-
PDF
- Split View
-
Views
-
Cite
Cite
Susan Gabay-Laughnan, Christine D Chase, Victor M Ortega, Liming Zhao, Molecular-Genetic Characterization of CMS-S Restorer-of-Fertility Alleles Identified in Mexican Maize and Teosinte, Genetics, Volume 166, Issue 2, 1 February 2004, Pages 959–970, https://doi.org/10.1093/genetics/166.2.959
Close - Share Icon Share
Abstract
Restorer-of-fertility (Rf) alleles for S-type cytoplasmic male sterility (CMS-S) are prevalent in Mexican races of maize and teosinte. Forty-five Rf alleles from 26 races of maize and 6 Rf alleles from different accessions of teosinte were found to be homozygous viable, consistent with the hypothesis that they are naturally occurring Rf alleles. Mapping and allelism studies were performed to assess the number of genes represented by these 51 alleles. Forty-two of the Rf alleles mapped to the long arm of chromosome 2 (2L), and 5 of these were further mapped to the whp1-rf3 region. The Rf3 restoring allele, found in some U.S. maize inbred lines, cosegregates with internal processing of CMS-S mitochondrial transcripts. Three of the 5 mapped Rf alleles were associated with a similar RNA processing event. Allelism or tight linkage was confirmed between Rf3 and 2 teosinte alleles (Rf K-69-6 and Rf 9477) and between Rf3 and the Cónico Norteño allele Rf C-N (GTO 22). The rf3 region of 2L potentially encodes a complex of linked rf genes. The prevalence of restoring alleles in this chromosomal region, among normal-cytoplasm accessions of Mexican maize and teosinte, supports the conclusion that these alleles have functions in normal mitochondrial gene expression that by chance allow them to restore male fertility in S cytoplasm.
CYTOPLASMIC male sterility (CMS) has been extensively investigated as a model for nuclear-mitochondrial genome interaction in higher plants (reviewed in Hanson and Conde 1985; Schnable and Wise 1998). This maternally inherited failure of a plant to produce functional pollen results from the expression of novel genes within the mitochondria (reviewed in Hanson 1991; Schnable and Wise 1998). CMS genes are “chimeric,” composed of short segments derived from various mitochondrial regions spliced together to give rise to new protein-coding genes. There are nuclear genes capable of suppressing CMS, each unique to a specific type of CMS (reviewed in Schnable and Wise 1998). In maize, these nuclear genes are designated as restorer-of-fertility (rf) genes.
In S-type cytoplasmic male sterility (CMS-S) of maize, the molecular-genetic events that determine phenotype with respect to male fertility take place in the developing male gametophyte or pollen grain (Buchert 1961). The nuclear allele capable of restoring fertility to CMS-S plants is designated Rf3 (Duvick 1965). In this system, the nonrestoring (rf3) allele does not transmit through the pollen. A CMS-S Rf3/rf3 plant produces 50% functional pollen and 50% aborted (collapsed) pollen; all of the functional pollen has the genotype Rf3 (Buchert 1961). Pollen of the genotype rf3 does, however, function in plants with normal cytoplasm.
CMS-S rf3 pollen collapses due to expression of a chimeric mitochondrial gene region designated orf355-orf77 (Zabala et al. 1997). Transcripts of 2.8 and 1.6 kb are associated with the expression of this region. The Rf3 allele cosegregates with internal processing and decreased accumulation of these transcripts (Zabala et al. 1997; Wen and Chase 1999a). The 2.8-kb orf355-orf77 transcript is processed to a 2.1-kb RNA in both sporophytic (leaf or immature ear) and gametophytic (microspore or pollen) tissues carrying an Rf3 allele.
The inbred lines CE1, Ky21, CI21E, and Tr each carry Rf alleles for CMS-S (Duvick 1965; Beckett 1971). These are considered to represent an allelic series at the rf3 locus (Duvick 1957; Laughnan and Gabay 1978), although the possibility does exist that different but tightly linked rf loci are involved. The rf3 locus has been mapped to the long arm of chromosome 2 (2L) proximal to the 2L.80 translocation breakpoint (Laughnan and Gabay 1978) and between the whp1 and bnl17.14 loci (Kamps and Chase 1997).
Naturally occurring Rf alleles for S cytoplasm maize can be found at loci other than rf3. At least 12 inbred lines in the Lancaster Surecrop heterotic group carry Rf alleles that do not map to 2L (reviewed in Chase and Gabay-Laughnan 2003). One of these lines, A619, carries a restoring allele designated RfA that acts by a mechanism distinct from that of Rf3 (E. Kuzmin and K. J. Newton, personal communication). Another Lancaster Surecrop line, Va20, carries a restorer that is not linked to either the whp1 or the bnl17.14 marker on 2L (Kamps and Chase 1996).
In addition to the naturally occurring Rf alleles, >60 restoring alleles for S cytoplasm have been recovered as spontaneous nuclear mutations (Laughnan and Gabay 1978; Gabay-Laughnan et al. 1995; Gabay-Laughnan 1997; reviewed in Chase and Gabay-Laughnan 2003). While 7 of these new restorers do map to 2L (Gabay-Laughnan et al. 1995; reviewed in Chase and Gabay-Laughnan 2003) others map to chromosomes 1, 3, 6, and 8 (Laughnan and Gabay 1978; Gabay-Laughnan 1997). All but one of these mutations have properties making them unsuitable for use in agriculture, mainly homozygous lethality and a deleterious effect on endosperm development. This class of restoring alleles has therefore been given the designation restorer-of-fertility lethal (rfl; Wen et al. 2003). The exceptional restorer is homozygous viable and belongs to a class of restoring alleles given the designation restorer-of-fertility viable (rfv).
As the result of an ancient allotetraploid event (Helentjaris et al. 1988; Ahn and Tanksley 1993; Gaut and Doebley 1997), maize contains many duplicated genes. Of 13 loci shared by chromosomes 2 and 7, most are found on 2L (Helentjaris et al. 1988). There are also loci on 2L that are duplicated on the long arm of chromosome 4 (4L). Two such gene pairs are of special interest here. One pair is white pollen1 (whp1) and color-less2 (c2), duplicated genes that encode chalcone synthase activity in the pathway for anthocyanin pigment (Coe et al. 1981; Coe 1985; Wienand et al. 1986; Franken et al. 1991). The second pair of genes is ns1 and ns2, responsible for the narrow sheath trait (Scanlon et al. 2000). The genes c2 and ns2 map very close to one another on 4L while whp1 and ns1 map close to one another on 2L (Scanlon et al. 2000). Because the rf3 locus also maps close to whp1 (Kamps and Chase 1997), it is possible that a duplicate rf locus may be present on 4L near c2 and ns2.
Gabay-Laughnan (2000) previously identified naturally occurring Rf alleles for CMS-S in Mexican races of maize and teosinte. Twenty-seven out of the 30 races of maize examined have accessions that carry Rf alleles. In the teosinte Zea mays ssp. parviglumis, the progenitor of maize (Matsuoka et al. 2002), four out of five accessions tested contained plants carrying Rf alleles for S cytoplasm. In Z. mays ssp. mexicana, the apparent source of S cytoplasm (reviewed in Gabay-Laughnan 2000), three out of seven accessions tested contained plants carrying Rf alleles. Forty-five maize Rf alleles and six teosinte Rf alleles, representing an undetermined number of genetic loci, were the subject of this study.
MATERIALS AND METHODS
Plant materials: The 26 races of Mexican maize contributing Rf alleles to this study are listed in Table 1. Rf alleles are designated by the three-letter abbreviation for the race of origin (see Table 1), e.g., Rf APA (see Doebley et al. 1985). Since multiple accessions of some races were included, this designation may also include the accession number, e.g., Rf C-N (GTO 22). These races of maize were obtained from the International Maize and Wheat Improvement Center (CIMMYT) and from M. M. Goodman, North Carolina State University, Raleigh. Multiple accessions of the two races known to have accessions with S cytoplasm, Cónico Norteño and Celaya, as well as races considered by Wellhausen et al. (1952) to be involved in their ancestry were employed (Gabay-Laughnan 2000). All Rf alleles used in this study have been introgressed into CMS-S versions of nonrestoring U.S. maize.
Races and accessions of Mexican maize used in this study
| Race . | Abbreviation . | Accession no.a . |
|---|---|---|
| Apachito | APA | CHH 159 |
| Arrocillo Amarillo | A-A | PUE 91 |
| Azul | AZU | CHH 133 |
| Bolita | BOL | OAX 28 |
| Celaya | CEL | GTO 20, 69, 75, 84, 88 |
| Chapalote | CHP | SIN 2 |
| Cristalino de Chihuahua | C-C | CHH 8 |
| Comiteco | COM | CHS 38 |
| Cónico | CON | MEX 72; PUE 48, 108 |
| Cónico Norteño | C-N | GTO 16, 22,b 34, 50, 60; QRO 1, 4, 5; ZAC 12 |
| Gordo | GOR | CHH 146 |
| Harinoso de Ocho | H-O | NAY 24; SON 102 |
| Jala | JAL | NAY 6 |
| Maíz Dulce | M-D | JAL 78 |
| Nal-Tel | N-T | YUC 7 |
| Olotillo | OLO | CHS 56 |
| Pepitilla | PEP | MOR 17 |
| Reventador | REV | NAY 15 |
| Tablilla de Ocho | T-O | NAY 193 |
| Tabloncillo | TAB | JAL 42, 63, 102 |
| Tehua | TEH | CHS 29 |
| Tepecintle | TEP | CHS 26, 76, 225 |
| Tuxpeño | TUX | VER 39 |
| Vandeño | VAN | CHS 25 |
| Zapalote Chico | Z-C | OAX 48 |
| Zapalote Grande | Z-G | CHS 104 |
| Race . | Abbreviation . | Accession no.a . |
|---|---|---|
| Apachito | APA | CHH 159 |
| Arrocillo Amarillo | A-A | PUE 91 |
| Azul | AZU | CHH 133 |
| Bolita | BOL | OAX 28 |
| Celaya | CEL | GTO 20, 69, 75, 84, 88 |
| Chapalote | CHP | SIN 2 |
| Cristalino de Chihuahua | C-C | CHH 8 |
| Comiteco | COM | CHS 38 |
| Cónico | CON | MEX 72; PUE 48, 108 |
| Cónico Norteño | C-N | GTO 16, 22,b 34, 50, 60; QRO 1, 4, 5; ZAC 12 |
| Gordo | GOR | CHH 146 |
| Harinoso de Ocho | H-O | NAY 24; SON 102 |
| Jala | JAL | NAY 6 |
| Maíz Dulce | M-D | JAL 78 |
| Nal-Tel | N-T | YUC 7 |
| Olotillo | OLO | CHS 56 |
| Pepitilla | PEP | MOR 17 |
| Reventador | REV | NAY 15 |
| Tablilla de Ocho | T-O | NAY 193 |
| Tabloncillo | TAB | JAL 42, 63, 102 |
| Tehua | TEH | CHS 29 |
| Tepecintle | TEP | CHS 26, 76, 225 |
| Tuxpeño | TUX | VER 39 |
| Vandeño | VAN | CHS 25 |
| Zapalote Chico | Z-C | OAX 48 |
| Zapalote Grande | Z-G | CHS 104 |
The three-letter abbreviation represents the state of Mexico in which the collection was made. CHH, Chihuahua; PUE, Puebla; OAX, Oaxaca; GTO, Guanajuato; SIN, Sinaloa; CMS, Chiapas; MEX, Mexico State; QRO, Querétaro; NAY, Nayarit; SON, Sonora; JAL, Jalisco; YUC, Yucatan; MOR, Morelos; VER, Veracruz; ZAC, Zacatecas.
Some plants in this accession carry CMS-S (Weissinger et al. 1983).
