-
PDF
- Split View
-
Views
-
Cite
Cite
Amanda L Seyfert, Melania E A Cristescu, Linda Frisse, Sarah Schaack, W Kelley Thomas, Michael Lynch, The Rate and Spectrum of Microsatellite Mutation in Caenorhabditis elegans and Daphnia pulex, Genetics, Volume 178, Issue 4, 1 April 2008, Pages 2113–2121, https://doi.org/10.1534/genetics.107.081927
Close - Share Icon Share
Abstract
The effective use of microsatellite loci as tools for microevolutionary analysis requires knowledge of the factors influencing the rate and pattern of mutation, much of which is derived from indirect inference from population samples. Interspecific variation in microsatellite stability also provides a glimpse into aspects of phylogenetic constancy of mutational processes. Using long-term series of mutation-accumulation lines, we have obtained direct estimates of the spectrum of microsatellite mutations in two model systems: the nematode Caenorhabditis elegans and the microcrustacean Daphnia pulex. Although the scaling of the mutation rate with the number of tandem repeats is highly consistent across distantly related species, including yeast and human, the per-cell-division mutation rate appears to be elevated in multicellular species. Contrary to the expectations under the stepwise mutation model, most microsatellite mutations in C. elegans and D. pulex involve changes of multiple repeat units, with expansions being much more common than contractions.
MICROSATELLITE loci, composed of short tandem repeats, are ubiquitous in eukaryotic genomes, and because of their highly polymorphic nature are the marker of choice in population genetics and mapping studies (see reviews in Chambers and MacAvoy 2000; Ellegren 2000b, 2004; Selkoe and Toonen 2006). In an effort to maximize their utility, elucidation of the mutational properties and evolution of this class of repetitive DNA has been an active subject of research (Chakrabortyet al. 1997; DiRienzoet al. 1998; Neff and Gross 2001; Renwicket al. 2001; Dettman and Taylor 2004; Xuet al. 2005). The large amount of allelic variation in the number of repeats is most likely due to instability during replication, which involves DNA-polymerase pausing and dissociating from the template strand and subsequent incorrect reannealing of template and/or newly synthesized strands (Vigueraet al. 2001). Such slippage adds or deletes repeat units from the newly synthesized strand at a frequency that depends on factors such as overall length, repeat type, flanking sequence, and recombination rate (Schlötterer 2000). Previous estimates of microsatellite mutation rates in animals [direct estimates derived from individuals with known parentage (Brinkmannet al. 1998; Sajantilaet al. 1999; Leopoldino and Pena 2003; Henke and Henke 2006; Hohoffet al. 2007) and indirect estimates based on the application of models of microsatellite evolution to standing patterns of variation (Chakrabortyet al. 1997)] are as high as 10−3 and 10−4/locus/generation. Most studies show that the spectrum of microsatellite mutations is heavily dependent on allele size (Xuet al. 2000; Lai and Sun 2003) and suggest that the mechanism may be more complex than the simple stepwise mutation model (Brohedeet al. 2002; Huanget al. 2002; Calabrese and Durrett 2003). To account for this complexity, some models incorporate a length limit for expansion or a balance between slippage and point mutations (Kruglyaket al. 1998; Rose and Falush 1998).
Although the necessity of assaying large numbers of parent–offspring pairs for each locus makes the direct estimation of mutation rates laborious, such an approach can yield a nearly unbiased view of the mutational behavior of each locus. The following study makes use of long-term mutation-accumulation (MA) lines to provide robust estimates of the rates and size spectra of microsatellite mutations in two model systems: the nematode Caenorhabditis elegans and the water flea Daphnia pulex. We combine the results from these experiments with data previously reported in the literature for yeast and mammals to demonstrate that, even when scaled to a per-cell-division basis, microsatellite mutation rates tend to be elevated in multicellular species.
MATERIALS AND METHODS
Mutation-accumulation lines:
C. elegans and D. pulex MA lines were propagated using single, randomly chosen offspring, thereby reducing the strength of selection and allowing mutations to accumulate over time. Although selection associated with rare lethal, sublethal, and sterility-causing mutations cannot be eliminated entirely, our method provides the best experimental proxy for assaying the rate of origin of spontaneous mutations that are likely to survive for more than one or two generations in nature.
The design and maintenance of the C. elegans MA lines is described in Vassilieva and Lynch (1999) in detail. In brief, 80 MA lines were initiated from a descendant of the well-characterized Bristol N2 strain. Nematodes were cultured at 20° on nematode growth medium plates seeded with Escherichia coli strain OP50 using standard techniques (Sulston and Hodgkin 1988), with lines being propagated by random single-progeny descent. To prevent accidental line loss, the two previous generations were maintained at 15° as potential backups. For this study, DNA was extracted from individuals in generation 140.
D. pulex MA lines were initiated from the Log50 clone collected from Slimy Log Pond, Oregon (43° 50′ N, 124° 07′ W) in March 2000. Second-generation descendants of a single female were used to initiate 268 experimental lines, which were maintained at constant temperature (20°) and fed Scenedesmus obliquus three times per week. All MA lines were clonally propagated for each generation soon after their first clutch (∼12 days) by transferring either one or five (alternating each generation) random 1- to 2-day-old female offspring to a new beaker. As with the C. elegans lines, a backup system was used to advance a line when focal animals were dead or infertile. After 27 generations (on average), three to five clonal individuals were isolated from each line and used for DNA extraction.
Microsatellite markers:
C. elegans microsatellite loci were identified by a series of BLASTN searches of the genome sequence (databases at the Sanger Center, Cambridge, UK, and at Washington University, St. Louis) using query sequences of 10 perfect repeating units for each unique di-, tri-, and tetranucleotide. The largest two loci identified for each trinucleotide and tetranucleotide motif were selected for mutational analysis. For the dinucleotide repeats, a range of sizes, from 14 to 191 repeats, was selected for mutational analysis. Primers were designed for each locus (see appendix) using MacVector Sequence Analysis software (Version 4.5; Kodak).
D. pulex microsatellite loci were identified by BLASTN searches of the complete genome sequence (WFleaBase) and ranged in size from 13 to 47 dinucleotide repeats. Primers were designed using the program Primer3 (Rozen and Skaletsky 2000) or were obtained from Cristescuet al. (2006) and Omilianet al. (2006) (see appendix).
DNA extraction and amplification:
Nematodes produced in the middle of the parental reproductive period (∼30–100/plate) were transferred to proteinase K digestion buffer (Wierdlet al. 1997). Polymerase chain reactions were performed [10 μl reactions containing 67 mm Tris, pH 8.8, 6.7 mm MgCl2, 16.6 mm [NH4]2SO4, 10 mm 2-mercaptoethanol, each dNTP at 1 mm, each primer at 1 μm, 2–5 units of Thermus aquaticus polymerase (Perkin-Elmer/Cetus), and 2 μl of DNA extract] at annealing temperatures ranging from 45° to 65°, depending on the melting temperature of specific primers (see appendix for primer sequences). Daphnia genomic DNA was extracted using a cetyl trimethyl ammonium bromide extraction method (Doyle and Doyle 1987) and PCR amplified as described in Cristescuet al. (2006).
Genotyping:
C. elegans genotyping was conducted on an ABI 277 using ABI Genescan analysis software with the internal size standard Tamara 500 for all loci except T08H4, which used the internal size standard Tamara 2500. Most loci were multiplexed prior to genotyping. The loci F59D112, F16A11, F09F3, K06G5, and T08H4 were amplified individually and desalted (Millipore, ULTRAFREE MC) prior to analysis to improve the resolution of large products.
For D. pulex, we employed M13(-21) fluorescent-labeled primers (Schuelke 2000). Loci were multiplexed in groups of four to six according to their size and fluorescent labels (NED, PET, FAM, VIC). Genotyping was conducted on an Applied Biosystems (Foster City, CA) 3730 capillary electrophoresis genotyper and sized using Applied Biosystems Genemapper 3.7 software with LIZ 500 size standard.
All putative mutations were visually scored on the electropherogram data to eliminate call errors and other genotyping artifacts and subsequently verified using independent DNA amplification and genotyping. Only unambiguous, confirmed mutations are included in the following analyses.
Mutation rate estimates:
For C. elegans, new mutations are rapidly either lost or brought to homozygosity by self-fertilization, and rates of mutation (per allele per generation) were estimated as μ = −{ln(1 − n/l)}/t, where n is the number of mutations, t is the number of generations, and l is the number of lines. This expression accounts for the possibility of multiple substitutions per locus, which can occur for large arrays with high mutation rates. For D. pulex, new mutations remain heterozygous as a consequence of asexual line propagation, and rates of mutation (per allele per generation) were estimated as μ = n/(2tl). For this species, the experimental duration was short enough that the possibility of multiple substitutions per site could safely be ignored.
RESULTS
C. elegans:
Our screen of 80 C. elegans lines at the 140th generation of mutation accumulation, across 23 microsatellite loci with various repeat structures (di-, tri-, and tetranucleotides) and various lengths (from 12 to 191 repeats), reveals that 237 mutations occurred. Within the overlapping size class containing 12–43 repeats, a comparison of mutation rates for di-, tri-, and tetranucleotide loci reveals no significant differences in mutation rates (one-way ANOVA; F = 0.311, P = 0.738, d.f. = 2, 14) so the following analyses involve the pooled data. Overall, there is a strong positive scaling relationship between the mutation rate and the number of repeats per ancestral locus (Figure 1).
