-
PDF
- Split View
-
Views
-
Cite
Cite
Xin Chen, Qinghong Li, Janice A Fischer, Genetic Analysis of the Drosophila DNAprim Gene: The Function of the 60-kD Primase Subunit of DNA Polymerase Opposes the fat facets Signaling Pathway in the Developing Eye, Genetics, Volume 156, Issue 4, 1 December 2000, Pages 1787–1795, https://doi.org/10.1093/genetics/156.4.1787
Close - Share Icon Share
Abstract
The Drosophila DNAprim gene encodes the large subunit (60 kD) of DNA primase, the part of DNA polymerase α that synthesizes RNA primers during DNA replication. The precise function of the 60-kD subunit is unknown. In a mutagenesis screen for suppressors of the fat facets (faf) mutant eye phenotype, we identified mutations in DNAprim. The faf gene encodes a deubiquitinating enzyme required specifically for patterning the compound eye. The DNA sequences of four DNAprim alleles were determined and these define essential protein domains. We show that while flies lacking DNAprim activity are lethal, flies with reduced DNAprim activity display morphological defects in their eyes, and unlike faf mutants, cell cycle abnormalities in larval eye discs. Mechanisms by which DNA primase levels might influence the faf-dependent cell communication pathway are discussed.
DNA polymerase α is a highly conserved protein composed of four subunits whose approximate molecular weights are 180, 75, 60, and 50 kD in eukaryotes (Lehman and Kaguni 1987). The 180-kD subunit contains the DNA polymerase activity (Cotterill et al. 1987a; Pizzagalli et al. 1988; Wong et al. 1988) and the two smallest subunits constitute DNA primase, an enzyme that synthesizes the RNA primers required during DNA replication (Conaway and Lehman 1982; Cotterill et al. 1987b; Bakkenist and Cotterill 1994). The 50-kD subunit alone has been shown to be sufficient for primase activity (Nasheuer and Grosse 1988; Santocanale et al. 1993; Bakkenist and Cotterill 1994). Although its precise function is unknown, the 60-kD subunit is considered to be part of DNA primase because it can bind ribonucleotides (Foiani et al. 1989) and it can alter the activity of the 50-kD subunit (Copeland 1997).
DNA primase is of interest from a structural viewpoint and also from a regulatory perspective: how does primase interact with DNA polymerase and DNA and how is the activity of DNA polymerase α coordinated with the cell cycle? Prerequisite to answering these questions, the Drosophila genes for all four DNA polymerase α subunits have been cloned and analyzed (Hirose et al. 1991; Cotterill et al. 1992; Melov et al. 1992; Bakkenist and Cotterill 1994; Huikeshoven and Cotterill 1999).
The Drosophila fat facets (faf) gene encodes a deubiquitinating enzyme essential in only two tissues of the fly, the eye and ovary (Fischer-Vize et al. 1992; Huang et al. 1995). In the compound eye, faf is required early during development for a cell communication event that limits to eight the number of photoreceptors in each unit eye (also called a facet or ommatidium; Fischer-Vize et al. 1992; Huang and Fischer-Vize 1996). Ubiquitin (Ub) is a highly conserved 76-amino-acid peptide that can be covalently linked to other proteins, usually via an isopeptide bond between the terminal Glycine residue of Ub and an internal Lysine residue of the substrate (Hershko 1998). A chain of isopeptide-linked Ub moieties targets the protein to which it is attached for degradation by the proteasome (Hershko 1998).
Deubiquitinating enzymes cleave Ub-protein bonds (Wilkinson and Hochstrasser 1998). Faf protein is one of a large family of deubiquitinating enzymes called ubiquitin-specific processing proteases (Ubps), which are recognizable by two small conserved amino acid domains that constitute the catalytic core (Wilkinson and Hochstrasser 1998); the Drosophila Genome Project has identified at least 18 Ubps in the fly genome (Rubin et al. 2000; Chen and Fischer 2000).
The faf gene is of particular interest because it is required for a specific step in eye patterning and also because, unique among Ubps, Faf appears to regulate the Ub pathway by deubiquitinating a specific substrate, thus preventing its degradation. The evidence for this model is genetic; mutations in a gene encoding a proteasome subunit [l(3)73Ai; Saville and Belote 1993] or in a gene encoding a Ub-conjugating enzyme required for attaching Ub to target proteins (UbcD1; Treier et al. 1992; Cenci et al. 1997; Neufeld et al. 1998) act as strong dominant suppressors of the faf mutant eye phenotype (Wu et al. 1999). These results indicate that faf function antagonizes the function of ubiquitination and the proteasome. In addition, slight overexpression of a gene called liquid facets (lqf) obviates the requirement for faf in the eye; this observation is consistent with a model where Faf protects a substrate from degradation and where that substrate is the Lqf protein (Cadavid et al. 2000).