Races and accessions of Mexican maize used in this study
| Race . | Abbreviation . | Accession no.a . |
|---|---|---|
| Apachito | APA | CHH 159 |
| Arrocillo Amarillo | A-A | PUE 91 |
| Azul | AZU | CHH 133 |
| Bolita | BOL | OAX 28 |
| Celaya | CEL | GTO 20, 69, 75, 84, 88 |
| Chapalote | CHP | SIN 2 |
| Cristalino de Chihuahua | C-C | CHH 8 |
| Comiteco | COM | CHS 38 |
| Cónico | CON | MEX 72; PUE 48, 108 |
| Cónico Norteño | C-N | GTO 16, 22,b 34, 50, 60; QRO 1, 4, 5; ZAC 12 |
| Gordo | GOR | CHH 146 |
| Harinoso de Ocho | H-O | NAY 24; SON 102 |
| Jala | JAL | NAY 6 |
| Maíz Dulce | M-D | JAL 78 |
| Nal-Tel | N-T | YUC 7 |
| Olotillo | OLO | CHS 56 |
| Pepitilla | PEP | MOR 17 |
| Reventador | REV | NAY 15 |
| Tablilla de Ocho | T-O | NAY 193 |
| Tabloncillo | TAB | JAL 42, 63, 102 |
| Tehua | TEH | CHS 29 |
| Tepecintle | TEP | CHS 26, 76, 225 |
| Tuxpeño | TUX | VER 39 |
| Vandeño | VAN | CHS 25 |
| Zapalote Chico | Z-C | OAX 48 |
| Zapalote Grande | Z-G | CHS 104 |
| Race . | Abbreviation . | Accession no.a . |
|---|---|---|
| Apachito | APA | CHH 159 |
| Arrocillo Amarillo | A-A | PUE 91 |
| Azul | AZU | CHH 133 |
| Bolita | BOL | OAX 28 |
| Celaya | CEL | GTO 20, 69, 75, 84, 88 |
| Chapalote | CHP | SIN 2 |
| Cristalino de Chihuahua | C-C | CHH 8 |
| Comiteco | COM | CHS 38 |
| Cónico | CON | MEX 72; PUE 48, 108 |
| Cónico Norteño | C-N | GTO 16, 22,b 34, 50, 60; QRO 1, 4, 5; ZAC 12 |
| Gordo | GOR | CHH 146 |
| Harinoso de Ocho | H-O | NAY 24; SON 102 |
| Jala | JAL | NAY 6 |
| Maíz Dulce | M-D | JAL 78 |
| Nal-Tel | N-T | YUC 7 |
| Olotillo | OLO | CHS 56 |
| Pepitilla | PEP | MOR 17 |
| Reventador | REV | NAY 15 |
| Tablilla de Ocho | T-O | NAY 193 |
| Tabloncillo | TAB | JAL 42, 63, 102 |
| Tehua | TEH | CHS 29 |
| Tepecintle | TEP | CHS 26, 76, 225 |
| Tuxpeño | TUX | VER 39 |
| Vandeño | VAN | CHS 25 |
| Zapalote Chico | Z-C | OAX 48 |
| Zapalote Grande | Z-G | CHS 104 |
The three-letter abbreviation represents the state of Mexico in which the collection was made. CHH, Chihuahua; PUE, Puebla; OAX, Oaxaca; GTO, Guanajuato; SIN, Sinaloa; CMS, Chiapas; MEX, Mexico State; QRO, Querétaro; NAY, Nayarit; SON, Sonora; JAL, Jalisco; YUC, Yucatan; MOR, Morelos; VER, Veracruz; ZAC, Zacatecas.
Some plants in this accession carry CMS-S (Weissinger et al. 1983).
The six accessions of teosinte utilized in these studies (Table 2) were provided by CIMMYT and by J. F. Doebley, University of Wisconsin, Madison (then at the University of Minnesota, St. Paul). All were previously shown to carry Rf alleles for CMS-S (Gabay-Laughnan 2000). These Rf alleles are designated by their accession numbers (see Table 2), e.g., Rf K-69-4. These Rf alleles have also been introgressed into CMS-S versions of nonrestoring U.S. maize.
Four U.S. inbred lines of maize known to be homozygous for the Rf3 allele were also employed. CE1, a Pioneer Hi-Bred inbred line, was a gift from D. N. Duvick (then at Pioneer Hi-Bred International, Johnston, Iowa). The public inbred lines Ky21, CI21E, and Tr were obtained from J. B. Beckett (University of Missouri, Columbia). For the molecular studies, the Rf3 alleles from each of these inbred lines were introgressed into Mo17, a nonrestoring inbred line, each having been crossed with Mo17 at least eight times.
Teosinte collections used in this study
| Zea mays subspeciesa . | Raceb . | Accession no. . |
|---|---|---|
| mexicana | Central Plateau | K-69-4 |
| mexicana | Central Plateau | K-69-6c |
| parviglumis | Jalisco | MO46 |
| parviglumis | Central Balsas | MO106 |
| parviglumis | Central Balsas | 9477 |
| parviglumis | Southern Guerrero | MO63 |
| Zea mays subspeciesa . | Raceb . | Accession no. . |
|---|---|---|
| mexicana | Central Plateau | K-69-4 |
| mexicana | Central Plateau | K-69-6c |
| parviglumis | Jalisco | MO46 |
| parviglumis | Central Balsas | MO106 |
| parviglumis | Central Balsas | 9477 |
| parviglumis | Southern Guerrero | MO63 |
Teosinte collections used in this study
| Zea mays subspeciesa . | Raceb . | Accession no. . |
|---|---|---|
| mexicana | Central Plateau | K-69-4 |
| mexicana | Central Plateau | K-69-6c |
| parviglumis | Jalisco | MO46 |
| parviglumis | Central Balsas | MO106 |
| parviglumis | Central Balsas | 9477 |
| parviglumis | Southern Guerrero | MO63 |
| Zea mays subspeciesa . | Raceb . | Accession no. . |
|---|---|---|
| mexicana | Central Plateau | K-69-4 |
| mexicana | Central Plateau | K-69-6c |
| parviglumis | Jalisco | MO46 |
| parviglumis | Central Balsas | MO106 |
| parviglumis | Central Balsas | 9477 |
| parviglumis | Southern Guerrero | MO63 |
Nomenclature: Loci or genes are indicated in lowercase (e.g., the rf3 locus), in compliance with the rules of maize genetics nomenclature. Alleles at a locus are also indicated in italics, with the first letter capitalized for dominant alleles (e.g., the Rf3 allele). We have indicated whether we are referring to a gene (locus) or an allele in cases where the nomenclature is ambiguous. Dominance relationships of gametophytic restoring and nonrestoring alleles cannot be directly assessed in haploid pollen (Kamps et al. 1996). However, in keeping with conventional usage, restorer-of-fertility genes (loci) are designated by rf, restoring alleles by Rf, and nonrestoring alleles by rf. The Rf alleles characterized in this study have been assigned Rf symbols indicating the race and/or accession number in which they were identified (see Plant materials) until their relationships to Rf3 have been clearly defined. Plant mitochondrial genes are indicated in lowercase italics (e.g., the atp9 gene).
The normal male-fertile cytoplasm of maize is given the designation N, while S male-sterile cytoplasm is designated CMS-S or, briefly, S. Since there have been numerous independent “discoveries” of S-type cytoplasms, subgroups or subtypes of CMS-S have been given unique letter designations (Beckett 1971; Sisco et al. 1985; reviewed in Gabay-Laughnan et al. 1995). The subtypes of CMS-S used in this study are S, I, R, and VG.
Analysis of homozygous viability: Newly identified Rf alleles were tested for homozygous viability by self-pollinating CMS-S Rf/rf plants. If the Rf allele is homozygous lethal, a semisterile seed set (one-half of the kernels missing) results. If an Rf allele is homozygous viable, the self-pollinated ear will have a normal set. Homozygous viability was confirmed by pollen phenotypes of plants grown from kernels on the self-pollinated ears. In the case of homozygous viability, one-half of the plants (Rf/rf) are expected to exhibit 50% aborted pollen and one-half of the plants (Rf/Rf) are expected to have 100% starch-filled pollen. All plants will exhibit 50% aborted pollen if the restorer is homozygous lethal.
Placement of Rf alleles to chromosomes: The waxy1 (wx1)-marked reciprocal translocation T2-9d was used to test Rf alleles for linkage with chromosome 2L. Linkage of Rf alleles with chromosome 4L was tested using the wx1-marked reciprocal translocation T4-9(5657). These translocations were obtained from the Maize Genetics Cooperation Stock Center, Urbana, Illinois. The translocation lines employed in our studies do not carry Rf alleles for S cytoplasm; i.e., they are rf/rf. The use of wx1-marked reciprocal translocations to ascertain the chromosome location of Rf alleles was previously described (Laughnan and Gabay 1978; Laughnan and Gabay-Laughnan 1994). Briefly, CMS-S plants carrying an unplaced Rf allele are crossed with rf pollen from plants carrying a wx1-marked translocation. CMS-S Rf/rf plants exhibit 50% aborted pollen grains due to gametophytic fertility restoration (Buchert 1961). Maize plants heterozygous for a reciprocal translocation also exhibit 50% pollen abortion (Patterson 1994). CMS-S plants heterozygous for both an Rf allele and a reciprocal translocation exhibit 75% aborted pollen. These are the plants of interest and their pollen is crossed onto wx1/wx1 tester plants. The proportion of waxy kernels on the resulting ear is a function of the recombination between Rf and wx. χ2 values for independence of rf and wx were calculated using Yates’ correction for continuity (Yates 1934) since there is only 1 d.f.
Restriction fragment length polymorphism analysis: Backcross (BC) populations segregating for CMS-S Rf/rf and CMS-S rf/rf plants were generated for restriction fragment length polymorphism (RFLP) analysis. CMS-S Mo17 rf/rf plants were fertilized with pollen carrying Rf alleles from Mexican maize or teosinte. The resulting CMS-S Rf/rf plants were fertilized with N-cytoplasm Mo17 rf pollen. In some cases, an additional generation of backcrossing was performed with N-cytoplasm Mo17 rf pollen. Populations of 50 plants were grown and analyzed for male fertility on the basis of visible pollen shed. The χ2 P for a 1:1 segregation of CMS-S Rf/rf and CMS-S rf/rf plants was at least 0.3 in each of the populations. The first 20 plants from each population row were analyzed for RFLPs at the whp1 locus as described by Kamps et al. (1996). The whp1 probe was obtained from U. Wienand (University of Hamburg).
Tests of allelism: Once two restoring alleles are located to the same chromosome, two approaches determine whether they are allelic. Both involve producing a CMS-S F1 plant containing the two Rf alleles. The first approach involves pollen examination. Due to the gametophytic nature of CMS-S restoration (Buchert 1961), a plant homozygous for an Rf allele exhibits 100% normal pollen, i.e., no pollen abortion. In contrast, plants heterozygous for restoring and nonrestoring alleles at two unlinked loci exhibit 25% pollen abortion. If the two restorers being tested are closely linked, but not allelic, the percentage of pollen abortion will be <25% and approach 0% in the case of very tightly linked rf loci. Pollen abortion is determined by examining fresh pollen with a pocket field microscope (×40). Since there may be a low level of background pollen abortion, a second approach is more accurate. However, this approach is labor intensive, requires extensive field space, and is therefore not always feasible. F1 plants exhibiting 100%, or nearly 100%, normal pollen are testcrossed by pollen from a normal-cytoplasm rf/rf tester, and the resulting kernels are planted. Fertility and sterility are scored by tassel examination at maturity. If the two Rf alleles being tested are at the same locus, the resulting progeny will all be fertile. If the two restorers are not allelic, the percentage of sterile plants among the progeny is a function of the distance between the two rf genes. Note that two nonallelic restorers may be so closely linked that a crossover between them may not be observed.