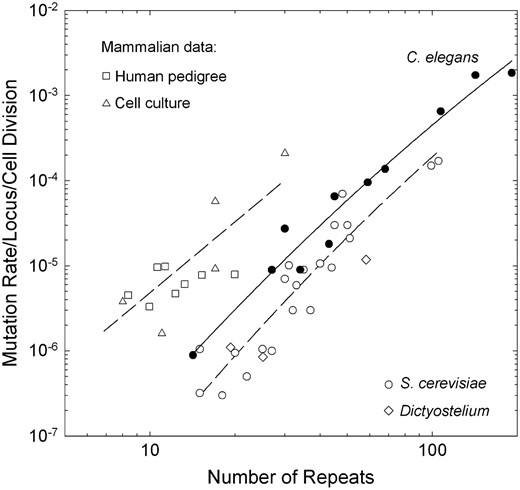
Scaling of the mutation rate with the number of repeat units in the ancestral locus on a per-cell-division basis. The data for C. elegans are taken directly from this study, with the data for the two shortest length classes being pooled, as most such loci exhibited no mutations. We assumed 10 germline cell divisions per generation for this species. The data for yeast are taken from direct observations on dinucleotide repeats from Wierdlet al. (1997) and Legendreet al. (2007). The data for Dictyostelium are taken from McConnellet al. (2007). The data for humans are pooled from several parent–offspring analyses (□) (Brinkmannet al. 1998; Leopoldino and Pena 2003; Henke and Henke 2006; Hohoffet al. 2007) and assume 200 germline cell divisions per generation. The mammalian cell-line data (▵) are taken from experimental investigations by Yamadaet al. (2002) and Hileet al. (2000) of mice and humans, respectively. Due to the lack of germline developmental data, we cannot include D. pulex in this comparison. The data for C. elegans and S. cerevisiae are fitted with second-order polynomials, respectively: y = −10.70 + 4.56x − 0.44x2 (r2 = 0.962) and y = −11.78 + 5.08x − 0.53x2 (r2 = 0.874). The mammalian data are fitted to the combined linear regression y = −8.09 + 2.77x (r2 = 0.749).
A substantial proportion of the mutations that we observed (73%; 173 of 237) involved gains or losses of more than one repeat (Table 1). This bias toward multi-step mutations is particularly evident in the largest loci (containing >100 repeats), where the vast majority of mutations (81%; 157 of 195) involve more than one repeat unit. The most frequent number of repeats gained for the largest loci is four, whereas smaller loci (containing <70 repeats) typically gained only one repeat. Overall, insertions greatly outnumbered deletions, accounting for 82% (195 of 237) of all mutations. Large insertions (>8 repeats) occurred only in loci >50 repeat units in length.
Allelic mutation rate estimates per generation for assayed loci in C. elegans and D. pulex
Locus . | Repeat . | No. of mutations . | Magnitude of mutation (no. of observations) . | Mutation rate . |
|---|---|---|---|---|
| C. elegans dimers | ||||
| C18F3 | (GT)14 | 0 | ||
| F10C2 | (GT)26 | 2 | +1 (1), +2 (1) | 0.000181 |
| F32B5 | (GT)34 | 1 | −1 (1) | 0.000090 |
| W01A6 | (GA)43 | 7 | −1 (4), +1 (2), +2 (1) | 0.000654 |
| F59D12 | (GT)59 | 10 | −20 (1), −1 (5), +2 (2), +7 (1), +8 (1) | 0.000954 |
| F16A11 | (GT)68 | 14 | −13 (1), −2 (1), −1 (5), +1 (4), +2 (2), +3 (1) | 0.001374 |
| F09F3 | (GT)107 | 48 | −62 (1), −28 (1), −2 (1), −1 (10), +1 (18), +2 (10), +3 (7), | 0.006545 |
| K06G5 | (GT)142 | 73 | −50 (1), −17 (1), −6 (1), −4 (1), −3 (1), −1 (2), +1 (6), +2 (8), +3 (18), +4 (29), +5 (2), +6 (3) | 0.017401 |
| T08H4 | (GT)191 | 74 | −4 (1), −3 (1), +1 (2), +2 (10), +3 (14), +4 (25), +5 (9), +6 (2), +7 (8), +10 (1), +22 (1) | 0.018502 |
| C. elegans trimers | ||||
| C27D9 | (AAT)12 | 0 | ||
| K08D12 | (ACC)12 | 0 | ||
| E02A10 | (GGA)13 | 0 | ||
| F47B8 | (GGA)13 | 0 | ||
| C24A3 | (TTC)13 | 1 | +1 (1) | 0.000090 |
| R01E6 | (TTC)14 | 0 | ||
| F10E7 | (CAA)15 | 0 | ||
| C03H5 | (AAT)28 | 2 | +1 (1), +8 (1) | 0.000181 |
| F35H10 | (GTA)30 | 3 | −1 (2), +2 (1) | 0.000273 |
| C. elegans tetramers | ||||
| C03B1 | (ATGG)17 | 0 | ||
| C16C8 | (TAGG)19 | 0 | ||
| C43B7 | (TAGG)26 | 0 | ||
| C13B7 | (TAGG)28 | 0 | ||
| B0564 | (AAAT)43 | 2 | +2 (2) | 0.000181 |
| D. pulex dimers | ||||
| P17D02 | (GA)13 | 2 | +2 (2) | 0.000139 |
| P17C11 | (GT)15 | 0 | ||
| P17C09 | (GA)15 | 0 | ||
| P17C08 | (GA)16 | 0 | ||
| P17B12 | (GA)19 | 1 | +2 (1) | 0.000071 |
| P17B11 | (GA)20 | 3 | +2 (2), +4 (1) | 0.000209 |
| P17A09 | (GA)26 | 0 | ||
| P17A07 | (GT)27 | 2 | +1 (2) | 0.000139 |
| P17A02 | (AG)30 | 2 | +2 (1), +5 (1) | 0.000154 |
| P17A01 | (GA)30 | 0 | ||
| P16H07 | (GA)34 | 1 | +1 (1) | 0.000100 |
| P16H05 | (GA)34 | 0 | ||
| P16H03 | (GA)34 | 2 | +1 (1), +7 (1) | 0.000140 |
| P16G10 | (GA)35 | 0 | ||
| P16F10 | (GA)36 | 0 | ||
| P16F02 | (AG)37 | 2 | −5 (1), +3 (1) | 0.000148 |
| P16F01 | (AG)38 | 4 | +1 (4) | 0.000282 |
| P16E08 | (GA)38 | 2 | +2 (1), +6 (1) | 0.000184 |
| P16E09 | (AG)38 | 1 | +2 (1) | 0.000075 |
| P16C05 | (AG)44 | 3 | −2 (1), +2 (1), +4 (1) | 0.000253 |
| P16C06 | (GA)44 | 3 | +1 (1), +4 (1), +5 (1) | 0.000217 |
| P16B10 | (GA)47 | 0 | ||
Locus . | Repeat . | No. of mutations . | Magnitude of mutation (no. of observations) . | Mutation rate . |
|---|---|---|---|---|
| C. elegans dimers | ||||
| C18F3 | (GT)14 | 0 | ||
| F10C2 | (GT)26 | 2 | +1 (1), +2 (1) | 0.000181 |
| F32B5 | (GT)34 | 1 | −1 (1) | 0.000090 |
| W01A6 | (GA)43 | 7 | −1 (4), +1 (2), +2 (1) | 0.000654 |
| F59D12 | (GT)59 | 10 | −20 (1), −1 (5), +2 (2), +7 (1), +8 (1) | 0.000954 |
| F16A11 | (GT)68 | 14 | −13 (1), −2 (1), −1 (5), +1 (4), +2 (2), +3 (1) | 0.001374 |
| F09F3 | (GT)107 | 48 | −62 (1), −28 (1), −2 (1), −1 (10), +1 (18), +2 (10), +3 (7), | 0.006545 |
| K06G5 | (GT)142 | 73 | −50 (1), −17 (1), −6 (1), −4 (1), −3 (1), −1 (2), +1 (6), +2 (8), +3 (18), +4 (29), +5 (2), +6 (3) | 0.017401 |
| T08H4 | (GT)191 | 74 | −4 (1), −3 (1), +1 (2), +2 (10), +3 (14), +4 (25), +5 (9), +6 (2), +7 (8), +10 (1), +22 (1) | 0.018502 |
| C. elegans trimers | ||||
| C27D9 | (AAT)12 | 0 | ||
| K08D12 | (ACC)12 | 0 | ||
| E02A10 | (GGA)13 | 0 | ||
| F47B8 | (GGA)13 | 0 | ||
| C24A3 | (TTC)13 | 1 | +1 (1) | 0.000090 |
| R01E6 | (TTC)14 | 0 | ||
| F10E7 | (CAA)15 | 0 | ||
| C03H5 | (AAT)28 | 2 | +1 (1), +8 (1) | 0.000181 |
| F35H10 | (GTA)30 | 3 | −1 (2), +2 (1) | 0.000273 |
| C. elegans tetramers | ||||
| C03B1 | (ATGG)17 | 0 | ||
| C16C8 | (TAGG)19 | 0 | ||
| C43B7 | (TAGG)26 | 0 | ||
| C13B7 | (TAGG)28 | 0 | ||
| B0564 | (AAAT)43 | 2 | +2 (2) | 0.000181 |
| D. pulex dimers | ||||
| P17D02 | (GA)13 | 2 | +2 (2) | 0.000139 |
| P17C11 | (GT)15 | 0 | ||
| P17C09 | (GA)15 | 0 | ||
| P17C08 | (GA)16 | 0 | ||
| P17B12 | (GA)19 | 1 | +2 (1) | 0.000071 |
| P17B11 | (GA)20 | 3 | +2 (2), +4 (1) | 0.000209 |
