified by the transposon resulting in mitotic hyper-recombination; most were hit directly, but some were hit in intergenic regions nearby (see Table S3 and Figure S1 for some examples). None of the 25,000 transformants reproducibly exhibited increased meiotic recombination. Additional features such as transposons and subtelomeric regions (such as the COS gene family) were also hit, but their positions could not be mapped precisely as a result of gene duplications at subtelomeric sequences (see http://www.nottingham.ac.uk/genetics/people/louis/research.php). Some of the transformants clearly had gross chromosomal rearrangements involving chromosome II or VI or both, which may have brought the heteroalleles into close proximity resulting in increased HR (Figure S2). The rDNA array was hit 14 times, which provides some measure of the rate of increased HR due to secondary events in the transformants. Many of the genes hit were involved in genetic processes such as DNA metabolism, chromatin remodeling, transcription, and replication; others were involved in additional cellular processes, such as cell membrane integrity, vesicle transport (particularly to the vacuole), mitochondrial proteins, and scaffold proteins or had unidentified functions. We investigated whether or not the genes isolated from our screen had previously been isolated in screens for genes that disrupt telomeric-related functions (Singer et al. 1998; Smith et al. 1999, 2004; Andrulis et al. 2002; Teixeira et al. 2002; Askree et al. 2004; Gatbonton et al. 2006; Addinall et al. 2008; Shachar et al. 2008). We found 13 genes: YKU80, STO1, HPR5, SGS1, RAD17, RAD24, HFI1, PHB2, APN1, IRC7, PCK6, RNR3, and UFD2 (Figure S1) that had previously been isolated in screens for genes that affect telomere length and capping mechanisms (Askree et al. 2004; Gatbonton et al. 2006; Addinall et al. 2008; Shachar et al. 2008).
YKU80 was the most frequently hit gene (five times) in our screen and is known to play a role in nuclear tethering, telomere maintenance, and TPE. However, although YKU70 also exhibited elevated HR in this assay when deleted, it was not hit, which shows that the screen was not saturating. We decided to delete additional genes involved in HR, TPE, telomere, and chromatin maintenance to try to decipher why a hyper-recombination phenotype was observed in a yku80-null mutant. Null alleles of SRS2, ESC1, RAD50, and TEL1 were introduced into either strain yAU15sir3Δ or strain yACT2. In addition, versions of yAU15 were constructed with sir2Δ, sir4Δ, or with a copy of MATa integrated at TRP1 (yAU36). None of the sir mutants or the haploid with both mating types exhibited elevated recombination, indicating that silencing per se is not involved in the repression of HR (Figure 3). Deletion of SRS2 resulted in a hyper-recombination phenotype in both strains, as expected due to its role in homologous recombination (Rong and Klein 1993). However, deletion of ESC1, involved in Sir4p-mediated tethering (Andrulis et al. 2002), and RAD50, which results in shortened telomeres (Askree et al. 2004), did not lead to hyper-recombination phenotypes in either strain. Therefore, neither tethering to the nuclear periphery nor short telomeres per se cause the elevated recombination. Deletion of TEL1, which also leads to shortened telomeres (Askree et al. 2004), did result in elevated recombination in the original assay strain yAU15sir3 but did not in yACT2 or yUA15. When tested in the sir mutants or the haploid version with both mating types, the deletion of TEL1 resulted in elevated recombination, indicating a diploid mating-type-specific repression of HR (Figure 3C). Similarly, the deletion of the gene encoding the chromatin-modifying enzyme, CHD1 (Tran et al. 2000; Luo et al. 2002), did not result in increased recombination in strain yACt2, yAU15, or yAU36, but did in strain yAU15sir3Δ. In contrast to TEL1, deletion of CHD1 resulted in elevated recombination only in the absence of sirs, indicating a role of chromatin rather than of mating type in repression of HR.