RNA blot hybridization: Backcross populations used for the RFLP analysis (described above) were also used to examine the association of Rf alleles with processing of orf355-orf77 mitochondrial transcripts. Immature ears were harvested from individual plants, frozen in liquid nitrogen, and stored at -80°. Total RNA was extracted as described by Wen and Chase (1999b) from one Rf/rf and one rf/rf plant in each segregating population. RNA samples of 15 μg were denatured by a 15-min incubation at 65° in 1× MEN buffer (20 mm MOPS, 5 mm sodium acetate, and 2 mm EDTA, pH 7.0) containing 40% v/v formamide, 12% v/v formaldehyde, and 10 μg/ml ethidium bromide. RNA was fractionated by electrophoresis through 1.5% agarose gels made with 1× MEN buffer containing 16% v/v formaldehyde and run in 1× MEN buffer. RNAs were transferred to GeneScreen Plus nylon membrane (Perkin-Elmer Life Sciences). 32P-labeled probes corresponding to orf355-orf77 or maize mitochondrial atp9 coding sequences were prepared as described by Wen and Chase (1999b). Membranes were prehybridized for 1 hr and hybridized for 18 hr at 65° in a solution containing 7% w/v SDS, 1% w/v BSA, 1 mm EDTA, and 0.5 m NaHPO4, pH 7.2. Hybridized membranes were washed twice for 15 min at 65° in 0.1% w/v SDS, 150 mm NaCl, and 15 mm sodium citrate, followed by a final 15-min wash at 65° in 0.1% w/v SDS, 15 mm NaCl, and 1.5 mm sodium citrate. Hybridized transcripts were visualized by autoradiography.
RESULTS
All newly identified Rf alleles are homozygous viable: Homozygous viability was previously reported for 47 Rf alleles occurring in 24 races of maize from Mexico and one race from Guatemala (Gabay-Laughnan 2000). All produced normal seed set when self-pollinated. At the time of that report, homozygous viability had not been confirmed by pollen analysis for 5 of the Rf alleles from Mexican maize or for the Rf allele from Guatemalan maize. Pollen analysis has now confirmed the homozygous viability of Rf CEL (GTO 88), Rf CHP, Rf CON (PUE 108), Rf C-N (AGS7), Rf TUX, and the Rf allele from the Guatamalan race Oloton. In addition, Rf alleles from the late-maturing races Comiteco (Rf COM) and Tehua (Rf TEH), identified since the previous report, were determined to be homozygous viable by both seed set and pollen criteria. No deleterious effects on endosperm development were observed on ears of self-pollinated plants carrying any of these Rf alleles. Furthermore, Rf alleles from ssp. mexicana, Rf K-69-4 and Rf K-69-6, and Rf alleles from ssp. parviglumis, Rf MO46, Rf MO106, Rf 9477, and Rf MO63, were homozygous viable by seed set and pollen criteria. In all cases, the Rf/Rf/Rf endosperm developed normally.
Most newly identified Rf alleles map to chromosome 2L: Because the rf3 locus maps to 2L, the translocation wx1 T2-9d with breakpoints at 2L.83 and 9L.27 was used to test the newly identified Rf alleles for linkage to chromosome 2 (see materials and methods). Previous mapping studies with Rf3 employed chromosome 2 translocations other than T2-9d (Laughnan and Gabay 1978). For this reason, we also mapped the Rf3 alleles from CE1, Ky21, CI21E, and Tr with wx1 T2-9d to better evaluate the data for the newly identified Rf alleles. Representative testcross data for placement of Rf3 alleles to chromosome 2L gave recombination values between the rf3 and wx1 loci ranging from 14.5 to 23.6% (Table 3). In all cases, placement of Rf3 alleles on 2L was supported. Because translocations alter the normal patterns of chromosome pairing and recombination (Patterson 1994), this technique is useful only for placing genes to a chromosome arm.
Representative testcross data for placement of Rf alleles from Mexican races of maize on 2L are presented in Table 4. Rf alleles from 45 accessions of 26 races were crossed with wx1 T2-9d. Testcrosses indicate that all but 7 alleles—Rf A-A, Rf C-N (GTO 34), Rf M-D, Rf N-T, Rf OLO, Rf TEP (CHS76), and Rf TUX—map to chromosome 2L. These 45 alleles therefore represent at least two different rf loci.
Representative testcross data for placement of Rf alleles from teosinte are presented in Table 5. Rf alleles from six sources were crossed with wx1 T2-9d. Test-crosses indicate that alleles Rf K-69-6, Rf MO46, Rf MO-106, and Rf 9477 map conclusively to chromosome 2L. The Rf K-69-4 allele is clearly not on chromosome 2L, and the data for the Rf MO63 restorer are inconclusive. Rf alleles isolated from teosinte therefore also represent at least two genetic loci.
Five newly identified Rf alleles are linked to whp1: The Rf3 allele from inbred lines Ky21 and CE1 is linked to the whp1 locus on 2L (Kamps and Chase 1997). One or more Rf alleles from Mexican maize and teosinte may therefore be allelic to Rf3. An RFLP analysis was used to test four maize alleles—Rf CEL (GTO 20), Rf CHP, Rf C-N (GTO 22), and Rf REV—and one teosinte allele, Rf K-69-6, for linkage with the whp1 locus (Table 6). Backcross populations segregating 1:1 for CMS-S Rf/rf and CMS-S rf/rf genotypes were examined for segregation of whp1 alleles from the Rf source and the recurrent Mo17 rf/rf parent. Linkage of whp1 with each of the tested rf loci was demonstrated by biased representation of the whp-Rf source allele in the CMS-S Rf/rf progeny, and χ2 analysis rejected independence of rf and whp1 loci at P < 0.01 in each population.
Rf alleles from teosinte are allelic or closely linked to Rf3: Two of the Rf alleles isolated from teosinte, Rf 9477 and Rf K-69-6, were tested for allelism with Rf3. Plants carrying both the Rf3 allele and the teosinte Rf allele were identified by pollen analysis (see materials and methods). Since we already know that both restorers map to 2L, we expect the plants to fall within a range of 0-25% pollen abortion, with a lower percentage of pollen abortion reflecting closer linkage of the two rf loci.
Plants of the genotype CMS-S (or CMS-I) Rf 9477/rf were crossed as female parents by the inbred line Tr (Rf3/Rf3) as male parent. Pollen of the resulting progeny plants was checked for level of abortion. The progeny included 4 plants exhibiting 50% pollen abortion (genotype Rf3/rf) and 15 plants exhibiting 0% pollen abortion (genotype Rf 9477/Rf3). Plants of the genotype CMS-S Rf K-69-6/rf were crossed as female parents by both the inbred line CE1 (Rf3/Rf3) and the inbred line Tr (Rf3/Rf3). We observed 17 progeny exhibiting 50% pollen abortion (genotype Rf3/rf), 5 plants with 0% pollen abortion (Rf K-69-6/Rf3), and 7 plants with >0% but <10% pollen abortion (also Rf K-69-6/Rf3). In a reciprocal cross, CMS-VG plants homozygous for Rf3 CE1 were crossed as female parents by plants of the genotype CMS-S (or CMS-R) Rf K-69-6/rf. Twenty-three progeny plants exhibited no pollen abortion and 9 exhibited a low percentage (<10%) of pollen abortion. As expected, no plants exhibiting 50% pollen abortion were observed. These results indicate that Rf 9477 and Rf K-69-6 are allelic or closely linked to the rf3 locus.
Representative testcross data for placement of Rf3 on chromosome 2L
| Rf3 sourcea . | Wx . | wxb . | % rf-wx recombination . | χ2c . | P value . |
|---|---|---|---|---|---|
| CE1 | 382 | 118 | 23.6 | 138.3 | <0.001 |
| 158 | 42 | 21.0 | 66.1 | <0.001 | |
| Ky21 | 164 | 36 | 18.0 | 80.6 | <0.001 |
| CI21E | 170 | 30 | 15.0 | 96.6 | <0.001 |
| 171 | 29 | 14.5 | 99.4 | <0.001 | |
| Tr | 307 | 93 | 23.3 | 113.4 | <0.001 |
| Rf3 sourcea . | Wx . | wxb . | % rf-wx recombination . | χ2c . | P value . |
|---|---|---|---|---|---|
| CE1 | 382 | 118 | 23.6 | 138.3 | <0.001 |
| 158 | 42 | 21.0 | 66.1 | <0.001 | |
| Ky21 | 164 | 36 | 18.0 | 80.6 | <0.001 |
| CI21E | 170 | 30 | 15.0 | 96.6 | <0.001 |
| 171 | 29 | 14.5 | 99.4 | <0.001 | |
| Tr | 307 | 93 | 23.3 | 113.4 | <0.001 |
Inbred line of genotype Rf3/Rf3.
Kernels were scored from contiguous regions on individual ears.
χ2 for independence of rf and wx loci calculated using Yates’ correction for continuity (Yates 1934).
Representative testcross data for placement of Rf3 on chromosome 2L
| Rf3 sourcea . | Wx . | wxb . | % rf-wx recombination . | χ2c . | P value . |
|---|---|---|---|---|---|
| CE1 | 382 | 118 | 23.6 | 138.3 | <0.001 |
| 158 | 42 | 21.0 | 66.1 | <0.001 | |
| Ky21 | 164 | 36 | 18.0 | 80.6 | <0.001 |
| CI21E | 170 | 30 | 15.0 | 96.6 | <0.001 |
| 171 | 29 | 14.5 | 99.4 | <0.001 | |
| Tr | 307 | 93 | 23.3 | 113.4 | <0.001 |
| Rf3 sourcea . | Wx . | wxb . | % rf-wx recombination . | χ2c . | P value . |
|---|---|---|---|---|---|
| CE1 | 382 | 118 | 23.6 | 138.3 | <0.001 |
| 158 | 42 | 21.0 | 66.1 | <0.001 | |
| Ky21 | 164 | 36 | 18.0 | 80.6 | <0.001 |
| CI21E | 170 | 30 | 15.0 | 96.6 | <0.001 |
| 171 | 29 | 14.5 | 99.4 | <0.001 | |
| Tr | 307 | 93 | 23.3 | 113.4 | <0.001 |
Inbred line of genotype Rf3/Rf3.
Kernels were scored from contiguous regions on individual ears.
χ2 for independence of rf and wx loci calculated using Yates’ correction for continuity (Yates 1934).
The second approach (see materials and methods) was undertaken to test more accurately allelism between Rf K-69-6 and Rf3. CMS-S Rf K-69-6/Rf3 plants, i.e., those exhibiting no or low levels of pollen abortion, were pollinated by N-cytoplasm rf/rf plants, either Mo17 or Oh51A. The resulting progeny were scored for the ratio of fertile to sterile plants. The Rf K-69-6 allele tested here is the allele that was linked to whp1 through the use of RFLP markers (Table 6). The data from seven populations carrying Rf K-69-6 and either Rf3 CE1 or Rf3 Tr are presented in Table 7. Each population was planted from a single testcross ear. Only 1 plant out of 2064 appeared to be rf/rf. This plant exhibited some thin (sterile) anthers, rating 2 on a scale of 1-5 where 1 is male sterile and 5 is fully fertile (see Gabay-Laughnan 2001). If this plant was truly of the genotype rf/rf, then Rf K-69-6 and Rf3 are very tightly linked but not allelic.
Rf C-N (GTO22) is allelic or closely linked to Rf3: The Rf C-N (GTO22) allele, linked to the RFLP marker whp1 (Table 6), was combined with Rf3 CE1 and with Rf3 Tr in tests for allelism. A plant of the genotype CMS-R Rf C-N (GTO 22)/rf was crossed as female parent with pollen from a plant of the genotype CMS-R Rf3 CE1/rf. Fifteen progeny were checked for pollen abortion. Due to the gametophytic nature of CMS-S restoration (Buchert 1961), all progeny received the Rf3 gene from the pollen parent. Seven plants exhibited 50% pollen abortion (Rf3/rf) while four plants exhibited no pollen abortion [Rf C-N (GTO 22)/Rf3] and two plants exhibited >0% but <10% pollen abortion [also Rf C-N (GTO 22)/Rf3].
Four of the putative Rf C-N (GTO 22)/Rf3 plants were crossed with pollen from the inbred line Oh51A (rf/rf). The resulting progeny were scored for the ratio of fertile to sterile plants. The data from the four populations, each planted from an individual ear, are presented in Table 8. All 512 of the resulting plants were male fertile.