| P17A09 | (GA)26 | 0 | ||
| P17A07 | (GT)27 | 2 | +1 (2) | 0.000139 |
| P17A02 | (AG)30 | 2 | +2 (1), +5 (1) | 0.000154 |
| P17A01 | (GA)30 | 0 | ||
| P16H07 | (GA)34 | 1 | +1 (1) | 0.000100 |
| P16H05 | (GA)34 | 0 | ||
| P16H03 | (GA)34 | 2 | +1 (1), +7 (1) | 0.000140 |
| P16G10 | (GA)35 | 0 | ||
| P16F10 | (GA)36 | 0 | ||
| P16F02 | (AG)37 | 2 | −5 (1), +3 (1) | 0.000148 |
| P16F01 | (AG)38 | 4 | +1 (4) | 0.000282 |
| P16E08 | (GA)38 | 2 | +2 (1), +6 (1) | 0.000184 |
| P16E09 | (AG)38 | 1 | +2 (1) | 0.000075 |
| P16C05 | (AG)44 | 3 | −2 (1), +2 (1), +4 (1) | 0.000253 |
| P16C06 | (GA)44 | 3 | +1 (1), +4 (1), +5 (1) | 0.000217 |
| P16B10 | (GA)47 | 0 | ||
Allelic mutation rate estimates per generation for assayed loci in C. elegans and D. pulex
Locus . | Repeat . | No. of mutations . | Magnitude of mutation (no. of observations) . | Mutation rate . |
|---|---|---|---|---|
| C. elegans dimers | ||||
| C18F3 | (GT)14 | 0 | ||
| F10C2 | (GT)26 | 2 | +1 (1), +2 (1) | 0.000181 |
| F32B5 | (GT)34 | 1 | −1 (1) | 0.000090 |
| W01A6 | (GA)43 | 7 | −1 (4), +1 (2), +2 (1) | 0.000654 |
| F59D12 | (GT)59 | 10 | −20 (1), −1 (5), +2 (2), +7 (1), +8 (1) | 0.000954 |
| F16A11 | (GT)68 | 14 | −13 (1), −2 (1), −1 (5), +1 (4), +2 (2), +3 (1) | 0.001374 |
| F09F3 | (GT)107 | 48 | −62 (1), −28 (1), −2 (1), −1 (10), +1 (18), +2 (10), +3 (7), | 0.006545 |
| K06G5 | (GT)142 | 73 | −50 (1), −17 (1), −6 (1), −4 (1), −3 (1), −1 (2), +1 (6), +2 (8), +3 (18), +4 (29), +5 (2), +6 (3) | 0.017401 |
| T08H4 | (GT)191 | 74 | −4 (1), −3 (1), +1 (2), +2 (10), +3 (14), +4 (25), +5 (9), +6 (2), +7 (8), +10 (1), +22 (1) | 0.018502 |
| C. elegans trimers | ||||
| C27D9 | (AAT)12 | 0 | ||
| K08D12 | (ACC)12 | 0 | ||
| E02A10 | (GGA)13 | 0 | ||
| F47B8 | (GGA)13 | 0 | ||
| C24A3 | (TTC)13 | 1 | +1 (1) | 0.000090 |
| R01E6 | (TTC)14 | 0 | ||
| F10E7 | (CAA)15 | 0 | ||
| C03H5 | (AAT)28 | 2 | +1 (1), +8 (1) | 0.000181 |
| F35H10 | (GTA)30 | 3 | −1 (2), +2 (1) | 0.000273 |
| C. elegans tetramers | ||||
| C03B1 | (ATGG)17 | 0 | ||
| C16C8 | (TAGG)19 | 0 | ||
| C43B7 | (TAGG)26 | 0 | ||
| C13B7 | (TAGG)28 | 0 | ||
| B0564 | (AAAT)43 | 2 | +2 (2) | 0.000181 |
| D. pulex dimers | ||||
| P17D02 | (GA)13 | 2 | +2 (2) | 0.000139 |
| P17C11 | (GT)15 | 0 | ||
| P17C09 | (GA)15 | 0 | ||
| P17C08 | (GA)16 | 0 | ||
| P17B12 | (GA)19 | 1 | +2 (1) | 0.000071 |
| P17B11 | (GA)20 | 3 | +2 (2), +4 (1) | 0.000209 |
| P17A09 | (GA)26 | 0 | ||
| P17A07 | (GT)27 | 2 | +1 (2) | 0.000139 |
| P17A02 | (AG)30 | 2 | +2 (1), +5 (1) | 0.000154 |
| P17A01 | (GA)30 | 0 | ||
| P16H07 | (GA)34 | 1 | +1 (1) | 0.000100 |
| P16H05 | (GA)34 | 0 | ||
| P16H03 | (GA)34 | 2 | +1 (1), +7 (1) | 0.000140 |
| P16G10 | (GA)35 | 0 | ||
| P16F10 | (GA)36 | 0 | ||
| P16F02 | (AG)37 | 2 | −5 (1), +3 (1) | 0.000148 |
| P16F01 | (AG)38 | 4 | +1 (4) | 0.000282 |
| P16E08 | (GA)38 | 2 | +2 (1), +6 (1) | 0.000184 |
| P16E09 | (AG)38 | 1 | +2 (1) | 0.000075 |
| P16C05 | (AG)44 | 3 | −2 (1), +2 (1), +4 (1) | 0.000253 |
| P16C06 | (GA)44 | 3 | +1 (1), +4 (1), +5 (1) | 0.000217 |
| P16B10 | (GA)47 | 0 | ||
Locus . | Repeat . | No. of mutations . | Magnitude of mutation (no. of observations) . | Mutation rate . |
|---|---|---|---|---|
| C. elegans dimers | ||||
| C18F3 | (GT)14 | 0 | ||
| F10C2 | (GT)26 | 2 | +1 (1), +2 (1) | 0.000181 |
| F32B5 | (GT)34 | 1 | −1 (1) | 0.000090 |
| W01A6 | (GA)43 | 7 | −1 (4), +1 (2), +2 (1) | 0.000654 |
| F59D12 | (GT)59 | 10 | −20 (1), −1 (5), +2 (2), +7 (1), +8 (1) | 0.000954 |
| F16A11 | (GT)68 | 14 | −13 (1), −2 (1), −1 (5), +1 (4), +2 (2), +3 (1) | 0.001374 |
| F09F3 | (GT)107 | 48 | −62 (1), −28 (1), −2 (1), −1 (10), +1 (18), +2 (10), +3 (7), | 0.006545 |
| K06G5 | (GT)142 | 73 | −50 (1), −17 (1), −6 (1), −4 (1), −3 (1), −1 (2), +1 (6), +2 (8), +3 (18), +4 (29), +5 (2), +6 (3) | 0.017401 |
| T08H4 | (GT)191 | 74 | −4 (1), −3 (1), +1 (2), +2 (10), +3 (14), +4 (25), +5 (9), +6 (2), +7 (8), +10 (1), +22 (1) | 0.018502 |
| C. elegans trimers | ||||
| C27D9 | (AAT)12 | 0 | ||
| K08D12 | (ACC)12 | 0 | ||
| E02A10 | (GGA)13 | 0 | ||
| F47B8 | (GGA)13 | 0 | ||
| C24A3 | (TTC)13 | 1 | +1 (1) | 0.000090 |
| R01E6 | (TTC)14 | 0 | ||
| F10E7 | (CAA)15 | 0 | ||
| C03H5 | (AAT)28 | 2 | +1 (1), +8 (1) | 0.000181 |
| F35H10 | (GTA)30 | 3 | −1 (2), +2 (1) | 0.000273 |
| C. elegans tetramers | ||||
| C03B1 | (ATGG)17 | 0 | ||
| C16C8 | (TAGG)19 | 0 | ||
| C43B7 | (TAGG)26 | 0 | ||
| C13B7 | (TAGG)28 | 0 | ||
| B0564 | (AAAT)43 | 2 | +2 (2) | 0.000181 |
| D. pulex dimers | ||||
| P17D02 | (GA)13 | 2 | +2 (2) | 0.000139 |
| P17C11 | (GT)15 | 0 | ||
| P17C09 | (GA)15 | 0 | ||
| P17C08 | (GA)16 | 0 | ||
| P17B12 | (GA)19 | 1 | +2 (1) | 0.000071 |
| P17B11 | (GA)20 | 3 | +2 (2), +4 (1) | 0.000209 |
| P17A09 | (GA)26 | 0 | ||
| P17A07 | (GT)27 | 2 | +1 (2) | 0.000139 |
| P17A02 | (AG)30 | 2 | +2 (1), +5 (1) | 0.000154 |
| P17A01 | (GA)30 | 0 | ||
| P16H07 | (GA)34 | 1 | +1 (1) | 0.000100 |
| P16H05 | (GA)34 | 0 | ||
| P16H03 | (GA)34 | 2 | +1 (1), +7 (1) | 0.000140 |
| P16G10 | (GA)35 | 0 | ||
| P16F10 | (GA)36 | 0 | ||
| P16F02 | (AG)37 | 2 | −5 (1), +3 (1) | 0.000148 |
| P16F01 | (AG)38 | 4 | +1 (4) | 0.000282 |
| P16E08 | (GA)38 | 2 | +2 (1), +6 (1) | 0.000184 |
| P16E09 | (AG)38 | 1 | +2 (1) | 0.000075 |
| P16C05 | (AG)44 | 3 | −2 (1), +2 (1), +4 (1) | 0.000253 |
| P16C06 | (GA)44 | 3 | +1 (1), +4 (1), +5 (1) | 0.000217 |
| P16B10 | (GA)47 | 0 | ||
D. pulex:
Our screen of 268 D. pulex MA lines, after an average of 27 generations of mutation accumulation, across 22 dinucleotide microsatellite loci yields a total of 28 mutations (Table 1). The number of repeat units (at the ancestral loci) analyzed ranges from 13 to 47, and most of the detected mutations involve additions of 2–7 repeats, with only 32% (9 of 28) showing single-repeat additions and no observed single-repeat losses. The mutation rate of large loci (34–47 repeats, averaging 38.2) is 1.16 (0.30) × 10−4/allele/generation, which is an order of magnitude higher than the average for loci with 30 or fewer repeats (averaging 21.1): 7.11 (2.59) × 10−5/allele/generation.