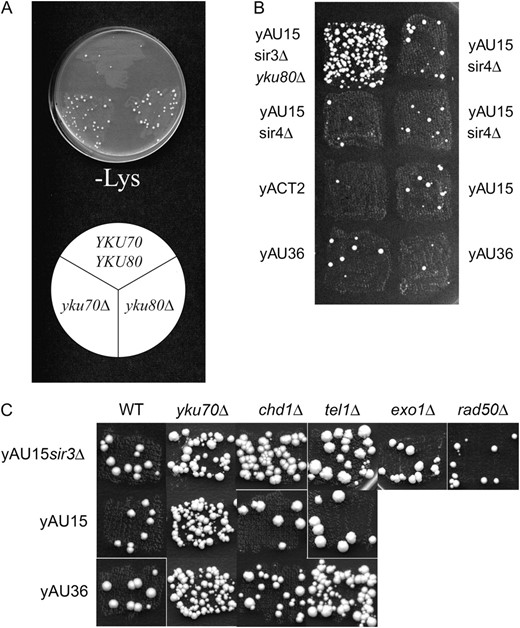
Increased recombination in mutant strains in comparison to sir mutants and parental strains. Single colonies were picked and patched onto YEPD medium and then replica plated onto complete synthetic medium or synthetic medium not containing lysine. The larger number of papillae formed on −Lys medium indicates a higher rate of mitotic ectopic recombination between the VIR∷URA3∷lys2-NdeI-1 and lys2-NdeI-2 heteroalleles in each strain. (A) The identical hyper-recombination phenotype displayed by both yku70 and yku80-null mutants in strain yAU15. (B) Deletion of sir2-4 does not increase the hyper-recombination phenotype displayed by yku80-mutants. The strains shown are the original hyper-recombination assay strain yAU15sir3Δ with yku80 tagged with the Tn3∷LEU2 transposon (sir3Δ yku80Δ); yAU15 harboring sir2, sir4, or sir3-null alleles (sir2Δ, sir4Δ, and sir3Δ); yACT2 (MATa); yAU15 (MATα); and yAU36 with MATa integrated at TRP1 (MATa-MATα). (C) A mutant strain harboring a tel1Δ allele displays elevated recombination in combination only with a sir3-null allele or in a MATa-MATα background (yAU36), and a strain harboring a chd1Δ allele displays elevated recombination in combination only with a sir3-null allele. Neither exo1 nor rad50 displayed hyper-recombination phenotypes.
In an attempt to interpret why YKU80 and the additional genes had been isolated from our screen, we decided to use information from the SGD to look for additional physical and genetic interactions that may explain the increased recombination rates (Table S3). Using data from the SGD, we found that 23 of the genes hit in our screen had some kind of interaction with another gene from our screen. Furthermore, 38 genes from our screen had physical or genetic interactions with the genes identified in previous screens (Askree et al. 2004; Gatbonton et al. 2006; Addinall et al. 2008; Shachar et al. 2008). We compiled a list using data from the SGD that contained all GO annotations, protein interactions, and genetic interactions for each of the genes hit in our screen (Table S3). A total of 1755 possible interactions were found, including 1121 protein interactions and 527 genetic interactions (File S1). Many of the additional interacting proteins had also been isolated in previous screens for genes maintaining telomere length (Askree et al. 2004; Gatbonton et al. 2006; Addinall et al. 2008; Shachar et al. 2008). In an attempt to determine whether or not any particular cell processes were significant in preventing recombination involving telomeric sequences, GO annotations for all the interacting genes were compiled and their frequency was determined (File S3). Many of the most significant processes were telomere or chromatin related, each processes that the yKu heterodimer is known to affect (Downs and Jackson 2004). We then decided to investigate the role of the yKu heterodimer in preventing genome instability by looking at recombination rates in its absence.
Mitotic ectopic recombination in yku70 and yku80-null mutants:
Having found evidence of a role for YKU80 in repressing ectopic recombination between subtelomeric and interstitial regions, we wanted to determine how this repression is mediated. The elevated recombination was not due solely to short telomeres, TPE, or tethering to the nuclear periphery on the basis of results from the deletion of sir2-4, rad50, tel1, and esc1. To determine whether or not the hyper-recombination could be attributed to YKU80 alone or to the yKu heterodimer in general, we introduced yku80 and yku70-null alleles into strain yAU15 and found that both mutants displayed identical hyper-recombination phenotypes on lysine drop-out medium (Figure 3).