A similar strategy was carried out to test allelism between Rf C-N (GTO 22) and Rf3 Tr. A CMS-R Rf C-N (GTO22)/rf plant was crossed with pollen from a CMS-R Rf3 Tr/rf plant. Five putative Rf C-N (GTO 22)/Rf3 Tr progeny were crossed with pollen from the inbred line Oh51A (rf/rf). The ratio of fertile to sterile plants was scored in the resulting progeny. The data from these five populations, each planted from an individual ear, are presented in Table 8. All 543 plants were male fertile.
There were no male-sterile plants out of a combined total of 1055 plants scored for fertility in the tests for allelism of Rf C-N (GTO 22) with Rf3 (Table 8). This would indicate that Rf C-N (GTO 22) is allelic to rf3. However, it is always possible that additional progeny would contain a sterile (rf/rf) recombinant. To err on the side of caution, we cannot discount the possibility that the two restorers are tightly linked but not allelic to each other.
Processing of mitochondrial RNA transcripts encoding CMS-S: Since most Mexican Rf alleles map to 2L and some are allelic or closely linked to Rf3, we performed RNA analyses to determine if the Mexican Rf alleles work by the same mechanism as Rf3. From populations segregating for Rf alleles linked to whp1 (Table 6), selected plants were examined for association of fertility restoration and mitochondrial RNA processing events.
Representative testcross data for placement of maize Rf alleles on chromosome 2L
| Rf sourcea . | Accession no.b . | Wx . | wxc . | % rf-wx recombination . | χ2d . | P value . |
|---|---|---|---|---|---|---|
| APA | CHH 159 | 177 | 23 | 11.5 | 117.0 | <0.001 |
| A-A | PUE 91 | 100 | 100 | 50.0 | 0.0 | >0.99 |
| AZU | CHH 133 | 182 | 18 | 9.0 | 132.8 | <0.001 |
| BOL | OAX 28 | 174 | 26 | 13.0 | 108.0 | <0.001 |
| CEL | GTO 20 | 170 | 30 | 15.0 | 96.6 | <0.001 |
| CEL | GTO 69 | 156 | 44 | 22.0 | 61.6 | <0.001 |
| CEL | GTO 75 | 172 | 28 | 14.0 | 102.3 | <0.001 |
| CEL | GTO 84 | 184 | 16 | 8.0 | 139.4 | <0.001 |
| CEL | GTO 88 | 135 | 65 | 32.5 | 23.8 | <0.001 |
| CHP | SIN 2 | 176 | 24 | 12.0 | 114.0 | <0.001 |
| C-C | CHH 8 | 144 | 56 | 28.0 | 37.8 | <0.001 |
| COM | CHS 38 | 125 | 75 | 37.5 | 12.0 | <0.001 |
| CON | MEX 72 | 184 | 16 | 8.0 | 139.4 | <0.001 |
| CON | PUE 48 | 170 | 30 | 15.0 | 96.6 | <0.001 |
| CON | PUE 108 | 170 | 30 | 15.0 | 96.6 | <0.001 |
| C-N | GTO 16 | 171 | 29 | 14.5 | 99.4 | <0.001 |
| C-N | GTO 22 | 163 | 37 | 18.5 | 78.1 | <0.001 |
| C-N | GTO 34 | 110 | 90 | 45.0 | 1.8 | 0.2-0.1 |
| C-N | GTO 50 | 159 | 41 | 20.5 | 68.4 | <0.001 |
| C-N | GTO 60 | 179 | 21 | 10.5 | 123.2 | <0.001 |
| C-N | QRO 1 | 161 | 39 | 19.5 | 73.2 | <0.001 |
| C-N | QRO 4 | 166 | 34 | 17.0 | 85.8 | <0.001 |
| C-N | QRO 5 | 158 | 42 | 21.0 | 66.1 | <0.001 |
| C-N | ZAC 12 | 157 | 43 | 21.5 | 63.8 | <0.001 |
| GOR | CHH 146 | 188 | 12 | 6.0 | 153.1 | <0.001 |
| H-O | NAY 24 | 132 | 68 | 34.0 | 19.8 | <0.001 |
| H-O | SON 102 | 154 | 46 | 23.0 | 57.2 | <0.001 |
| JAL | NAY 6 | 171 | 26 | 14.5 | 99.4 | <0.001 |
| M-D | JAL 78 | 95 | 105 | 52.5 | 0.4 | 0.8-0.7 |
| N-T | YUC 7 | 93 | 107 | 53.5 | 0.8 | 0.8-0.7 |
| OLO | CHS 56 | 107 | 93 | 46.5 | 0.8 | 0.8-0.7 |
| PEP | MOR 17 | 161 | 39 | 19.5 | 73.2 | <0.001 |
| REV | NAY 15 | 181 | 19 | 9.5 | 129.6 | <0.001 |
| T-O | NAY 193 | 147 | 53 | 26.5 | 43.2 | <0.001 |
| TAB | JAL 42 | 182 | 18 | 9.0 | 132.8 | <0.001 |
| TAB | JAL 63 | 151 | 49 | 24.5 | 51.0 | <0.001 |
| TAB | JAL 102 | 170 | 24 | 12.0 | 108.4 | <0.001 |
| TEH | CHS 29 | 138 | 62 | 31.0 | 28.1 | <0.001 |
| TEP | CHS 26 | 181 | 19 | 9.5 | 129.6 | <0.001 |
| TEP | CHS 76 | 114 | 86 | 43.0 | 3.6 | 0.1-0.05 |
| TEP | CHS 225 | 179 | 21 | 10.5 | 123.2 | <0.001 |
| TUX | VER 39 | 87 | 113 | 56.5 | 3.1 | 0.1-0.05 |
| VAN | CHS 25 | 150 | 50 | 25.0 | 49.0 | <0.001 |
| Z-C | OAX 48 | 165 | 35 | 17.5 | 83.2 | <0.001 |
| Z-G | CHS 104 | 117 | 83 | 41.5 | 5.4 | 0.02 |
| Rf sourcea . | Accession no.b . | Wx . | wxc . | % rf-wx recombination . | χ2d . | P value . |
|---|---|---|---|---|---|---|
| APA | CHH 159 | 177 | 23 | 11.5 | 117.0 | <0.001 |
| A-A | PUE 91 | 100 | 100 | 50.0 | 0.0 | >0.99 |
| AZU | CHH 133 | 182 | 18 | 9.0 | 132.8 | <0.001 |
| BOL | OAX 28 | 174 | 26 | 13.0 | 108.0 | <0.001 |
| CEL | GTO 20 | 170 | 30 | 15.0 | 96.6 | <0.001 |
| CEL | GTO 69 | 156 | 44 | 22.0 | 61.6 | <0.001 |
| CEL | GTO 75 | 172 | 28 | 14.0 | 102.3 | <0.001 |
| CEL | GTO 84 | 184 | 16 | 8.0 | 139.4 | <0.001 |
| CEL | GTO 88 | 135 | 65 | 32.5 | 23.8 | <0.001 |
| CHP | SIN 2 | 176 | 24 | 12.0 | 114.0 | <0.001 |
| C-C | CHH 8 | 144 | 56 | 28.0 | 37.8 | <0.001 |
| COM | CHS 38 | 125 | 75 | 37.5 | 12.0 | <0.001 |
| CON | MEX 72 | 184 | 16 | 8.0 | 139.4 | <0.001 |
| CON | PUE 48 | 170 | 30 | 15.0 | 96.6 | <0.001 |
| CON | PUE 108 | 170 | 30 | 15.0 | 96.6 | <0.001 |
| C-N | GTO 16 | 171 | 29 | 14.5 | 99.4 | <0.001 |
| C-N | GTO 22 | 163 | 37 | 18.5 | 78.1 | <0.001 |
| C-N | GTO 34 | 110 | 90 | 45.0 | 1.8 | 0.2-0.1 |
| C-N | GTO 50 | 159 | 41 | 20.5 | 68.4 | <0.001 |
| C-N | GTO 60 | 179 | 21 | 10.5 | 123.2 | <0.001 |
| C-N | QRO 1 | 161 | 39 | 19.5 | 73.2 | <0.001 |
| C-N | QRO 4 | 166 | 34 | 17.0 | 85.8 | <0.001 |
| C-N | QRO 5 | 158 | 42 | 21.0 | 66.1 | <0.001 |
| C-N | ZAC 12 | 157 | 43 | 21.5 | 63.8 | <0.001 |
| GOR | CHH 146 | 188 | 12 | 6.0 | 153.1 | <0.001 |
| H-O | NAY 24 | 132 | 68 | 34.0 | 19.8 | <0.001 |
| H-O | SON 102 | 154 | 46 | 23.0 | 57.2 | <0.001 |
| JAL | NAY 6 | 171 | 26 | 14.5 | 99.4 | <0.001 |
| M-D | JAL 78 | 95 | 105 | 52.5 | 0.4 | 0.8-0.7 |
| N-T | YUC 7 | 93 | 107 | 53.5 | 0.8 | 0.8-0.7 |
| OLO | CHS 56 | 107 | 93 | 46.5 | 0.8 | 0.8-0.7 |
| PEP | MOR 17 | 161 | 39 | 19.5 | 73.2 | <0.001 |
| REV | NAY 15 | 181 | 19 | 9.5 | 129.6 | <0.001 |
| T-O | NAY 193 | 147 | 53 | 26.5 | 43.2 | <0.001 |
| TAB | JAL 42 | 182 | 18 | 9.0 | 132.8 | <0.001 |
| TAB | JAL 63 | 151 | 49 | 24.5 | 51.0 | <0.001 |
| TAB | JAL 102 | 170 | 24 | 12.0 | 108.4 | <0.001 |
| TEH | CHS 29 | 138 | 62 | 31.0 | 28.1 | <0.001 |
| TEP | CHS 26 | 181 | 19 | 9.5 | 129.6 | <0.001 |
| TEP | CHS 76 | 114 | 86 | 43.0 | 3.6 | 0.1-0.05 |
| TEP | CHS 225 | 179 | 21 | 10.5 | 123.2 | <0.001 |
| TUX | VER 39 | 87 | 113 | 56.5 | 3.1 | 0.1-0.05 |
| VAN | CHS 25 | 150 | 50 | 25.0 | 49.0 | <0.001 |
| Z-C | OAX 48 | 165 | 35 | 17.5 | 83.2 | <0.001 |
| Z-G | CHS 104 | 117 | 83 | 41.5 | 5.4 | 0.02 |
Three-character abbreviation represents the race of maize (see Table 1).
Three-letter abbreviation represents the state of Mexico in which the collection was made (see Table 1).
Kernels were scored from contiguous regions on individual ears.
χ2 for independence of rf and wx loci calculated using Yates’ correction for continuity (Yates 1934).