DISCUSSION
Despite the central significance of microsatellite mutations to issues of genomic instability, forensic testing, and population genetic analyses, the rate of origin and spectrum of effects of such mutations are still poorly understood, with many estimates being derived from reporter constructs in yeast cultures (e.g., Henderson and Petes 1992; Strandet al. 1995; Wierdlet al. 1996; Siaet al. 1997). Our long-term series of mutation-accumulation lines of C. elegans and D. pulex provide a useful platform for a more direct evaluation of the properties of microsatellite mutations in two key model organisms.
As in previous studies (Wierdlet al. 1997; Brinkmannet al. 1998; Kayseret al. 2000; Becket al. 2003; Whittakeret al. 2003; Legendreet al. 2007), we find a strong correlation between allele size (repeat number) and mutation rate in C. elegans (Figure 1). In addition, although the variation of repeat numbers among loci sampled in D. pulex does not permit a formal evaluation of length dependence over a large range, as noted above, this organism also exhibits an order of magnitude span in the average mutation rate for small vs. large loci (averaging 21.1 and 38.2 repeats, respectively). On a per-generation basis, the average rates for these two size classes in D. pulex are not significantly different from the C. elegans rates (from the regression line in Figure 1).
Because all but one of the dinucleotide repeats assayed in C. elegans were GT, and all but two loci assayed in D. pulex were GA repeats, in addition to there being no significant differences in mutation rates among loci with different repeat-unit lengths in C. elegans, the scaling noted above does not appear to be a spurious consequence of changes in base composition with repeat lengths. Nevertheless, further investigation of a multiplicity of repeat types is desirable to determine definitively whether the size-specific scaling that we have noted can be generalized across repeat types. Because some evidence suggests that this may not be the case (Bicharaet al. 2000; Hileet al. 2000; Eckertet al. 2002; Gragget al. 2002), any comparative analysis of mutation-rate differences among species is best performed with ancestral microsatellites with similar nucleotide composition and size.
On a per-generation basis, the mutation rates that we report for C. elegans and D. pulex are much lower than those previously estimated for humans (Heyeret al. 1997; Brinkmannet al. 1998; Sajantilaet al. 1999; Kayseret al. 2000; Leopoldino and Pena 2003; Dupuyet al. 2004; Ballardet al. 2005; Gusmaoet al. 2005; Henke and Henke 2006; Yanet al. 2006; Hohoffet al. 2007; Leeet al. 2007), but much higher than those for yeast (Henderson and Petes 1992; Strandet al. 1993; Johnsonet al. 1996; Siaet al. 1997, 2001; Wierdlet al. 1997; Hawket al. 2005; Legendreet al. 2007). In principle, such differences could simply arise as a consequence of the pronounced variation in the numbers of germline cell divisions per generation that exists among these species (1, 10, and ∼200, respectively, in Saccharomyces cerevisiae, C. elegans, and Homo sapiens; Kimble and Ward 1998; Crow 2000). Unfortunately, the necessary germline developmental data for this calculation do not exist for Daphnia. However, although the mutation rates for C. elegans and S. cerevisiae per cell division scale in an almost identical fashion with repeat number in the range of overlapping data (loci with ∼15–100 repeats), the rates for the former are 2.5–5.1 times the latter (Figure 1). Because all of the experimental results for yeast rely on GT dinucleotide repeats, as was the case for almost all such loci for C. elegans, these results cannot be an artifact of nucleotide-composition differences among the loci surveyed in these two taxa.
Although the range of data is more limited, the size scaling of mammalian microsatellite mutation rates is also similar to that of C. elegans and yeast (Figure 1). However, the mammalian rates are elevated even further on a per-cell-division basis, and this is true for loci monitored in mitotically reproducing cell cultures as well as for surveys of parent–offspring relationships. For loci with 15 repeats, the mutation rates for S. cerevisiae, C. elegans, and mammals scale as 1:5:67. Three of the five mammalian cell-line estimates involve GT repeats, and two of these (involving 8 and 30 repeats) reside above the regression line for mammals, so the elevated mammalian rate does not appear to be an artifact of relying on repeat types with unusual properties.
With these analyses being based on just three organisms, it is reasonable to ask whether the observed increase in the per-cell-division microsatellite mutation rate in organisms with increasing levels of multicellularity is simply a chance relationship, but the limited data that are available for other taxa support such an assertion. First, although few thorough experimental investigations have been performed with other unicellular organisms, the predominantly single-celled slime mold Dictyostelium discoideum exhibits patterns of microsatellite instability (McConnellet al. 2007) that are not discernibly different from those of yeast (Figure 1). Second, several surveys have been performed on small di- and trinucleotide repeats in Drosophila melanogaster (Schlöttereret al. 1998; Schuget al. 1998; FernandoVázquezet al. 2000). On average, these loci harbor 13.4 repeats and exhibit an average mutation rate of 1.87 (0.77) × 10−6/allele/cell division (after accounting for the ∼36 germline cell divisions/generation in this species; Drost and Lee 1995), which is ∼2.5 times the expected rate for C. elegans. Third, avian microsatellite rates appear to be very high as well. For example, a (GT)26 repeat in the superb fairy-wren (Malurus cyaneus) has a mutation rate of 0.011/allele/generation (Becket al. 2003), and rates in the range of 0.01–0.05/allele/generation have been reported for tetra- and pentanucleotide repeat loci containing 20–60 units in the barn swallow (Hirundo rustica) (Brohedeet al. 2002). Even assuming 200 germline cell divisions (the number in humans), the per-cell-division rates for these birds are in the range of the estimates for mammalian genomes.
Finally, although limited work has been done on the microsatellite mutation rate in plants, the data are again consistent with an elevated rate of instability relative to the situation in unicellular species. For example, a whole-plant analysis of a (GT)17 reporter construct in Arabidopsis thaliana implies a mutation rate of 9.10 × 10−6/allele/cell division (Leonardet al. 2003), which is ∼5.2 times the expected rate for C. elegans loci with this number of repeats. The per-generation rate of mutation for a (G)16 mononucleotide repeat in the same species (Azaiezet al. 2006) is ∼350 times the per-cell-division rate for a (G)18 repeat in S. cerevisiae (Siaet al. 1997, 2001). In a survey of maize microsatellites with a wide variety of motifs, Vigourouxet al. (2002) estimate an average per-generation mutation rate for loci with an average of 24.5 repeats, which after accounting for the estimated 49 cell divisions per life cycle (in the cellular line of descent from seed to seed) (Otto and Walbot 1990), yields a rate per cell division of 1.57 × 10−5/allele, ∼2.6 times the expected rate for C. elegans loci of comparable length. In durum wheat (Triticum turgidum), the average mutation rate for a variety of dinucleotide repeats with an average ancestral state of 23.2 repeats is 2.4 × 10−4/allele/generation (Thuilletet al. 2002), whereas the per-generation rates for (TAA) repeats with average lengths of 27.4 and 42.0 repeats are 2.39 (0.72) × 10−3 and 1.90 (0.60) × 10−2/allele, respectively (Udupa and Baum 2001). As the latter three rates are, respectively, 472, 270, and 545 times the expected C. elegans rates per cell division—assuming the number of cell divisions in these annual plants is comparable to that of maize—they, too, strongly support the idea that microsatellites are much less stable in land plants than in unicellular species.