The lqf gene, identified in mutagenesis screens for dominant enhancers of faf mutations, encodes the Drosophila homolog of vertebrate epsin, a component of the endocytosis complex (Fischer et al. 1997; Chen et al. 1998; Cadavid et al. 2000). Although it is not certain whether Lqf is the substrate of Faf in the eye, Lqf must be a critical player, downstream of faf, in the faf-dependent cell communication pathway.
To identify additional components of the faf pathway, we performed a screen for dominant suppressors of the faf mutant eye phenotype. We found that mutations in DNAprim (Spradling et al. 1999), which encodes the 60-kD subunit of DNA primase, are strong dominant suppressors of faf mutations. We have determined the DNA sequences of several mutant DNAprim genes and these reveal important functional domains of the protein. In addition, we found that while strong DNAprim mutations are lethal, flies with reduced DNAprim activity have defects in eye development. Moreover, we show that in larval eye discs with reduced DNAprim activity, the cell cycle is perturbed. We discuss possible roles for DNA primase in the faf-dependent cell communication pathway.
MATERIALS AND METHODS
Drosophila strains: The Bloomington Stock Center provided l(3)j10B2 and other lethal P{w+} insertions in the region (Spradling et al. 1999), In(3L)78Cb1 (Russell et al. 1996), the third chromosome deficiency kit, which includes Df(3L)rdgC-co2 (Flybase 1999) and Df(3L)ri-79c (Jurgens et al. 1984), the Δ2-3 Sb chromosome (Robertson et al. 1988) used for P-element mobilization, the th st cu sr e chromosome, the P{w+} insertion in 28E used for meiotic mapping, and the Act5C-Gal4 transgene described in Flybase (1999). The UbcD1 (Cenci et al. 1997; Neufeld et al. 1998) and l(3)73Ai (Saville and Belote 1993) alleles used for complementation tests are described in Huang et al. (1995) and Wu et al. (1999). The three alleles of l(3)77ABe (Lukinova et al. 1999) were obtained from N. Lukinova and M. Fortini. The P{w+} insertions in 25D, 30C, and 57B used for meiotic mapping were provided by K. Moses. The faf alleles and P{w+, faf+} transformants are maintained in our laboratory and are described in Fischer-Vize et al. (1992). Marker mutations and balancers are described in Lindsley and Zimm (1992). All flies were grown on standard media at 25° and strains were constructed using standard cross schemes.
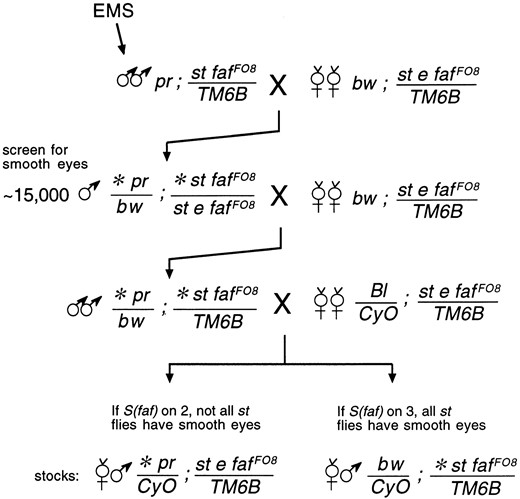
EMS mutagenesis screen: Males were mutagenized with EMS according to the method of Lewis and Bacher (1968), crossed to females, and progeny were collected for 4 days. As shown in Figure 1, ∼15,000 mutagenized second and third chromosomes (∼80% of the genome) were screened for S(faf) mutations. All of the suppressors were tested for complementation with UbcD1 and l(3)73Ai mutations, which were shown previously to act as strong suppressors of faf (Huang et al. 1995; Wu et al. 1999).