Representative testcross data for placement of maize Rf alleles on chromosome 2L
| Rf sourcea . | Accession no.b . | Wx . | wxc . | % rf-wx recombination . | χ2d . | P value . |
|---|---|---|---|---|---|---|
| APA | CHH 159 | 177 | 23 | 11.5 | 117.0 | <0.001 |
| A-A | PUE 91 | 100 | 100 | 50.0 | 0.0 | >0.99 |
| AZU | CHH 133 | 182 | 18 | 9.0 | 132.8 | <0.001 |
| BOL | OAX 28 | 174 | 26 | 13.0 | 108.0 | <0.001 |
| CEL | GTO 20 | 170 | 30 | 15.0 | 96.6 | <0.001 |
| CEL | GTO 69 | 156 | 44 | 22.0 | 61.6 | <0.001 |
| CEL | GTO 75 | 172 | 28 | 14.0 | 102.3 | <0.001 |
| CEL | GTO 84 | 184 | 16 | 8.0 | 139.4 | <0.001 |
| CEL | GTO 88 | 135 | 65 | 32.5 | 23.8 | <0.001 |
| CHP | SIN 2 | 176 | 24 | 12.0 | 114.0 | <0.001 |
| C-C | CHH 8 | 144 | 56 | 28.0 | 37.8 | <0.001 |
| COM | CHS 38 | 125 | 75 | 37.5 | 12.0 | <0.001 |
| CON | MEX 72 | 184 | 16 | 8.0 | 139.4 | <0.001 |
| CON | PUE 48 | 170 | 30 | 15.0 | 96.6 | <0.001 |
| CON | PUE 108 | 170 | 30 | 15.0 | 96.6 | <0.001 |
| C-N | GTO 16 | 171 | 29 | 14.5 | 99.4 | <0.001 |
| C-N | GTO 22 | 163 | 37 | 18.5 | 78.1 | <0.001 |
| C-N | GTO 34 | 110 | 90 | 45.0 | 1.8 | 0.2-0.1 |
| C-N | GTO 50 | 159 | 41 | 20.5 | 68.4 | <0.001 |
| C-N | GTO 60 | 179 | 21 | 10.5 | 123.2 | <0.001 |
| C-N | QRO 1 | 161 | 39 | 19.5 | 73.2 | <0.001 |
| C-N | QRO 4 | 166 | 34 | 17.0 | 85.8 | <0.001 |
| C-N | QRO 5 | 158 | 42 | 21.0 | 66.1 | <0.001 |
| C-N | ZAC 12 | 157 | 43 | 21.5 | 63.8 | <0.001 |
| GOR | CHH 146 | 188 | 12 | 6.0 | 153.1 | <0.001 |
| H-O | NAY 24 | 132 | 68 | 34.0 | 19.8 | <0.001 |
| H-O | SON 102 | 154 | 46 | 23.0 | 57.2 | <0.001 |
| JAL | NAY 6 | 171 | 26 | 14.5 | 99.4 | <0.001 |
| M-D | JAL 78 | 95 | 105 | 52.5 | 0.4 | 0.8-0.7 |
| N-T | YUC 7 | 93 | 107 | 53.5 | 0.8 | 0.8-0.7 |
| OLO | CHS 56 | 107 | 93 | 46.5 | 0.8 | 0.8-0.7 |
| PEP | MOR 17 | 161 | 39 | 19.5 | 73.2 | <0.001 |
| REV | NAY 15 | 181 | 19 | 9.5 | 129.6 | <0.001 |
| T-O | NAY 193 | 147 | 53 | 26.5 | 43.2 | <0.001 |
| TAB | JAL 42 | 182 | 18 | 9.0 | 132.8 | <0.001 |
| TAB | JAL 63 | 151 | 49 | 24.5 | 51.0 | <0.001 |
| TAB | JAL 102 | 170 | 24 | 12.0 | 108.4 | <0.001 |
| TEH | CHS 29 | 138 | 62 | 31.0 | 28.1 | <0.001 |
| TEP | CHS 26 | 181 | 19 | 9.5 | 129.6 | <0.001 |
| TEP | CHS 76 | 114 | 86 | 43.0 | 3.6 | 0.1-0.05 |
| TEP | CHS 225 | 179 | 21 | 10.5 | 123.2 | <0.001 |
| TUX | VER 39 | 87 | 113 | 56.5 | 3.1 | 0.1-0.05 |
| VAN | CHS 25 | 150 | 50 | 25.0 | 49.0 | <0.001 |
| Z-C | OAX 48 | 165 | 35 | 17.5 | 83.2 | <0.001 |
| Z-G | CHS 104 | 117 | 83 | 41.5 | 5.4 | 0.02 |
| Rf sourcea . | Accession no.b . | Wx . | wxc . | % rf-wx recombination . | χ2d . | P value . |
|---|---|---|---|---|---|---|
| APA | CHH 159 | 177 | 23 | 11.5 | 117.0 | <0.001 |
| A-A | PUE 91 | 100 | 100 | 50.0 | 0.0 | >0.99 |
| AZU | CHH 133 | 182 | 18 | 9.0 | 132.8 | <0.001 |
| BOL | OAX 28 | 174 | 26 | 13.0 | 108.0 | <0.001 |
| CEL | GTO 20 | 170 | 30 | 15.0 | 96.6 | <0.001 |
| CEL | GTO 69 | 156 | 44 | 22.0 | 61.6 | <0.001 |
| CEL | GTO 75 | 172 | 28 | 14.0 | 102.3 | <0.001 |
| CEL | GTO 84 | 184 | 16 | 8.0 | 139.4 | <0.001 |
| CEL | GTO 88 | 135 | 65 | 32.5 | 23.8 | <0.001 |
| CHP | SIN 2 | 176 | 24 | 12.0 | 114.0 | <0.001 |
| C-C | CHH 8 | 144 | 56 | 28.0 | 37.8 | <0.001 |
| COM | CHS 38 | 125 | 75 | 37.5 | 12.0 | <0.001 |
| CON | MEX 72 | 184 | 16 | 8.0 | 139.4 | <0.001 |
| CON | PUE 48 | 170 | 30 | 15.0 | 96.6 | <0.001 |
| CON | PUE 108 | 170 | 30 | 15.0 | 96.6 | <0.001 |
| C-N | GTO 16 | 171 | 29 | 14.5 | 99.4 | <0.001 |
| C-N | GTO 22 | 163 | 37 | 18.5 | 78.1 | <0.001 |
| C-N | GTO 34 | 110 | 90 | 45.0 | 1.8 | 0.2-0.1 |
| C-N | GTO 50 | 159 | 41 | 20.5 | 68.4 | <0.001 |
| C-N | GTO 60 | 179 | 21 | 10.5 | 123.2 | <0.001 |
| C-N | QRO 1 | 161 | 39 | 19.5 | 73.2 | <0.001 |
| C-N | QRO 4 | 166 | 34 | 17.0 | 85.8 | <0.001 |
| C-N | QRO 5 | 158 | 42 | 21.0 | 66.1 | <0.001 |
| C-N | ZAC 12 | 157 | 43 | 21.5 | 63.8 | <0.001 |
| GOR | CHH 146 | 188 | 12 | 6.0 | 153.1 | <0.001 |
| H-O | NAY 24 | 132 | 68 | 34.0 | 19.8 | <0.001 |
| H-O | SON 102 | 154 | 46 | 23.0 | 57.2 | <0.001 |
| JAL | NAY 6 | 171 | 26 | 14.5 | 99.4 | <0.001 |
| M-D | JAL 78 | 95 | 105 | 52.5 | 0.4 | 0.8-0.7 |
| N-T | YUC 7 | 93 | 107 | 53.5 | 0.8 | 0.8-0.7 |
| OLO | CHS 56 | 107 | 93 | 46.5 | 0.8 | 0.8-0.7 |
| PEP | MOR 17 | 161 | 39 | 19.5 | 73.2 | <0.001 |
| REV | NAY 15 | 181 | 19 | 9.5 | 129.6 | <0.001 |
| T-O | NAY 193 | 147 | 53 | 26.5 | 43.2 | <0.001 |
| TAB | JAL 42 | 182 | 18 | 9.0 | 132.8 | <0.001 |
| TAB | JAL 63 | 151 | 49 | 24.5 | 51.0 | <0.001 |
| TAB | JAL 102 | 170 | 24 | 12.0 | 108.4 | <0.001 |
| TEH | CHS 29 | 138 | 62 | 31.0 | 28.1 | <0.001 |
| TEP | CHS 26 | 181 | 19 | 9.5 | 129.6 | <0.001 |
| TEP | CHS 76 | 114 | 86 | 43.0 | 3.6 | 0.1-0.05 |
| TEP | CHS 225 | 179 | 21 | 10.5 | 123.2 | <0.001 |
| TUX | VER 39 | 87 | 113 | 56.5 | 3.1 | 0.1-0.05 |
| VAN | CHS 25 | 150 | 50 | 25.0 | 49.0 | <0.001 |
| Z-C | OAX 48 | 165 | 35 | 17.5 | 83.2 | <0.001 |
| Z-G | CHS 104 | 117 | 83 | 41.5 | 5.4 | 0.02 |
Three-character abbreviation represents the race of maize (see Table 1).
Three-letter abbreviation represents the state of Mexico in which the collection was made (see Table 1).
Kernels were scored from contiguous regions on individual ears.
χ2 for independence of rf and wx loci calculated using Yates’ correction for continuity (Yates 1934).
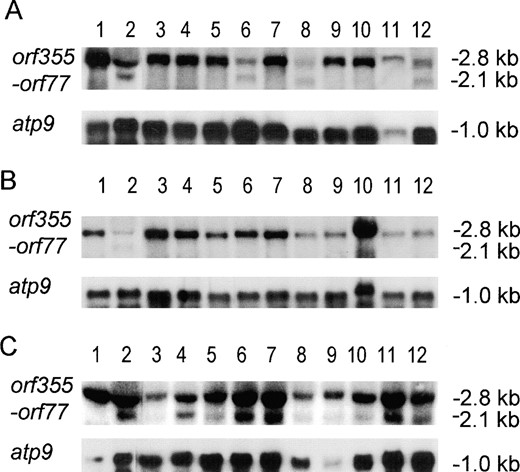
Northern blot hybridization of total cellular RNAs extracted from unfertilized ears of male-sterile and restored plants revealed that processing of the 2.8-kb orf355-orf77 transcript occurs in the presence of Rf3, Rf REV, Rf CHP, and Rf K-69-6. Processing is revealed by the presence of a 2.1-kb transcript in addition to the residual 2.8-kb transcript. No RNA processing was associated with Rf CEL (GTO 20) or Rf C-N (GTO 22) in immature cob tissue (Figure 1A). To further confirm the association between RNA processing and fertility restoration by Rf K-69-6, additional plants were analyzed. Ten rf/rf plants exhibited no 2.1-kb transcript (Figure 1B) whereas 10 individual Rf/rf plants all exhibited the 2.1-kb transcript, albeit in varying abundance (Figure 1C). The Rf3 alleles from Ky21, CI21E, and Tr (data not shown) are all associated with the RNA processing events previously described for the Rf3 allele from CE1 (Wen and Chase 1999a).