Thus, a broad set of observations suggests that, relative to the situation in unicellular species, there is a substantial decline in the ability to repair premutations resulting from replication slippage in both metazoans, particularly mammals, and land plants. This apparent reduction in the ability to repair a common form of DNA instability may be a consequence of the compromised ability of natural selection to maintain optimal features of genomic architecture in lineages of multicellular organisms with reduced effective population sizes (Lynch 2006, 2007). For example, because mismatch repair (MMR) plays a central role in the elimination of post-replication slippage errors, specifically in microsatellite loci (Strandet al. 1995; Wierdlet al. 1996, 1997; Eshleman and Markowitz 1996; Boyeret al. 2002; Yamadaet al. 2002), a decline in the efficiency of MMR might be responsible in part for increased mutation rates in multicellular species.
A large number of studies have compared microsatellite mutation rates in MMR-proficient vs. deficient lines of yeast (Strandet al. 1993, 1995; Johnsonet al. 1996; Siaet al. 1997, 2001; Tranet al. 1997; Wierdlet al. 1997; Drotschmannet al. 1999; Gragget al. 2002; Hawket al. 2005; Gammieet al. 2007), and, collectively, these imply an inflation in the average mutation rate by a factor of ∼1346 (194) in the absence of MMR. In contrast, the loss of MMR appears to inflate microsatellite mutation rates by a factor of only ∼257 (132) in C. elegans (Degtyarevaet al. 2002; Denveret al. 2005), implying a reduced efficiency of MMR. Because these studies have been performed with a wide variety of experimental methods, this comparison should be interpreted with caution, but it is notable that C. elegans, unlike yeast, is lacking the MSH3 gene (Eisen 1998), whose product plays a central role in the mismatch repair of small insertion/deletions. On the other hand, MSH3 is present in mammalian genomes, and very limited work with MMR-deficient mammalian cell lines suggests an inflation in microsatellite mutation rates in the range of 2600–33,000 times (Hanfordet al. 1998; Abuinet al. 2000), so it is not clear that the elevated microsatellite mutation rates in mammals can be attributed to a low efficiency of MMR. One alternative is that replication errors (prior to mismatch repair) are generated at a higher rate in animals than in yeast, because of either a higher rate of template slippage or a lower efficiency of proofreading by the DNA polymerase. In yeast, a knockout of the polymerase proofreading domain increases microsatellite mutation rates by only a few fold (Strandet al. 1993; Pavlovet al. 2006). Unfortunately, no studies of this nature have been performed with mammalian DNA polymerases, but the overall fidelity of mammalian polymerases does not appear to be unusually low in comparison to yeast (Kunkel 1985; Kunkelet al. 1987, 1991; Thomaset al. 1991). Thus, the mechanistic basis for the apparent differences in microsatellite stability among taxa remains largely unresolved.
In contrast with the situation in maize (Vigourouxet al. 2002), but in accordance with observations in humans (Huanget al. 2002), we found that ∼72% (192 of 265) of microsatellite mutations in C. elegans and D. pulex involve multi-step changes, with a strong tendency toward insertions, especially in ancestral alleles with large numbers of repeats. Thus, a simple stepwise mutation model appears to inadequately explain the pattern of mutation in both species. Wierdlet al. (1997) also noted a greatly elevated bias toward insertions with increasing numbers of repeat units, and insertion biases have been found within human microsatellites as well (Ellegren 2000a; Kayseret al. 2000; Dupuyet al. 2004). Such observations appear to be inconsistent with the proposed existence of a critical repeat length above which contraction mutations outnumber expansions (Rose and Falush 1998; Xuet al. 2000).
Finally, with an overall insertion:deletion ratio of 195:42 in C. elegans and 26:2 in D. pulex, it is clear that microsatellite loci in the genomes of these two species are not in drift-mutation equilibrium, as under the latter hypothesis the numbers of observed insertions and deletions should be equal. Although, in principle, variants with very large repeat numbers may also be broken up by point mutations (Kruglyaket al. 1998), if the size distributions of alleles in both species are in equilibrium in nature, some other force, presumably either natural selection or biased gene conversion, must be opposing the accumulation of insertions.
APPENDIX
Primers used in the microsatellite survey
Locus . | Repeat . | Forward primer 5′-3′/reverse primer 5′-3′ . |
|---|---|---|
| C. elegans dimers | ||
| C18F3 | (GT)14 | AAGGGCATGAGGAGAAGTCAACGGGAGCAGCTCTCGTTCTGATTGC |
| F10C2 | (GT)26 | GCTCCACTTTTCAACATCCTTCCTTGTCCCCGCTATTTCGCTC |
| F32B5 | (GT)34 | GCACTTTTCCAACAAAAAAATCCGCCCACACGAGACATGCAATAACTACC |
| W01A6 | (GA)45 | CCCCCGAATCTCATTTCAGTTGTTAGCACCCTTCCGTTTTCACGCC |
| F59D12 | (GT)59 | ACCCCAATGGACTACGTTGTGCACGTGACGCCTTTGGTTCATAG |
| F16A11 | (GT)68 | CTTCTTCTTCTTCCTCCGACCGCAGAGGGGGTGATCTTTATGCC |
| F09F3 | (GT)107 | GCAATCCTACGGTACACCCATACCCGTATTTTGACATTAGGCAGGTGC |
| K06G5 | (GT)142 | AGCTTGAGGAAGAAGGTGGGGTGTTCGTCCATTTGAGCACC |
| T08H4 | (GT)191 | TGGCTCATCACTTGTGTTCAACCCGGCTCCTCTCTGATTTGTCTG |
| C. elegans trimers | ||
| C27D9 | (AAT)12 | AATCGTCGTCGGAACTCAGATAAGGCAAGTGTTGAGGCAAACAGAAG |
| K08D12 | (ACC)12 | GAACGTGCATCAAGTGTAATTCGCTGGCACCCGATATGTGTGAAG |