To further characterize ectopic recombination in the absence of yKu, we constructed diploid strains carrying homozygous yku70-null alleles and heteroalleles of leu2 at gap 1 and site 12 in all three combinations at chromosomes XVR and XVIL (Figure 1B) and performed quantitative recombination assays. Our data showed that ectopic recombination rates were elevated between heteroalleles when they were placed at the Y′ ACS, with similar trends observed for both allelic and ectopic rates (Table 2). Recombination between leu2 heteroalleles both placed at the Y′ ACS was elevated 10-fold, whereas, when one heteroallele was place at an interstitial location, the elevation was only 4-fold. We also observed differences between the insertions at different ends; the allelic recombination rate between leu2 heteroalleles at Y′ ACS was elevated 16-fold at chromosome XVR, but only 4-fold at chromosome XVIL. A similar difference in rate between interstitial and Y′ ACS heteroalleles at both chromosome ends was also observed. These findings suggested that repression of recombination at XVR is more dependent on yKu than at XVIL; however, the wild-type recombination rate at XVIL is much higher than at XVR. In the absence of yKu, they both give similar rates of recombination. In contrast, heteroalleles placed at interstitial regions did not exhibit elevated recombination in either mutant strain nor did the allelic recombination rates between the control lys2 heteroalleles change in the absence of yKu.
An increase in recombination rate in yku80 mutants is not exclusive to heteroalleles placed in Y′ elements:
Using a leu2 heteroallele embedded in the Y′ ACS of chromosome XVIL, we observed that yKu helps to prevent recombination events between Y′ sequences and other interstitial sequences in the genome. To ascertain whether or not our observations were exclusive to Y′ elements or could be extended to additional subtelomeric elements such as core X, we performed recombination assays on strains harboring a heteroallele placed at an end not containing the Y′ element, XIIIR (Figure 1C and Figure 4). Our results showed that ectopic recombination in these strains was elevated in the absence of yKu, showing that the effect of yKu is not exclusive to Y′ elements.
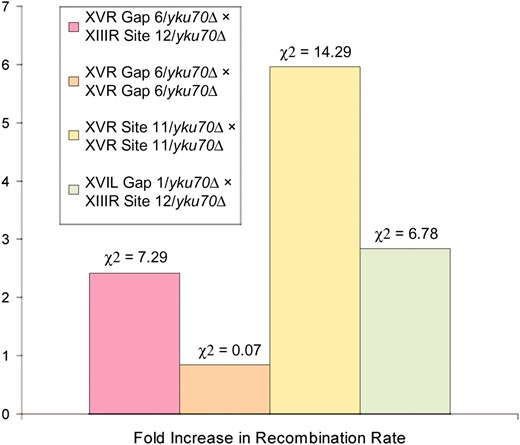
Recombination rates at X and X-Y′ ends in response to the deletion of YKU70. Results show the fold increase in recombination rate in yku70-null mutants relative to their corresponding wild-type strains. Four different types of recombination were tested in the presence and absence of YKU70 (see Figure 1C): ectopic recombination between X-Y′ and X-only ends (XVR gap 6 × XIIIR site 12 and XVIL gap 1 × XIIIR site 12); allelic recombination between internal sites at X-Y′ ends (XVR gap 6 × XVR gap 6); allelic recombination between subtelomeric sites at X-Y′ ends (XVR site 11 × XVR site 11). The rank-ordered chi-square values are shown for each data set.
Both yKu and core X are involved in repression of HR at telomeres:
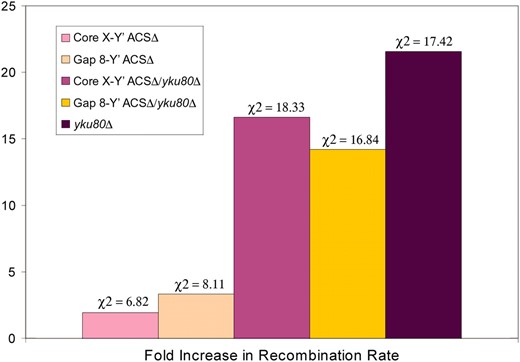
It has been suggested that a folding back of the terminal TG1-3 repeat sequences to subtelomeric sequences may help to protect chromosome ends and prevent genome instability (Pryde and Louis 1999; de Bruin et al. 2000, 2001). At native chromosome ends this fold-back may be targeted to the core X element (Pryde and Louis 1999), which may be involved in genome stability as well as TPE. To decipher whether or not core X sequences help to stabilize the genome by repressing recombination events, we performed recombination assays on strains harboring deleted subtelomeric core X sequences in the presence and absence of yku80-null alleles (Figure 5). Two core X deletions were tested, the first a deletion between gap 8 and the Y′ ACS and the second a deletion between the beginning of core X and the Y′ ACS (Figure 1). Strains carrying either of the core X deletions displayed a slight but significant increase in recombination rate when compared to a wild-type strain. In addition, the particular telomere that we tested (XVR) was slightly elongated (data not shown). No significant difference in recombination rate was observed between the two core X deletions. Deletion of YKU80 in the core X deletion strains resulted in an elevated recombination rate when compared to the same strain harboring a functional yKu80 protein, but one that was lower than that displayed in a strain harboring a yku80-null allele alone. The fact that there was no increase in HR in the double mutant over either the core X deletion or the yku80 deletion alone suggests that their repression of HR is via the same pathway.