Representative testcross data for placement of teosinte Rf alleles on chromosome 2L
| Rf source . | Accession no. . | Wx . | wx . | % rf-wx recombination . | χ2a . | P value . |
|---|---|---|---|---|---|---|
| mexicana | K-69-4 | 98 | 102 | 51.0 | 0.04 | 0.90-0.80 |
| mexicana | K-69-4 | 97 | 103 | 51.5 | 0.1 | 0.80-0.70 |
| mexicana | K-69-4 | 100 | 100 | 50.0 | 0.0 | >0.99 |
| mexicana | K-69-6 | 122 | 78 | 39.0 | 9.2 | 0.01-0.001 |
| mexicana | K-69-6 | 131 | 69 | 34.5 | 18.6 | <0.001 |
| mexicana | K-69-6 | 162 | 38 | 19.0 | 75.6 | <0.001 |
| mexicana | K-69-6 | 175 | 25 | 12.5 | 111.0 | <0.001 |
| mexicana | K-69-6 | 149 | 51 | 25.5 | 47.0 | <0.001 |
| mexicana | K-69-6 | 127 | 73 | 36.5 | 10.1 | 0.01-0.001 |
| mexicana | K-69-6 | 154 | 46 | 23.0 | 57.2 | <0.001 |
| mexicana | K-69-6 | 149 | 51 | 25.5 | 47.0 | <0.001 |
| mexicana | K-69-6 | 158 | 42 | 21.0 | 66.1 | <0.001 |
| mexicana | K-69-6 | 156 | 44 | 22.0 | 61.6 | <0.001 |
| parviglumis | MO46 | 175 | 25 | 12.5 | 111.0 | <0.001 |
| parviglumis | MO46 | 170 | 30 | 15.0 | 96.6 | <0.001 |
| parviglumis | MO46 | 134 | 66 | 33.0 | 22.4 | <0.001 |
| parviglumis | MO46 | 173 | 27 | 13.5 | 105.1 | <0.001 |
| parviglumis | MO63 | 114 | 86 | 43.0 | 3.6 | 0.10-0.05 |
| parviglumis | MO63 | 109 | 91 | 45.5 | 1.4 | 0.30-0.20 |
| parviglumis | MO63 | 113 | 87 | 43.5 | 3.13 | 0.10-0.05 |
| parviglumis | MO106 | 132 | 68 | 34.0 | 19.8 | <0.001 |
| parviglumis | MO106 | 129 | 71 | 35.5 | 16.2 | <0.001 |
| parviglumis | 9477 | 184 | 16 | 8.0 | 139.4 | <0.001 |
| parviglumis | 9477 | 187 | 13 | 6.5 | 149.6 | <0.001 |
| parviglumis | 9477 | 186 | 14 | 7.0 | 146.2 | <0.001 |
| parviglumis | 9477 | 179 | 21 | 10.5 | 123.2 | <0.001 |
| Rf source . | Accession no. . | Wx . | wx . | % rf-wx recombination . | χ2a . | P value . |
|---|---|---|---|---|---|---|
| mexicana | K-69-4 | 98 | 102 | 51.0 | 0.04 | 0.90-0.80 |
| mexicana | K-69-4 | 97 | 103 | 51.5 | 0.1 | 0.80-0.70 |
| mexicana | K-69-4 | 100 | 100 | 50.0 | 0.0 | >0.99 |
| mexicana | K-69-6 | 122 | 78 | 39.0 | 9.2 | 0.01-0.001 |
| mexicana | K-69-6 | 131 | 69 | 34.5 | 18.6 | <0.001 |
| mexicana | K-69-6 | 162 | 38 | 19.0 | 75.6 | <0.001 |
| mexicana | K-69-6 | 175 | 25 | 12.5 | 111.0 | <0.001 |
| mexicana | K-69-6 | 149 | 51 | 25.5 | 47.0 | <0.001 |
| mexicana | K-69-6 | 127 | 73 | 36.5 | 10.1 | 0.01-0.001 |
| mexicana | K-69-6 | 154 | 46 | 23.0 | 57.2 | <0.001 |
| mexicana | K-69-6 | 149 | 51 | 25.5 | 47.0 | <0.001 |
| mexicana | K-69-6 | 158 | 42 | 21.0 | 66.1 | <0.001 |
| mexicana | K-69-6 | 156 | 44 | 22.0 | 61.6 | <0.001 |
| parviglumis | MO46 | 175 | 25 | 12.5 | 111.0 | <0.001 |
| parviglumis | MO46 | 170 | 30 | 15.0 | 96.6 | <0.001 |
| parviglumis | MO46 | 134 | 66 | 33.0 | 22.4 | <0.001 |
| parviglumis | MO46 | 173 | 27 | 13.5 | 105.1 | <0.001 |
| parviglumis | MO63 | 114 | 86 | 43.0 | 3.6 | 0.10-0.05 |
| parviglumis | MO63 | 109 | 91 | 45.5 | 1.4 | 0.30-0.20 |
| parviglumis | MO63 | 113 | 87 | 43.5 | 3.13 | 0.10-0.05 |
| parviglumis | MO106 | 132 | 68 | 34.0 | 19.8 | <0.001 |
| parviglumis | MO106 | 129 | 71 | 35.5 | 16.2 | <0.001 |
| parviglumis | 9477 | 184 | 16 | 8.0 | 139.4 | <0.001 |
| parviglumis | 9477 | 187 | 13 | 6.5 | 149.6 | <0.001 |
| parviglumis | 9477 | 186 | 14 | 7.0 | 146.2 | <0.001 |
| parviglumis | 9477 | 179 | 21 | 10.5 | 123.2 | <0.001 |
χ2 for independence of rf and wx loci calculated using Yates’ correction for continuity (Yates 1934).
Representative testcross data for placement of teosinte Rf alleles on chromosome 2L
| Rf source . | Accession no. . | Wx . | wx . | % rf-wx recombination . | χ2a . | P value . |
|---|---|---|---|---|---|---|
| mexicana | K-69-4 | 98 | 102 | 51.0 | 0.04 | 0.90-0.80 |
| mexicana | K-69-4 | 97 | 103 | 51.5 | 0.1 | 0.80-0.70 |
| mexicana | K-69-4 | 100 | 100 | 50.0 | 0.0 | >0.99 |
| mexicana | K-69-6 | 122 | 78 | 39.0 | 9.2 | 0.01-0.001 |
| mexicana | K-69-6 | 131 | 69 | 34.5 | 18.6 | <0.001 |
| mexicana | K-69-6 | 162 | 38 | 19.0 | 75.6 | <0.001 |
| mexicana | K-69-6 | 175 | 25 | 12.5 | 111.0 | <0.001 |
| mexicana | K-69-6 | 149 | 51 | 25.5 | 47.0 | <0.001 |
| mexicana | K-69-6 | 127 | 73 | 36.5 | 10.1 | 0.01-0.001 |
| mexicana | K-69-6 | 154 | 46 | 23.0 | 57.2 | <0.001 |
| mexicana | K-69-6 | 149 | 51 | 25.5 | 47.0 | <0.001 |
| mexicana | K-69-6 | 158 | 42 | 21.0 | 66.1 | <0.001 |
| mexicana | K-69-6 | 156 | 44 | 22.0 | 61.6 | <0.001 |
| parviglumis | MO46 | 175 | 25 | 12.5 | 111.0 | <0.001 |
| parviglumis | MO46 | 170 | 30 | 15.0 | 96.6 | <0.001 |
| parviglumis | MO46 | 134 | 66 | 33.0 | 22.4 | <0.001 |
| parviglumis | MO46 | 173 | 27 | 13.5 | 105.1 | <0.001 |
| parviglumis | MO63 | 114 | 86 | 43.0 | 3.6 | 0.10-0.05 |
| parviglumis | MO63 | 109 | 91 | 45.5 | 1.4 | 0.30-0.20 |
| parviglumis | MO63 | 113 | 87 | 43.5 | 3.13 | 0.10-0.05 |
| parviglumis | MO106 | 132 | 68 | 34.0 | 19.8 | <0.001 |
| parviglumis | MO106 | 129 | 71 | 35.5 | 16.2 | <0.001 |
| parviglumis | 9477 | 184 | 16 | 8.0 | 139.4 | <0.001 |
| parviglumis | 9477 | 187 | 13 | 6.5 | 149.6 | <0.001 |
| parviglumis | 9477 | 186 | 14 | 7.0 | 146.2 | <0.001 |
| parviglumis | 9477 | 179 | 21 | 10.5 | 123.2 | <0.001 |
| Rf source . | Accession no. . | Wx . | wx . | % rf-wx recombination . | χ2a . | P value . |
|---|---|---|---|---|---|---|
| mexicana | K-69-4 | 98 | 102 | 51.0 | 0.04 | 0.90-0.80 |
| mexicana | K-69-4 | 97 | 103 | 51.5 | 0.1 | 0.80-0.70 |
| mexicana | K-69-4 | 100 | 100 | 50.0 | 0.0 | >0.99 |
| mexicana | K-69-6 | 122 | 78 | 39.0 | 9.2 | 0.01-0.001 |
| mexicana | K-69-6 | 131 | 69 | 34.5 | 18.6 | <0.001 |
| mexicana | K-69-6 | 162 | 38 | 19.0 | 75.6 | <0.001 |
| mexicana | K-69-6 | 175 | 25 | 12.5 | 111.0 | <0.001 |
| mexicana | K-69-6 | 149 | 51 | 25.5 | 47.0 | <0.001 |
| mexicana | K-69-6 | 127 | 73 | 36.5 | 10.1 | 0.01-0.001 |
| mexicana | K-69-6 | 154 | 46 | 23.0 | 57.2 | <0.001 |
| mexicana | K-69-6 | 149 | 51 | 25.5 | 47.0 | <0.001 |
| mexicana | K-69-6 | 158 | 42 | 21.0 | 66.1 | <0.001 |
| mexicana | K-69-6 | 156 | 44 | 22.0 | 61.6 | <0.001 |
| parviglumis | MO46 | 175 | 25 | 12.5 | 111.0 | <0.001 |
| parviglumis | MO46 | 170 | 30 | 15.0 | 96.6 | <0.001 |
| parviglumis | MO46 | 134 | 66 | 33.0 | 22.4 | <0.001 |
| parviglumis | MO46 | 173 | 27 | 13.5 | 105.1 | <0.001 |
| parviglumis | MO63 | 114 | 86 | 43.0 | 3.6 | 0.10-0.05 |
| parviglumis | MO63 | 109 | 91 | 45.5 | 1.4 | 0.30-0.20 |
| parviglumis | MO63 | 113 | 87 | 43.5 | 3.13 | 0.10-0.05 |
| parviglumis | MO106 | 132 | 68 | 34.0 | 19.8 | <0.001 |
| parviglumis | MO106 | 129 | 71 | 35.5 | 16.2 | <0.001 |
| parviglumis | 9477 | 184 | 16 | 8.0 | 139.4 | <0.001 |
| parviglumis | 9477 | 187 | 13 | 6.5 | 149.6 | <0.001 |
| parviglumis | 9477 | 186 | 14 | 7.0 | 146.2 | <0.001 |
| parviglumis | 9477 | 179 | 21 | 10.5 | 123.2 | <0.001 |
χ2 for independence of rf and wx loci calculated using Yates’ correction for continuity (Yates 1934).
No newly identified Rf alleles map to 4L: While most of the newly identified Rf alleles map to 2L, maize alleles Rf A-A, Rf C-N (GTO 34), Rf M-D, Rf N-T, Rf OLO, Rf TEP (CHS 76), and Rf TUX (Table 4) and the teosinte alleles Rf K-69-4 and Rf MO63 (Table 5) are not linked to 2L. Although no rf loci for CMS-S have been mapped to chromosome 4, a region on 2L close to rf3 is duplicated on 4L (Helentjaris et al. 1988; Ahn and Tanksley 1993; Gaut and Doebley 1997; Scanlon et al. 2000). We therefore used the wx1-marked translocation T4-9(5657), with breakpoints at 4L.33 and 9S.25, to test the unplaced Rf alleles for linkage to 4L. For unknown reasons, the Rf N-T, Rf OLO, Rf TEP (CHS 76), and Rf TUX alleles do not restore fertility in combination with this translocation. Testcross data for the Rf A-A, Rf C-N (GTO 34), Rf M-D, Rf K-69-4, and Rf MO63 alleles demonstrated independence from 4L (data not shown).
Testcross data for linkage of Rf alleles and the RFLP locus whp1
| . | Rf/rf . | rf/rf . | ||
|---|---|---|---|---|
| Population . | . | . | . | . |
| [(R)Mo17/Rf CEL (GTO 20)] × (N) Mo17 (2) | 0 | 11 | 10 | 1 |
| [(S)Mo17/Rf CHP] × (N) Mo17(2) | 2 | 12 | 6 | 0 |
| [(S)Mo17 × (S) WF9/Rf C-N (GTO 22)] × (N) Mo17 | 1 | 15 | 5 | 0 |
| [(S)Mo17/Rf REV] × (N) Mo17 (2) | 2 | 8 | 12 | 0 |
| [(S)Mo17 × (S)WF9/Rf K-69-6] × (N) Mo17 | 0 | 17 | 5 | 1 |
| . | Rf/rf . | rf/rf . | ||
|---|---|---|---|---|
| Population . | . | . | . | . |
| [(R)Mo17/Rf CEL (GTO 20)] × (N) Mo17 (2) | 0 | 11 | 10 | 1 |
| [(S)Mo17/Rf CHP] × (N) Mo17(2) | 2 | 12 | 6 | 0 |
| [(S)Mo17 × (S) WF9/Rf C-N (GTO 22)] × (N) Mo17 | 1 | 15 | 5 | 0 |
| [(S)Mo17/Rf REV] × (N) Mo17 (2) | 2 | 8 | 12 | 0 |
| [(S)Mo17 × (S)WF9/Rf K-69-6] × (N) Mo17 | 0 | 17 | 5 | 1 |
χ 2 P for independence of rf and whp loci was <0.01 for all populations.