| E02A10 | (GGA)13 | GTAGCAAGAGCCGAAAGCTGAGTGTCAAGCCTACACCACCACTAGC |
| F47B8 | (GGA)13 | GCTCCTCCAGAATATCCAGCTTCCTATGGTCTGCTCGATCAGTTGTG |
| C24A3 | (TTC)13 | CGCATCTTCTTTTCGGAGCAAGTCATCCTCCTTCCAACACTGCC |
| R01E6 | (TTC)14 | TCCCCTCGTAATTCCCAAAATCCCAAAAGGAGATGCAGAAGAGTAGG |
| F10E7 | (CAA)15 | ACCAAAGTCAACCCGACTACAGCTCGAGTGTGATGCGAACAGTTG |
| C03H5 | (AAT)28 | AACAAAAATGTGGCAGGGAGGGGTTACGGTAGTGGTACTGTAGG |
| F35H10 | (GTA)30 | CGCTCAGAACCATGTTTAAGATTTCCTTTTATGAGTAACTTTGGTGCAGG |
| C. elegans tetramers | ||
| C03B1 | (ATGG)17 | GAATCTGTTCTTCCTTCCTTCGCCGTTCTCCTTTTATCATCAAC |
| C16C8 | (TAGG)19 | TCCTGCCTACCTGCCCATTAGGGCGTGCAGACATAAGATAGG |
| C43B7 | (TAGG)26 | GATGCAGACAAAAATACCGAGAGGGACTAATCACCCACAAACCGAAAC |
| C13B7 | (TAGG)28 | CAAAGTTAGCAAGAACCCAGTTGGTCGCACCTACCTACACCCTTTCTG |
| B0564 | (AAAT)43 | GACCATTAGATGAATTTCCCGAGCAACAGAATTTTTAC |
| D. pulex dimers | ||
| P17D02 | (GA)13 | CCGAATGTCTAACCCACACCTGCACATATTGCCCGTGTAT |
| P17C11 | (GT)15 | ACGGTAGCGGTATTGGAAGATGCAGCGATTCAAATAATGG |
| P17C09 | (GA)15 | GTATGGGCCGGCTAAGGATAGGAATTTGGCGCATCGAG |
| P17C08 | (GA)16 | CCGTCTCCAACCAATTCACTTAAACTTCGGGGACCTTGTG |
| P17B12 | (GA)19 | TCCCGGTTTATCACAGTTCCGCAAAAATCGCAGGTTTCTC |
| P17B11 | (GA)20 | TAACAAGTGGCTGCATCCTGCAGAGGAGGTAACGCAAAGG |
| P17A09 | (GA)26 | CGTCCATCATGACTTGGATCTCACCATGAGTGAAAGCCAGA |
| P17A07 | (GT)27 | AGAGTGAAGGCAGGAAACCACGGAGACGACCAAGGTGTA |
| P17A02 | (GA)30 | TTATTCATGTACGCCCGTTGGCAGTGCTGTCCTGTGAGAC |
| P17A01 | (GA)30 | TCATGTACGCCCGTTGTAAAGCCATGGACGTGACGTATAA |
| P16H07 | (GA)34 | GCACGTGACACATTTGAAGCGTTCAGCACGAAAATGACGA |
| P16H05 | (GA)34 | AGCCGTACCTGTGATGTGTGACGAGCAATACACGATGACG |
| P16H03 | (GA)34 | TGAACGAGGGGAAAATCAAAAACCTGCTCTTCACGACGAC |
| P16G10 | (GA)35 | CTGAGAGCGAGACCGACAGCGACATGGATGATGTTGAGG |
| P16F10 | (GA)36 | GGTGTATCATTTTTGATTGAATTTTCTAGTCGCATTTGCGCCTTT |
| P16F02 | (GA)37 | AGTCTGGAGAGCCAAAGCAGCTCGTCGACCATGAGAACAA |
| P16F01 | (GA)38 | ATTCGAACCGACGGAAAACTCAACGGCAGTGGCAATACTA |
| P16E08 | (GA)38 | CAACGGATGGTGGAATAACCTCCGCGTAGCCTCATAGAAT |
| P16E09 | (GA)38 | CTCGGGAAGGTTTCAGACAGACCCAAAAGAGCGAGGAACT |
| P16C05 | (GA)44 | GATTCGAACCACCCAACTGTGTGGGGTACTCAGCGAAAGA |
| P16C06 | (GA)44 | CCGAGAGCTTCTCACTGCTTACGAAATCGATCCCTCTTCC |
| P16B10 | (GA)47 | CAAGTGGGCGTGTACAACAGTGTCAGCGCGTCTAGAGAAA |
Locus . | Repeat . | Forward primer 5′-3′/reverse primer 5′-3′ . |
|---|---|---|
| C. elegans dimers | ||
| C18F3 | (GT)14 | AAGGGCATGAGGAGAAGTCAACGGGAGCAGCTCTCGTTCTGATTGC |
| F10C2 | (GT)26 | GCTCCACTTTTCAACATCCTTCCTTGTCCCCGCTATTTCGCTC |
| F32B5 | (GT)34 | GCACTTTTCCAACAAAAAAATCCGCCCACACGAGACATGCAATAACTACC |
| W01A6 | (GA)45 | CCCCCGAATCTCATTTCAGTTGTTAGCACCCTTCCGTTTTCACGCC |
| F59D12 | (GT)59 | ACCCCAATGGACTACGTTGTGCACGTGACGCCTTTGGTTCATAG |
| F16A11 | (GT)68 | CTTCTTCTTCTTCCTCCGACCGCAGAGGGGGTGATCTTTATGCC |
| F09F3 | (GT)107 | GCAATCCTACGGTACACCCATACCCGTATTTTGACATTAGGCAGGTGC |
| K06G5 | (GT)142 | AGCTTGAGGAAGAAGGTGGGGTGTTCGTCCATTTGAGCACC |
| T08H4 | (GT)191 | TGGCTCATCACTTGTGTTCAACCCGGCTCCTCTCTGATTTGTCTG |
| C. elegans trimers | ||
| C27D9 | (AAT)12 | AATCGTCGTCGGAACTCAGATAAGGCAAGTGTTGAGGCAAACAGAAG |
| K08D12 | (ACC)12 | GAACGTGCATCAAGTGTAATTCGCTGGCACCCGATATGTGTGAAG |
| E02A10 | (GGA)13 | GTAGCAAGAGCCGAAAGCTGAGTGTCAAGCCTACACCACCACTAGC |
| F47B8 | (GGA)13 | GCTCCTCCAGAATATCCAGCTTCCTATGGTCTGCTCGATCAGTTGTG |
| C24A3 | (TTC)13 | CGCATCTTCTTTTCGGAGCAAGTCATCCTCCTTCCAACACTGCC |
| R01E6 | (TTC)14 | TCCCCTCGTAATTCCCAAAATCCCAAAAGGAGATGCAGAAGAGTAGG |
| F10E7 | (CAA)15 | ACCAAAGTCAACCCGACTACAGCTCGAGTGTGATGCGAACAGTTG |
| C03H5 | (AAT)28 | AACAAAAATGTGGCAGGGAGGGGTTACGGTAGTGGTACTGTAGG |
| F35H10 | (GTA)30 | CGCTCAGAACCATGTTTAAGATTTCCTTTTATGAGTAACTTTGGTGCAGG |
| C. elegans tetramers | ||
| C03B1 | (ATGG)17 | GAATCTGTTCTTCCTTCCTTCGCCGTTCTCCTTTTATCATCAAC |
| C16C8 | (TAGG)19 | TCCTGCCTACCTGCCCATTAGGGCGTGCAGACATAAGATAGG |
| C43B7 | (TAGG)26 | GATGCAGACAAAAATACCGAGAGGGACTAATCACCCACAAACCGAAAC |
| C13B7 | (TAGG)28 | CAAAGTTAGCAAGAACCCAGTTGGTCGCACCTACCTACACCCTTTCTG |
| B0564 | (AAAT)43 | GACCATTAGATGAATTTCCCGAGCAACAGAATTTTTAC |
| D. pulex dimers | ||
| P17D02 | (GA)13 | CCGAATGTCTAACCCACACCTGCACATATTGCCCGTGTAT |
| P17C11 | (GT)15 | ACGGTAGCGGTATTGGAAGATGCAGCGATTCAAATAATGG |
| P17C09 | (GA)15 | GTATGGGCCGGCTAAGGATAGGAATTTGGCGCATCGAG |
| P17C08 | (GA)16 | CCGTCTCCAACCAATTCACTTAAACTTCGGGGACCTTGTG |
| P17B12 | (GA)19 | TCCCGGTTTATCACAGTTCCGCAAAAATCGCAGGTTTCTC |
| P17B11 | (GA)20 | TAACAAGTGGCTGCATCCTGCAGAGGAGGTAACGCAAAGG |
| P17A09 | (GA)26 | CGTCCATCATGACTTGGATCTCACCATGAGTGAAAGCCAGA |
| P17A07 | (GT)27 | AGAGTGAAGGCAGGAAACCACGGAGACGACCAAGGTGTA |
| P17A02 | (GA)30 | TTATTCATGTACGCCCGTTGGCAGTGCTGTCCTGTGAGAC |
| P17A01 | (GA)30 | TCATGTACGCCCGTTGTAAAGCCATGGACGTGACGTATAA |
| P16H07 | (GA)34 | GCACGTGACACATTTGAAGCGTTCAGCACGAAAATGACGA |
| P16H05 | (GA)34 | AGCCGTACCTGTGATGTGTGACGAGCAATACACGATGACG |
| P16H03 | (GA)34 | TGAACGAGGGGAAAATCAAAAACCTGCTCTTCACGACGAC |
| P16G10 | (GA)35 | CTGAGAGCGAGACCGACAGCGACATGGATGATGTTGAGG |
| P16F10 | (GA)36 | GGTGTATCATTTTTGATTGAATTTTCTAGTCGCATTTGCGCCTTT |
| P16F02 | (GA)37 | AGTCTGGAGAGCCAAAGCAGCTCGTCGACCATGAGAACAA |
| P16F01 | (GA)38 | ATTCGAACCGACGGAAAACTCAACGGCAGTGGCAATACTA |
| P16E08 | (GA)38 | CAACGGATGGTGGAATAACCTCCGCGTAGCCTCATAGAAT |
| P16E09 | (GA)38 | CTCGGGAAGGTTTCAGACAGACCCAAAAGAGCGAGGAACT |
| P16C05 | (GA)44 | GATTCGAACCACCCAACTGTGTGGGGTACTCAGCGAAAGA |
| P16C06 | (GA)44 | CCGAGAGCTTCTCACTGCTTACGAAATCGATCCCTCTTCC |
| P16B10 | (GA)47 | CAAGTGGGCGTGTACAACAGTGTCAGCGCGTCTAGAGAAA |
Primers used in the microsatellite survey
Locus . | Repeat . | Forward primer 5′-3′/reverse primer 5′-3′ . |