Recombination rates between telomeres in response to deletion of core X and/or YKU80. Results show the fold increase in recombination rates between heteroalleles placed at site 12 at chromosome XVR relative to a wild-type YKU80. The corresponding rank-ordered chi-square values are shown above each data set.
DISCUSSION
In this study, we found that the rate of mitotic recombination near telomeres is elevated above the average rate for the entire genome. This is in contrast to the repression near telomeres seen for meiotic recombination (Barton et al. 2008). There appears to be a gradient of HR rate between heteroalleles when one copy is placed at a telomere and the second copy is moved toward the centromere. This is consistent with, but not exactly like, the recombination barrier previously proposed in our laboratory, where telomeres are sequestered from the rest of the genome, perhaps via tethering to the nuclear periphery (Pryde et al. 1997). A screen for mutations that allow increased HR between telomeric and internal sequences resulted in the identification of several genes involved in HR, some of which may be telomere specific. Many of these genes were general HR factors such as SRS2 and SGS1, several have telomeric properties, and several are involved in chromatin remodeling. The most frequently hit gene was YKU80, which has been shown to possess a telomere-specific role in suppressing HR (Polotnianka et al. 1998; Fellerhoff et al. 2000; Grandin and Charbonneau 2003; Celli et al. 2006). We can now modify our previously proposed fold-back model (Pryde and Louis 1999) to incorporate a yKu–core X interaction, perhaps via additional interactions with proteins bound to the X element (Sir2p-4p, ORC, etc.) rather than via a direct DNA-binding association.
Tethering per se is not the physical cause of repression of HR at telomeres as sir4 and esc1 mutants do not display a hyper-recombination phenotype. TPE per se is also not the cause of HR repression as none of the sir mutants exhibit elevated recombination. Telomere length is not directly involved in HR repression as rad50 and tel1 mutants with similar shortening to yKu mutants do not result in elevated recombination. The absence of an effect in rad50 also shows that NHEJ per se is not involved in the repression of HR. Although an exo1 mutant alone had no effect on HR, we cannot rule out that exo1-dependent single-stranded DNA in yKu mutant cells causes an increase in HR (Gravel et al. 1998). Interestingly, a chd1-sir3 null mutant did display elevated recombination, pointing to a possible link with silent heterochromatin. Perhaps the physical architecture preventing HR is related to the fold-back structure proposed for the chromosome ends, where the TG1-3 repeats interact with the core X element. This is supported by the fact that deletion of core X results in an increase in HR and that this increase is in the same pathway as yKu. Therefore, a novel role for yKu in maintaining genome stability by reducing HR has been identified in this study, and this may be manifested by a protective fold-back structure at chromosome ends. The interaction between core X and yKu may be involved in telomere length regulation in addition to repressing HR. This model provides testable predictions of the possible mechanism of HR repression and the yKu–core X interaction, such as whether yKu is physically associated with core X, whether this depends on the binding of other known core-X-associated proteins such as ORC and Abf1p, and whether or not the repression of HR requires yKu, NHEJ, or telomere biology-specific functions. Whatever the mechanism, the repression of HR in the vicinity of telomeres is yet another function of yKu.
Footnotes
Supporting information is available online at http://www.genetics.org/cgi/content/full/genetics.109.106674/DC1.
Footnotes
Communicating editor: E. Alani
Acknowledgements
We thank Anthony Underwood and Anna Timbrell for strain construction and early stages of screening. We also thank Sarah Sharp, Gianni Liti, and members of the Louis Laboratory for comments and criticisms. This work was supported by the Wellcome Trust.