Testcross data for linkage of Rf alleles and the RFLP locus whp1
| . | Rf/rf . | rf/rf . | ||
|---|---|---|---|---|
| Population . | . | . | . | . |
| [(R)Mo17/Rf CEL (GTO 20)] × (N) Mo17 (2) | 0 | 11 | 10 | 1 |
| [(S)Mo17/Rf CHP] × (N) Mo17(2) | 2 | 12 | 6 | 0 |
| [(S)Mo17 × (S) WF9/Rf C-N (GTO 22)] × (N) Mo17 | 1 | 15 | 5 | 0 |
| [(S)Mo17/Rf REV] × (N) Mo17 (2) | 2 | 8 | 12 | 0 |
| [(S)Mo17 × (S)WF9/Rf K-69-6] × (N) Mo17 | 0 | 17 | 5 | 1 |
| . | Rf/rf . | rf/rf . | ||
|---|---|---|---|---|
| Population . | . | . | . | . |
| [(R)Mo17/Rf CEL (GTO 20)] × (N) Mo17 (2) | 0 | 11 | 10 | 1 |
| [(S)Mo17/Rf CHP] × (N) Mo17(2) | 2 | 12 | 6 | 0 |
| [(S)Mo17 × (S) WF9/Rf C-N (GTO 22)] × (N) Mo17 | 1 | 15 | 5 | 0 |
| [(S)Mo17/Rf REV] × (N) Mo17 (2) | 2 | 8 | 12 | 0 |
| [(S)Mo17 × (S)WF9/Rf K-69-6] × (N) Mo17 | 0 | 17 | 5 | 1 |
χ 2 P for independence of rf and whp loci was <0.01 for all populations.
DISCUSSION
Rf alleles for CMS-S are widespread in Mexican maize and present in some teosinte accessions (Gabay-Laughnan 2000), whereas fewer than one-half of U.S. maize inbred lines carry Rf alleles for CMS-S (Beckett 1971; Gracen and Grogan 1974; Gracen 1982; Gabay-Laughnan and Laughnan 1994; our unpublished data). These observations are consistent with studies indicating that the development of commercial varieties and inbred lines has resulted in a greater loss of diversity than has any domestication bottleneck (reviewed in Gabay-Laughnan 2000).
Testcross data for allelism of Rf K-69-9 and Rf3
| Populationa . | Fertile . | Sterile . | Total . |
|---|---|---|---|
| [(VG) Rf3 CE1/Rf K-69-6] × (N) Mo17 line | 292 | 1b | 293 |
| [(S) Rf K-69-6/Rf3 CE1] × (N) Mo17 line | 389 | 0 | 389 |
| [(S) Rf K-69-6/Rf3 Tr] × (N) Mo17 line | 233 | 0 | 233 |
| [(VG) Rf3 CE1/Rf K-69-6] × (N) Oh51A line | 296 | 0 | 296 |
| [(VG) Rf3 CE1/Rf K-69-6 × (N) Oh51A line | 295 | 0 | 295 |
| [(R) Rf K-69-6/Rf3 CE1] × (N) Oh51A line | 283 | 0 | 283 |
| [(R) Rf K-69-6/Rf3 Tr] × (N) Oh51A line | 275 | 0 | 275 |
| Populationa . | Fertile . | Sterile . | Total . |
|---|---|---|---|
| [(VG) Rf3 CE1/Rf K-69-6] × (N) Mo17 line | 292 | 1b | 293 |
| [(S) Rf K-69-6/Rf3 CE1] × (N) Mo17 line | 389 | 0 | 389 |
| [(S) Rf K-69-6/Rf3 Tr] × (N) Mo17 line | 233 | 0 | 233 |
| [(VG) Rf3 CE1/Rf K-69-6] × (N) Oh51A line | 296 | 0 | 296 |
| [(VG) Rf3 CE1/Rf K-69-6 × (N) Oh51A line | 295 | 0 | 295 |
| [(R) Rf K-69-6/Rf3 CE1] × (N) Oh51A line | 283 | 0 | 283 |
| [(R) Rf K-69-6/Rf3 Tr] × (N) Oh51A line | 275 | 0 | 275 |
Each population was planted from an individual ear.
Sterile (thin) anthers were exserted, less than half the total number.
Testcross data for allelism of Rf K-69-9 and Rf3
| Populationa . | Fertile . | Sterile . | Total . |
|---|---|---|---|
| [(VG) Rf3 CE1/Rf K-69-6] × (N) Mo17 line | 292 | 1b | 293 |
| [(S) Rf K-69-6/Rf3 CE1] × (N) Mo17 line | 389 | 0 | 389 |
| [(S) Rf K-69-6/Rf3 Tr] × (N) Mo17 line | 233 | 0 | 233 |
| [(VG) Rf3 CE1/Rf K-69-6] × (N) Oh51A line | 296 | 0 | 296 |
| [(VG) Rf3 CE1/Rf K-69-6 × (N) Oh51A line | 295 | 0 | 295 |
| [(R) Rf K-69-6/Rf3 CE1] × (N) Oh51A line | 283 | 0 | 283 |
| [(R) Rf K-69-6/Rf3 Tr] × (N) Oh51A line | 275 | 0 | 275 |
| Populationa . | Fertile . | Sterile . | Total . |
|---|---|---|---|
| [(VG) Rf3 CE1/Rf K-69-6] × (N) Mo17 line | 292 | 1b | 293 |
| [(S) Rf K-69-6/Rf3 CE1] × (N) Mo17 line | 389 | 0 | 389 |
| [(S) Rf K-69-6/Rf3 Tr] × (N) Mo17 line | 233 | 0 | 233 |
| [(VG) Rf3 CE1/Rf K-69-6] × (N) Oh51A line | 296 | 0 | 296 |
| [(VG) Rf3 CE1/Rf K-69-6 × (N) Oh51A line | 295 | 0 | 295 |
| [(R) Rf K-69-6/Rf3 CE1] × (N) Oh51A line | 283 | 0 | 283 |
| [(R) Rf K-69-6/Rf3 Tr] × (N) Oh51A line | 275 | 0 | 275 |
Each population was planted from an individual ear.
Sterile (thin) anthers were exserted, less than half the total number.
We expected that we might identify unique Rf alleles in some races of maize and teosinte from Mexico (reviewed in Gabay-Laughnan 2000). Cornbelt hybrids derive primarily from Southern Dent and Northern Flint materials (Doebley et al. 1988). The highland races of maize from central Mexico are not closely related to either component of these Cornbelt hybrids (Eagles and Lothrop 1994). Isozyme studies reveal more variation in Mexican races of maize than in Southern Dent and Northern Flint maize (Doebley et al. 1988). The races of maize from Mexico exhibit 60-75% of the isozyme variation observed among the teosintes (Doebley et al. 1985; Eyre-Walker et al. 1998; Hilton and Gaut 1998). While domestication resulted in some loss of diversity, the development of commercial U.S. varieties and inbred lines from limited source material resulted in further losses.
Testcross data for allelism of Rf C-N (GTO22) and Rf3
| Populationa . | Fertile . | Sterile . | Total . |
|---|---|---|---|
| [(R) Rf C-N/Rf3 CE1] × (N) Oh51A line | 95 | 0 | 95 |
| [(R) Rf C-N/Rf3 CE1] × (N) Oh51A line | 130 | 0 | 130 |
| [(R) Rf C-N/Rf3 CE1] × (N) Oh51A line | 169 | 0 | 169 |
| [(R) Rf C-N/Rf3 CE1] × (N) Oh51A line | 118 | 0 | 118 |
| [(R) Rf C-N/Rf3 Tr] × (N) Oh51A line | 106 | 0 | 106 |
| [(R) Rf C-N/Rf3 Tr] × (N) Oh51A line | 100 | 0 | 100 |
| [(R) Rf C-N/Rf3 Tr] × (N) Oh51A line | 108 | 0 | 108 |
| [(R) Rf C-N/Rf3 Tr] × (N) Oh51A line | 112 | 0 | 112 |
| [(R) Rf C-N/Rf3 Tr] × (N) Oh51A line | 117 | 0 | 117 |
| Populationa . | Fertile . | Sterile . | Total . |
|---|---|---|---|
| [(R) Rf C-N/Rf3 CE1] × (N) Oh51A line | 95 | 0 | 95 |
| [(R) Rf C-N/Rf3 CE1] × (N) Oh51A line | 130 | 0 | 130 |
| [(R) Rf C-N/Rf3 CE1] × (N) Oh51A line | 169 | 0 | 169 |
| [(R) Rf C-N/Rf3 CE1] × (N) Oh51A line | 118 | 0 | 118 |
| [(R) Rf C-N/Rf3 Tr] × (N) Oh51A line | 106 | 0 | 106 |
| [(R) Rf C-N/Rf3 Tr] × (N) Oh51A line | 100 | 0 | 100 |
| [(R) Rf C-N/Rf3 Tr] × (N) Oh51A line | 108 | 0 | 108 |
| [(R) Rf C-N/Rf3 Tr] × (N) Oh51A line | 112 | 0 | 112 |
| [(R) Rf C-N/Rf3 Tr] × (N) Oh51A line | 117 | 0 | 117 |
Each population was planted from an individual ear.
Testcross data for allelism of Rf C-N (GTO22) and Rf3
| Populationa . | Fertile . | Sterile . | Total . |
|---|---|---|---|
| [(R) Rf C-N/Rf3 CE1] × (N) Oh51A line | 95 | 0 | 95 |
| [(R) Rf C-N/Rf3 CE1] × (N) Oh51A line | 130 | 0 | 130 |
| [(R) Rf C-N/Rf3 CE1] × (N) Oh51A line | 169 | 0 | 169 |
| [(R) Rf C-N/Rf3 CE1] × (N) Oh51A line | 118 | 0 | 118 |
| [(R) Rf C-N/Rf3 Tr] × (N) Oh51A line | 106 | 0 | 106 |
| [(R) Rf C-N/Rf3 Tr] × (N) Oh51A line | 100 | 0 | 100 |
| [(R) Rf C-N/Rf3 Tr] × (N) Oh51A line | 108 | 0 | 108 |
| [(R) Rf C-N/Rf3 Tr] × (N) Oh51A line | 112 | 0 | 112 |
| [(R) Rf C-N/Rf3 Tr] × (N) Oh51A line | 117 | 0 | 117 |
| Populationa . | Fertile . | Sterile . | Total . |
|---|---|---|---|
| [(R) Rf C-N/Rf3 CE1] × (N) Oh51A line | 95 | 0 | 95 |
| [(R) Rf C-N/Rf3 CE1] × (N) Oh51A line | 130 | 0 | 130 |
| [(R) Rf C-N/Rf3 CE1] × (N) Oh51A line | 169 | 0 | 169 |
| [(R) Rf C-N/Rf3 CE1] × (N) Oh51A line | 118 | 0 | 118 |
| [(R) Rf C-N/Rf3 Tr] × (N) Oh51A line | 106 | 0 | 106 |
| [(R) Rf C-N/Rf3 Tr] × (N) Oh51A line | 100 | 0 | 100 |
| [(R) Rf C-N/Rf3 Tr] × (N) Oh51A line | 108 | 0 | 108 |
| [(R) Rf C-N/Rf3 Tr] × (N) Oh51A line | 112 | 0 | 112 |
| [(R) Rf C-N/Rf3 Tr] × (N) Oh51A line | 117 | 0 | 117 |
Each population was planted from an individual ear.