|---|---|---|
| C. elegans dimers | ||
| C18F3 | (GT)14 | AAGGGCATGAGGAGAAGTCAACGGGAGCAGCTCTCGTTCTGATTGC |
| F10C2 | (GT)26 | GCTCCACTTTTCAACATCCTTCCTTGTCCCCGCTATTTCGCTC |
| F32B5 | (GT)34 | GCACTTTTCCAACAAAAAAATCCGCCCACACGAGACATGCAATAACTACC |
| W01A6 | (GA)45 | CCCCCGAATCTCATTTCAGTTGTTAGCACCCTTCCGTTTTCACGCC |
| F59D12 | (GT)59 | ACCCCAATGGACTACGTTGTGCACGTGACGCCTTTGGTTCATAG |
| F16A11 | (GT)68 | CTTCTTCTTCTTCCTCCGACCGCAGAGGGGGTGATCTTTATGCC |
| F09F3 | (GT)107 | GCAATCCTACGGTACACCCATACCCGTATTTTGACATTAGGCAGGTGC |
| K06G5 | (GT)142 | AGCTTGAGGAAGAAGGTGGGGTGTTCGTCCATTTGAGCACC |
| T08H4 | (GT)191 | TGGCTCATCACTTGTGTTCAACCCGGCTCCTCTCTGATTTGTCTG |
| C. elegans trimers | ||
| C27D9 | (AAT)12 | AATCGTCGTCGGAACTCAGATAAGGCAAGTGTTGAGGCAAACAGAAG |
| K08D12 | (ACC)12 | GAACGTGCATCAAGTGTAATTCGCTGGCACCCGATATGTGTGAAG |
| E02A10 | (GGA)13 | GTAGCAAGAGCCGAAAGCTGAGTGTCAAGCCTACACCACCACTAGC |
| F47B8 | (GGA)13 | GCTCCTCCAGAATATCCAGCTTCCTATGGTCTGCTCGATCAGTTGTG |
| C24A3 | (TTC)13 | CGCATCTTCTTTTCGGAGCAAGTCATCCTCCTTCCAACACTGCC |
| R01E6 | (TTC)14 | TCCCCTCGTAATTCCCAAAATCCCAAAAGGAGATGCAGAAGAGTAGG |
| F10E7 | (CAA)15 | ACCAAAGTCAACCCGACTACAGCTCGAGTGTGATGCGAACAGTTG |
| C03H5 | (AAT)28 | AACAAAAATGTGGCAGGGAGGGGTTACGGTAGTGGTACTGTAGG |
| F35H10 | (GTA)30 | CGCTCAGAACCATGTTTAAGATTTCCTTTTATGAGTAACTTTGGTGCAGG |
| C. elegans tetramers | ||
| C03B1 | (ATGG)17 | GAATCTGTTCTTCCTTCCTTCGCCGTTCTCCTTTTATCATCAAC |
| C16C8 | (TAGG)19 | TCCTGCCTACCTGCCCATTAGGGCGTGCAGACATAAGATAGG |
| C43B7 | (TAGG)26 | GATGCAGACAAAAATACCGAGAGGGACTAATCACCCACAAACCGAAAC |
| C13B7 | (TAGG)28 | CAAAGTTAGCAAGAACCCAGTTGGTCGCACCTACCTACACCCTTTCTG |
| B0564 | (AAAT)43 | GACCATTAGATGAATTTCCCGAGCAACAGAATTTTTAC |
| D. pulex dimers | ||
| P17D02 | (GA)13 | CCGAATGTCTAACCCACACCTGCACATATTGCCCGTGTAT |
| P17C11 | (GT)15 | ACGGTAGCGGTATTGGAAGATGCAGCGATTCAAATAATGG |
| P17C09 | (GA)15 | GTATGGGCCGGCTAAGGATAGGAATTTGGCGCATCGAG |
| P17C08 | (GA)16 | CCGTCTCCAACCAATTCACTTAAACTTCGGGGACCTTGTG |
| P17B12 | (GA)19 | TCCCGGTTTATCACAGTTCCGCAAAAATCGCAGGTTTCTC |
| P17B11 | (GA)20 | TAACAAGTGGCTGCATCCTGCAGAGGAGGTAACGCAAAGG |
| P17A09 | (GA)26 | CGTCCATCATGACTTGGATCTCACCATGAGTGAAAGCCAGA |
| P17A07 | (GT)27 | AGAGTGAAGGCAGGAAACCACGGAGACGACCAAGGTGTA |
| P17A02 | (GA)30 | TTATTCATGTACGCCCGTTGGCAGTGCTGTCCTGTGAGAC |
| P17A01 | (GA)30 | TCATGTACGCCCGTTGTAAAGCCATGGACGTGACGTATAA |
| P16H07 | (GA)34 | GCACGTGACACATTTGAAGCGTTCAGCACGAAAATGACGA |
| P16H05 | (GA)34 | AGCCGTACCTGTGATGTGTGACGAGCAATACACGATGACG |
| P16H03 | (GA)34 | TGAACGAGGGGAAAATCAAAAACCTGCTCTTCACGACGAC |
| P16G10 | (GA)35 | CTGAGAGCGAGACCGACAGCGACATGGATGATGTTGAGG |
| P16F10 | (GA)36 | GGTGTATCATTTTTGATTGAATTTTCTAGTCGCATTTGCGCCTTT |
| P16F02 | (GA)37 | AGTCTGGAGAGCCAAAGCAGCTCGTCGACCATGAGAACAA |
| P16F01 | (GA)38 | ATTCGAACCGACGGAAAACTCAACGGCAGTGGCAATACTA |
| P16E08 | (GA)38 | CAACGGATGGTGGAATAACCTCCGCGTAGCCTCATAGAAT |
| P16E09 | (GA)38 | CTCGGGAAGGTTTCAGACAGACCCAAAAGAGCGAGGAACT |
| P16C05 | (GA)44 | GATTCGAACCACCCAACTGTGTGGGGTACTCAGCGAAAGA |
| P16C06 | (GA)44 | CCGAGAGCTTCTCACTGCTTACGAAATCGATCCCTCTTCC |
| P16B10 | (GA)47 | CAAGTGGGCGTGTACAACAGTGTCAGCGCGTCTAGAGAAA |
Locus . | Repeat . | Forward primer 5′-3′/reverse primer 5′-3′ . |
|---|---|---|
| C. elegans dimers | ||
| C18F3 | (GT)14 | AAGGGCATGAGGAGAAGTCAACGGGAGCAGCTCTCGTTCTGATTGC |
| F10C2 | (GT)26 | GCTCCACTTTTCAACATCCTTCCTTGTCCCCGCTATTTCGCTC |
| F32B5 | (GT)34 | GCACTTTTCCAACAAAAAAATCCGCCCACACGAGACATGCAATAACTACC |
| W01A6 | (GA)45 | CCCCCGAATCTCATTTCAGTTGTTAGCACCCTTCCGTTTTCACGCC |
| F59D12 | (GT)59 | ACCCCAATGGACTACGTTGTGCACGTGACGCCTTTGGTTCATAG |
| F16A11 | (GT)68 | CTTCTTCTTCTTCCTCCGACCGCAGAGGGGGTGATCTTTATGCC |
| F09F3 | (GT)107 | GCAATCCTACGGTACACCCATACCCGTATTTTGACATTAGGCAGGTGC |
| K06G5 | (GT)142 | AGCTTGAGGAAGAAGGTGGGGTGTTCGTCCATTTGAGCACC |
| T08H4 | (GT)191 | TGGCTCATCACTTGTGTTCAACCCGGCTCCTCTCTGATTTGTCTG |
| C. elegans trimers | ||
| C27D9 | (AAT)12 | AATCGTCGTCGGAACTCAGATAAGGCAAGTGTTGAGGCAAACAGAAG |
| K08D12 | (ACC)12 | GAACGTGCATCAAGTGTAATTCGCTGGCACCCGATATGTGTGAAG |
| E02A10 | (GGA)13 | GTAGCAAGAGCCGAAAGCTGAGTGTCAAGCCTACACCACCACTAGC |
| F47B8 | (GGA)13 | GCTCCTCCAGAATATCCAGCTTCCTATGGTCTGCTCGATCAGTTGTG |
| C24A3 | (TTC)13 | CGCATCTTCTTTTCGGAGCAAGTCATCCTCCTTCCAACACTGCC |
| R01E6 | (TTC)14 | TCCCCTCGTAATTCCCAAAATCCCAAAAGGAGATGCAGAAGAGTAGG |
| F10E7 | (CAA)15 | ACCAAAGTCAACCCGACTACAGCTCGAGTGTGATGCGAACAGTTG |
| C03H5 | (AAT)28 | AACAAAAATGTGGCAGGGAGGGGTTACGGTAGTGGTACTGTAGG |
| F35H10 | (GTA)30 | CGCTCAGAACCATGTTTAAGATTTCCTTTTATGAGTAACTTTGGTGCAGG |
| C. elegans tetramers | ||
| C03B1 | (ATGG)17 | GAATCTGTTCTTCCTTCCTTCGCCGTTCTCCTTTTATCATCAAC |
| C16C8 | (TAGG)19 | TCCTGCCTACCTGCCCATTAGGGCGTGCAGACATAAGATAGG |
| C43B7 | (TAGG)26 | GATGCAGACAAAAATACCGAGAGGGACTAATCACCCACAAACCGAAAC |
| C13B7 | (TAGG)28 | CAAAGTTAGCAAGAACCCAGTTGGTCGCACCTACCTACACCCTTTCTG |
| B0564 | (AAAT)43 | GACCATTAGATGAATTTCCCGAGCAACAGAATTTTTAC |
| D. pulex dimers | ||
| P17D02 | (GA)13 | CCGAATGTCTAACCCACACCTGCACATATTGCCCGTGTAT |
| P17C11 | (GT)15 | ACGGTAGCGGTATTGGAAGATGCAGCGATTCAAATAATGG |
| P17C09 | (GA)15 | GTATGGGCCGGCTAAGGATAGGAATTTGGCGCATCGAG |
| P17C08 | (GA)16 | CCGTCTCCAACCAATTCACTTAAACTTCGGGGACCTTGTG |
| P17B12 | (GA)19 | TCCCGGTTTATCACAGTTCCGCAAAAATCGCAGGTTTCTC |
| P17B11 | (GA)20 | TAACAAGTGGCTGCATCCTGCAGAGGAGGTAACGCAAAGG |
| P17A09 | (GA)26 | CGTCCATCATGACTTGGATCTCACCATGAGTGAAAGCCAGA |
| P17A07 | (GT)27 | AGAGTGAAGGCAGGAAACCACGGAGACGACCAAGGTGTA |
| P17A02 | (GA)30 | TTATTCATGTACGCCCGTTGGCAGTGCTGTCCTGTGAGAC |
| P17A01 | (GA)30 | TCATGTACGCCCGTTGTAAAGCCATGGACGTGACGTATAA |