References
Addinall, S. G., M. Downey, M. Yu, M. K. Zubko, J. Dewar et al.,
Andrulis, E. D., D. C. Zappulla, A. Ansari, S. Perrod, C. V. Laiosa et al.,
Askree, S. H., T. Yehuda, S. Smolikov, R. Gurevich, J. Hawk et al.,
Barton, A. B., M. R. Pekosz, R. S. Kurvathi and D. B. Kaback,
Bertuch, A. A., and V. Lundblad,
Boulton, S. J., and S. P. Jackson,
Burns, N., B. Grimwade, P. B. Ross-Macdonald, E. Y. Choi, K. Finberg et al.,
Celli, G. B., E. L. Denchi and T. de Lange,
Daley, J. M., P. L. Palmbos, D. Wu and T. E. Wilson,
de Bruin, D., S. M. Kantrow, R. A. Liberatore and V. A. Zakian,
de Bruin, D., Z. Zaman, R. A. Liberatore and M. Ptashne,
Downs, J. A., and S. P. Jackson,
Fellerhoff, B., F. Eckardt-Schupp and A. A. Friedl,
Flint, J., G. P. Bates, K. Clark, A. Dorman, D. Willingham et al.,
Fourel, G., E. Revardel, C. E. Koering and E. Gilson,
Gatbonton, T., M. Imbesi, M. Nelson, J. M. Akey, D. M. Ruderfer et al.,
Gietz, D., A. St Jean, R. A. Woods and R. H. Schiestl,
Goldman, A. S., and M. Lichten,
Grandin, N., and M. Charbonneau,
Gravel, S., M. Larrivee, P. Labrecque and R. J. Wellinger,
Hediger, F., F. R. Neumann, G. Van Houwe, K. Dubrana and S. M. Gasser,
Kramer, K. M., and J. E. Haber,
Krogh, B. O., and L. S. Symington,
Lea, D. E., and C. A. Coulson,
Lichten, M., and J. E. Haber,
Liti, G., and E. J. Louis,
Longtine, M. S., A. McKenzie, III, D. J. Demarini, N. G. Shah, A. Wach et al.,
Louis, E. J., and R. H. Borts,
Louis, E. J., and J. E. Haber,
Louis, E. J., and J. E. Haber,
Louis, E. J., E. S. Naumova, A. Lee, G. Naumov and J. E. Haber,
Luo, K., M. A. Vega-Palas and M. Grunstein,
McCusker, J. H., and J. E. Haber,
Palmbos, P. L., J. M. Daley and T. E. Wilson,
Paques, F., and J. E. Haber,
Polotnianka, R. M., J. Li and A. J. Lustig,
Pryde, F. E., and E. J. Louis,
Pryde, F. E., and E. J. Louis,
Pryde, F. E., H. C. Gorham and E. J. Louis,
Reenan, R. A., and R. D. Kolodner,
Ribes-Zamora, A., I. Mihalek, O. Lichtarge and A. A. Bertuch,
Rine, J., and I. Herskowitz,
Rong, L., and H. L. Klein,
Rothstein, R. J.,
Rutkowski, L. H., and R. E. Esposito,
Shachar, R., L. Ungar, M. Kupiec, E. Ruppin and R. Sharan,
Sherman, F., G. R. Fink and J. B. Hicks,
Singer, M. S., A. Kahana, A. J. Wolf, L. L. Meisinger, S. E. Peterson et al.,
Smith, J. S., E. Caputo and J. D. Boeke,
Smith, S., J. Y. Hwang, S. Banerjee, A. Majeed, A. Gupta et al.,
Stavenhagen, J. B., and V. A. Zakian,
Stellwagen, A. E., Z. W. Haimberger, J. R. Veatch and D. E. Gottschling,
Taddei, A., F. Hediger, F. R. Neumann, C. Bauer and S. M. Gasser,
Tam, A. T., B. L. Pike, A. Hammet and J. Heierhorst,
Teixeira, M. T., B. Dujon and E. Fabre,
Tham, W. H., J. S. Wyithe, P. Ko Ferrigno, P. A. Silver and V. A. Zakian,
Tran, H. G., D. J. Steger, V. R. Iyer and A. D. Johnson,