The newly identified Rf alleles occur naturally: The 45 Rf alleles from Mexican maize and 6 Rf alleles from teosinte are all homozygous viable and condition no deleterious effects on endosperm development. In contrast, 51 of 60 restoring alleles recovered by spontaneous mutation are homozygous lethal (rfl alleles; Laughnan and Gabay 1978; Gabay-Laughnan et al. 1995; Gabay-Laughnan 1997; reviewed in Chase and Gabay-Laughnan 2003). In most cases, the Rfl/rfl/rfl endosperm, carrying two doses of the restoring allele, is reduced in size compared to Rfl/Rfl/rfl endosperm carrying only 1 restoring allele (Laughnan and Gabay 1975; Gabay-Laughnan et al. 1995; reviewed in Chase and Gabay-Laughnan 2003). Furthermore, in the case of 9 of the homozygous-viable (rfv) restoring alleles recovered by spontaneous mutation, rfv/rfv plants are reduced in stature and late maturing while kernels homozygous for the restoring allele exhibit poor endosperm development (Gabay-Laughnan et al. 1995; reviewed in Chase and Gabay-Laughnan 2003).
The restoring alleles from Mexican maize and teosinte could potentially occur naturally or result from spontaneous mutations. The wild-type rowing and kernel size on ears resulting from self-pollination of heterozygous (Rf/rf) plants and the recovery of homozygous (Rf/Rf) progeny, verified by pollen analysis, support the in Mexican materials.
—Orf355-orf77 transcript processing in the presence or absence of Rf alleles. Total cellular RNAs were extracted from immature cobs of paired siblings that did or did not carry an Rf allele. RNA samples were denatured, fractionated, and hybridized as described in materials and methods. The 2.1-kb RNA is indicative of Rf-associated processing; the 2.8-kb RNA is residual, unprocessed transcript. (A) Screening of Rf alleles for RNA processing activity. RNA samples were extracted from individual plants of the following genotypes: (1) CMS-S Mo17 rf/rf, (2) CMS-S Mo17 Rf3/Rf3, (3) rf/rf sibling of plant 4, (4) CMS-S Rf CEL (GTO 20)/rf, (5) rf/rf sibling of plant 6, (6) CMS-S Rf REV/rf, (7) rf/rf sibling of plant 8, (8) CMS-S Rf CHP/rf, (9) rf/rf sibling of plant 10, (10) CMS-S Rf C-N (GTO 22)/rf, (11) rf/rf sibling of plant 12, and (12) CMS-S Rf K-69-6/rf. (B and C) Cosegregation of Rf K-69-6 with RNA processing activity. Lanes 1 and 2 are as described for corresponding lanes in A. (B) Lanes 3-12 contain RNA samples extracted from male-sterile siblings in the population segregating for Rf K-69-6. (C) Lanes 3-12 contain RNA samples extracted from male-fertile siblings carrying the Rf K-69-6 gene. All blots were first hybridized with radiolabeled orf355-orf77 probe and were subsequently stripped and rehybridized with the atp9 probe as a control for the loading of RNA samples.
The Rf alleles on 2L may be allelic to Rf3: The 51 Rf alleles from Mexican maize and teosinte represent an undetermined number of genetic loci. Because they behave as naturally occurring Rf alleles, they are potentially alleles of the standard CMS-S restorer, Rf3, which maps to 2L (Laughnan and Gabay 1978; Kamps and Chase 1997; Table 3). Of the 45 Rf alleles from Mexican maize, 38 mapped to 2L (Table 4). In addition, four of the six teosinte Rf alleles mapped conclusively to 2L (Table 5). In contrast, of 43 spontaneous restorer mutations tested, only 7 map to 2L, and all 7 are homozygous lethal (Gabay-Laughnan et al. 1995; reviewed in Chase and Gabay-Laughnan 2003).
All of the newly identified Rf alleles on 2L are potentially allelic to Rf3. Four Rf alleles from Mexican maize and one from teosinte were linked to the whp1 locus (Table 6), which is also linked to the rf3 locus (Kamps and Chase 1997). Direct tests of allelism demonstrated that two Rf alleles from teosinte and the Rf C-N (GTO 22) allele from maize were allelic or tightly linked to Rf3 (Tables 7 and 8).
The prevalence of Rf alleles for CMS-S in N-cytoplasm maize and teosinte from Mexico indicates that these alleles likely have a function separate from restoration of fertility. An additional function for Rf alleles has been invoked to explain the fact that the Rf2 allele, one of two required for restoration of fertility in CMS-T maize, has been maintained during evolution (Schnable and Wise 1994; Wise et al. 1999). The only known source of the nonrestoring rf2 allele is the maize inbred line Wf9 (Wise et al. 1999). Thus, Rf alleles of genes that function in plant development or metabolism may by chance also function to restore male fertility (Hanson et al. 1999; Wise et al. 1999; Gabay-Laughnan 2000). This hypothesis has recently been refined to include the possibility that restorers may arise by duplication and modification of such genes (Bentolila et al. 2002).
The unplaced Rf alleles do not group by race or geographic region: We have been unable to determine a relationship among the sources of the unplaced Rf alleles. The seven maize alleles that do not map to 2L derive from two of the three distinct racial groups characterized by Goodman and Brown (1988). Rf A-A and Rf C-N (GTO 34) were identified in closely related races within the group that also includes the race carrying Rf M-D. Rf TEP (CHS 76) and Rf TUX were identified in closely related races of a separate racial group that also contains the races carrying Rf OLO and Rf N-T. Note that other accessions of the races Cónico Norteño and Tepecintle carry restorers that do map to 2L.
The seven unplaced maize restorers were identified in accessions from six different states of Mexico (see Figure 5 in Wellhausen et al. 1952). Rf A-A was identified in an accession from Puebla, Rf C-N (GTO 34) from Guanajuato, Rf M-D from Jalisco, Rf N-T from Yucatan, Rf OLO and Rf TEP (CHS 76) from Chiapas, and Rf TUX from Veracruz. Yucatan is a state in the Gulf Coastal Plain while the remaining states are all from southern Mexico. These states encompass a variety of geographic regions (see Figure 6 in Wellhausen et al. 1952) and a range of altitudes, from 31 m to >2000 m elevation. Two unplaced restorers were also identified in the teosintes, Rf K-69-4 in ssp. mexicana and Rf MO63 in ssp. parviglumis. Rf K-69-4 was identified in teosinte from the Central Plateau while Rf MO63 is present in teosinte collected from southern Guerrero.
While most of the Rf alleles map to 2L, at least one other allele is relatively rare but geographically dispersed in Mexican maize and teosinte. Three unplaced maize Rf alleles and two unplaced teosinte alleles were tested and do not map to the duplicate region of 2L found on 4L. We are currently testing allelism among the unplaced Rf alleles and the RfA allele, to determine whether there are Mexican maize and teosinte Rf alleles that are not represented in present-day U.S. maize.
A possible variant Rf3 allele: Restoration in S-cytoplasm maize is but one example of systems in which transcripts of the CMS-associated mitochondrial region are altered by a specific Rf allele. Similar observations have been made in CMS systems of brassica (Singh and Brown 1991; Singh et al. 1996; Li et al. 1998), petunia (Pruitt and Hanson 1991), rice (Iwabuchi et al. 1993), sorghum (Tang et al. 1996; 1998), sunflower (Moneger et al. 1994), and T-cytoplasm maize (Dewey et al. 1987; Kennel et al. 1987; Kennel and Pring 1989; Wise et al. 1996; Dill et al. 1997). Furthermore, Rf alleles have been associated with processing of normal mitochondrial gene transcripts in several of these systems, including CMS-S maize (reviewed in Kempken and Pring 1999; Wise and Pring 2002). The Rf3 allele cosegregates with novel shorter atp6 and cob mitochondrial gene transcripts in addition to processed orf355-orf77 transcripts (Wen and Chase 1999a).
At this time, it is not known whether the variation in normal mitochondrial gene transcripts that is associated with fertility restoration results from the action of nuclear Rf alleles or from the action of closely linked modifier-of-mitochondrial-transcript (mmt) loci (reviewed in Kempken and Pring 1999). On the basis of the rf loci recently cloned from petunia (Bentolila et al. 2002), brassica (Brown et al. 2003), and rice (Kazama and Toriyama 2003), either explanation is possible. Each of these cloned Rf alleles encodes a protein of the pentatricopeptide repeat (PPR) class, and additional ppr genes are tightly linked to each of the cloned rf genes. PPR proteins are encoded by a large multigene family in Arabidopsis thaliana, and most of these gene products are predicted to be organelle targeted (reviewed in Small and Peters 2000). Furthermore, PPR proteins are implicated in the expression of organelle genomes from organisms as diverse as fungi and maize (reviewed in Barkan and Goldschmidt-Clermont 2000). Mmt activity associated with Rf alleles could be due to the action of tightly linked genes that encode PPR proteins having no role in fertility restoration or to the action of an Rf gene product performing multiple functions within the mitochondria.
Through examination of RNA processing events, we identified a possible variant of the maize Rf3 allele. The Rf3 allele from the inbred lines CE1, Ky21, CI21E, and Tr, as well as Rf REV, Rf CHP, and Rf K-69-6, is associated with processing of the 2.8-kb orf355-orf77 transcript in immature cobs (Figure 1; data not shown). No RNA processing was observed in immature cobs of plants carrying Rf CEL (GTO 20) or Rf C-N (GTO22) alleles (Figure 1), even though both are linked to the whp1 region of 2L (Table 6) and Rf C-N (GTO22) is allelic or tightly linked to Rf3 (Table 8).
In petunia, the Rf locus is composed of duplicated genes containing PPR proteins (Bentolila et al. 2002). Of interest here is the fact that a nonrestoring allele carries a deletion in the promoter region that changes its tissue specificity. While the petunia restoring allele is expressed in floral buds, the transcript from the nonrestoring allele is detectable only in root tissue. In CMS-S maize, mitochondria of both immature cobs and microspores accumulate the 2.8-kb orf355-orf77 transcript, and Rf3-associated processing is observed in both tissue types (Zabala et al. 1997). Analysis of orf355-orf77 transcripts in Rf C-N (GTO 22) pollen will be needed to determine if we have identified an Rf allele with different tissue specificity. Alternatively, Rf C-N (GTO 22) may be tightly linked to the rf3 locus and restore male fertility by a different molecular mechanism.
The identification of a variant allele at or tightly linked to the rf3 locus would be an unusual finding. There is only one reported case of linked variant restoring alleles. Rf1 (chromosome 3) and Rf2 (chromosome 9) are unlinked restorers of CMS-T maize, and both are required for fertility restoration (Duvick 1965). The rare Rf8 and Rf* alleles can each substitute for the Rf1 allele to partially restore fertility in the presence of Rf2 (Dill et al. 1997). Rf8 and Rf* are allelic either to each other or to alleles at tightly linked loci on 2L (Wise and Pring 2002). Rf1, Rf8, and Rf* each independently mediate the processing of RNAs transcribed from the mitochondrial CMS-T gene, T-urf13 (Dill et al. 1997). Interestingly, Rf8, Rf*, Rf3, and Rf C-N (GTO 22) all map to the whp1 region on chromosome 2L. On the basis of these observations, the whp1 region of 2L in maize potentially encodes an array of rf genes that are larger and more complex than those characterized in petunia (Bentolila et al. 2002), brassica (Brown et al. 2003), or rice (Kazama and Toriyama 2003).
Acknowledgement
We thank the International Maize and Wheat Improvement Center (CIMMYT), Jack Beckett, John Doebley, Don Duvick, Major Goodman, and the Maize Genetics Cooperation Stock Center for providing the stocks used in these studies. We thank Udo Wienand for the whp1 probe and Eugene Kuzmin and Kathleen Newton for sharing unpublished data. We appreciate the excellent technical assistance of Janet Day Jackson and Shane Zimmerman. Rebecca Heid provided expert secretarial assistance to S.G.-L. Our work was funded in part by U.S. Department of Agriculture National Research Initiative Competitive Grants Program awards 95-37301-2039 to C.D.C., 96-35300-3778 to S.G.-L., and 00-35300-9409 to S.G.-L. and C.D.C. Additional support was provided by Illinois Foundation Seeds and by the University of Florida Agriculture Experiment Station. This article is no. R-09491 in the Florida Agriculture Experiment Station Journal Series.
Footnotes
Communicating editor: J. A. Birchler
LITERATURE CITED