| P16H07 | (GA)34 | GCACGTGACACATTTGAAGCGTTCAGCACGAAAATGACGA |
| P16H05 | (GA)34 | AGCCGTACCTGTGATGTGTGACGAGCAATACACGATGACG |
| P16H03 | (GA)34 | TGAACGAGGGGAAAATCAAAAACCTGCTCTTCACGACGAC |
| P16G10 | (GA)35 | CTGAGAGCGAGACCGACAGCGACATGGATGATGTTGAGG |
| P16F10 | (GA)36 | GGTGTATCATTTTTGATTGAATTTTCTAGTCGCATTTGCGCCTTT |
| P16F02 | (GA)37 | AGTCTGGAGAGCCAAAGCAGCTCGTCGACCATGAGAACAA |
| P16F01 | (GA)38 | ATTCGAACCGACGGAAAACTCAACGGCAGTGGCAATACTA |
| P16E08 | (GA)38 | CAACGGATGGTGGAATAACCTCCGCGTAGCCTCATAGAAT |
| P16E09 | (GA)38 | CTCGGGAAGGTTTCAGACAGACCCAAAAGAGCGAGGAACT |
| P16C05 | (GA)44 | GATTCGAACCACCCAACTGTGTGGGGTACTCAGCGAAAGA |
| P16C06 | (GA)44 | CCGAGAGCTTCTCACTGCTTACGAAATCGATCCCTCTTCC |
| P16B10 | (GA)47 | CAAGTGGGCGTGTACAACAGTGTCAGCGCGTCTAGAGAAA |
Footnotes
Communicating editor: M. S. McPeek
Acknowledgement
We thank A. Omilian and D. Allen for engaging discussions and helpful suggestions on the manuscript; E. Williams, V. Cloud, E. Choi, and B. Molter for assistance with animal maintenance; and W. Sung for technical assistance. The project was supported by National Institutes of Health grants to M.L. and W.K.T., a National Science Foundation grant to M.L., a Natural Sciences and Engineering Research Council of Canada grant to M.E.A.C., a National Science Foundation Doctoral Dissertation Improvement grant to S.S., and a Howard Hughes Medical Institute Capstone award to A.L.S.
References
Abuin, A., H. Zhang and A. Bradley,
Azaiez, A., E. F. Bouchard, M. Jean and F. J. Belzile,
Ballard, D. J., C. Phillips, G. Wright, C. R. Thacker, C. Robson et al.,
Beck, N. R., M. C. Double and A. Cockburn,
Bichara, M., I. Pinet, S. Schumacher and R. P. Fuchs,
Boyer, J. C., N. A. Yamada, C. N. Roques, S. B. Hatch, K. Riess et al.,
Brinkmann, B., M. Klintschar, F. Neuhuber, J. Huhne and B. Rolf,
Brohede, J., C. R. Primmer, A. Moller and H. Ellegren,
Calabrese, P., and R. Durrett,
Chakraborty, R., M. Kimmel, D. N. Stivers, L. J. Davison and R. Deka,
Chambers, G. K., and E. S. MacAvoy,
Cristescu, M. E., J. K. Colbourne, J. Radivojac and M. Lynch,
Crow, J. F.,
Degtyareva, N. P., P. Greenwell, E. R. Hofmann, M. O. Hengartner, L. Zhang et al.,
Denver, D. R., S. Feinberg, S. Estes, W. K. Thomas and M. Lynch,
Dettman, J. R., and J. W. Taylor,
Di Rienzo, A., P. Donnelly, C. Toomajian, B. Sisk, A. Hill et al.,
Doyle, J. J., and J. L. Doyle,
Drost, J. B., and W. R. Lee,
Drotschmann, K., A. B. Clark, H. T. Tran, M. A. Resnick, D. A. Gordenin et al.,
Dupuy, B. M., M. Stenersen, T. Egeland and B. Olaisen,
Eckert, K. A., A. Mowery and S. E. Hile,
Eisen, J. A.,
Ellegren, H.,
Ellegren, H.,
Ellegren, H.,
Eshleman, J. R., and S. D. Markowitz,
Fernando Vázquez, J., T. Perez, J. Albornoz and A. Dominguez,
Gammie, A. E., N. Erdeniz, J. Beaver, B. Devlin, A. Nanji et al.,
Gragg, H., B. D. Harfe and S. Jinks-Robertson,
Gusmao, L., P. Sanchez-Diz, F. Calafell, P. Martin, C. A. Alonso et al.,
Hanford, M. G., B. C. Rushton, L. C. Gowen and R. A. Farber,
Hawk, J. D., L. Stefanovic, J. C. Boyer, T. D. Petes and R. A. Farber,
Henderson, S. T., and T. D. Petes,
Henke, L., and J. Henke,
Heyer, E., J. Puymirat, P. Dieltjes, E. Bakker and P. de Knijff,
Hile, S. E., G. Yan and K. A. Eckert,
Hohoff, C., K. Dewa, U. Sibbing, K. Hoppe, P. Forster et al.,
Huang, Q. Y., F. H. Xu, H. Shen, H. Y. Deng, Y. J. Liu et al.,
Johnson, R. E., G. K. Kovvali, L. Prakash and S. Prakash,
Kayser, M., L. Roewer, M. Hedman, L. Henke, J. Henke et al.,
Kimble, J., and S. Ward,
Kruglyak, S., R. T. Durrett, M. D. Schug and C. F. Aquadro,
Kunkel, T. A.,
Kunkel, T. A., R. D. Sabatino and R. A. Bambara,
Kunkel, T. A., J. D. Roberts and A. Sugino,
Lai, Y., and F. Sun,
Lee, H. Y., M. J. Park, U. Chung, H. Y. Lee, W. I. Yang et al.,
Legendre, M., N. Pochet, T. Pak and K. J. Verstrepen,
Leonard, J. M., S. R. Bollmann and J. B. Hays,
Leopoldino, A. M., and S. D. Pena,
Lynch, M.,
McConnell, R., S. Middlemist, C. Scala, J. E. Strassmann and D. C. Queller,
Neff, B. D., and M. R. Gross,
Omilian, A. R., M. E. Cristescu, J. L. Dudycha and M. Lynch,
Otto, S. P., and V. Walbot,
Pavlov, Y. I., C. Frahm, S. A. Nick McElhinny, A. Niimi, M. Suzuki et al.,
Renwick, A., L. Davison, H. Spratt, J. P. King and M. Kimmel,
Rose, O., and D. Falush,
Rozen, S., and H. J. Skaletsky,
Sajantila, A., M. Lukka and A. C. Syvanen,
Schlötterer, C.,
Schlötterer, C., R. Ritter, B. Harr and G. Brem,
Schuelke, M.,
Schug, M. D., C. M. Hutter, K. A. Wetterstrand, M. S. Gaudette, T. F. Mackay et al.,
Selkoe, K. A., and R. J. Toonen,
Sia, E. A., R. J. Kokoska, M. Dominska, P. Greenwell and T. D. Petes,
Sia, E. A., M. Dominska, L. Stefanovic and T. D. Petes,
Strand, M., T. A. Prolla, R. M. Liskay and T. D. Petes,
Strand, M., M. C. Earley, G. F. Crouse and T. D. Petes,
Sulston, J., and J. Hodgkin,
Thomas, D. C., J. D. Roberts, R. D. Sabatino, T. W. Myers, C. K. Tan et al.,
Thuillet, A. C., D. Bru, J. David, P. Roumet, S. Santoni et al.,
Tran, H. T., J. D. Keen, M. Kricker, M. A. Resnick and D. A. Gordenin,
Udupa, S. M., and M. Baum,
Vassilieva, L. L., and M. Lynch,
Vigouroux, Y., J. S. Jaqueth, Y. Matsuoka, O. S. Smith, W. D. Beavis et al.,
Viguera, E., D. Canceill and S. D. Ehrlich,
Whittaker, J. C., R. M. Harbord, N. Boxall, I. Mackay, G. Dawson et al.,
Wierdl, M., C. N. Greene, A. Datta, S. Jinks-Robertson and T. D. Petes,
Wierdl, M., M. Dominska and T. D. Petes,
Xu, H., R. Chakraborty and Y. X. Fu,
Xu, X., M. Peng and Z. Fang,
Yamada, N. A., G. A. Smith, A. Castro, C. N. Roques, J. C. Boyer et al.,
Yan, J., Y. Liu, H. Tang, Q. Zhang, Z. Huo et al.,