Mapping S(faf) mutants: Second chromsome-linked S(faf) mutants were mapped roughly with respect to four P{w+} insertions in 25D, 28E, 30C, and 57B as follows. Females of the genotype w; S(faf)/P{w+}; st faf FO8/TM6B were crossed with w; st faf FO8/TM6B males. Approximately 100 male progeny with pigmented eyes [thus of the genotype w; P{w+}, S(faf)?/+; st faf FO8/TM6B] were crossed individually with w; st faf FO8/TM6B females. The faf FO8 progeny were scored for the presence or absence of the S(faf) mutation. Third chromosome mutations were localized with respect to four recessive markers (th, cu, sr, and e) as follows. Female progeny of the genotype w, P{w+, faf+}/w; S(faf), st faf FO8/th st cu sr e faf FO8 were crossed with w; th st cu sr e faf FO8/TM6B males. [P{w+, faf+} is a P-element insertion containing a wild-type faf gene that complements the faf mutant phenotype completely; it was used here to complement the sterility of faf - females.] All non-TM3, white-eyed flies [without P{w+, faf+}] with smooth eyes [with S(faf)] were scored for the presence of the four recessive markers. After meiotic mapping, physical mapping of some of the S(faf) mutations was performed. Individual S(faf) lines were crossed with appropriate third chromosome deficiencies and S(faf)/Df progeny were examined for lethality or a mutant eye phenotype.
Mobilization of P{lacW}j10B2: One hundred individual crosses were performed in which two males of the genotype w/Y; P{lacW}j10B2/Δ2-3 Sb were crossed with w; st e faf FO8/TM6B females. Progeny with chromosomes in which the P element had hopped out were recognized as white-eyed flies; one white-eyed progeny male of the genotype w; P{lacW}j10B2/TM6B from each of the 100 original crosses was crossed to w; S(faf)240/TM3 females. The w; P{lacW}j10B2Δ/TM3 progeny were used to generate stocks of each hop-out chromosome and the other progeny were examined to determine if P{lacW}j10B2Δ/S(faf)240 progeny were viable and if so, if they display a mutant phenotype.
Plasmid rescue: Molecular biology procedures were carried out using standard techniques (Sambrook et al. 1989). Total genomic DNA prepared from P{lacW}j10B2/+ flies was restricted to completion with EcoRI, ligated, and electroporated into Escherichia coli. The plasmid recovered was digested with EcoRI and XbaI to release the Drosophila genomic DNA fragment adjacent to the P element. The end sequences of the genomic DNA fragment were determined, and as one of them contains some plasmid sequence, the precise insertion point of the P element within the genomic DNA could be determined.
Molecular analysis of DNAprim alleles: The DNAprim gene, whose open reading frame is contained in a single exon, was amplified by polymerase chain reaction (PCR) from genomic DNA prepared from the following heterozygotes: S(faf) 240/+, l(3)77ABeJ1/+, l(3)77ABeJ9/+, l(3)77ABeL5/+, and l(3) 77ABeT2/+. The primers used for PCR were (1) 5′-ATTAC TATAGCAATTCCCGCGAC-3′ and (2) 5′CGTGTACTTGTG CCGGAGGTG-3′. Genomic DNA was prepared by adding one fly to a microfuge tube containing 50 μl of buffer (10 mm Tris pH 8.2, 1 mm EDTA, 25 mm NaCl) and 1 μl of proteinase K (20 mg/ml). The fly was homogenized with a pipette tip, incubated at 37° for 1 hr, and then at 100° for 2 min to inactivate the proteinase K. A 4-ml aliquot of this homogenate was used in a single PCR reaction. The PCR products, mixtures of two different DNA populations, were subcloned into pBLUESCRIPT (Stratagene, La Jolla, CA) and the DNA sequences of several subclones from each PCR reaction were determined. The sequences were compared with the DNA sequence in Huikeshoven and Cotterill (1999) and also with those determined by the Drosophila Genome Project (genomic clone AC017679 and cDNA clone GM13640). For clones where DNA sequence differences from wild type were detected, that sequence was confirmed by sequencing subclones from a second PCR reaction. The inversion breakpoint of In(3L)78Cb1 was localized to a ∼2.8-kb EcoRI fragment of genomic DNA that includes the first ∼160 bp of the DNAprim open reading frame and ∼2.6 kb of sequence upstream by the following genomic DNA blotting experiment. A blot of genomic DNA prepared from wild-type and In(3L)78Cb1/+ flies restricted to completion with EcoRI was hybridized with a probe corresponding to the open reading frame of the DNAprim gene. The level of the ∼2.8-kb EcoRI fragment in the In(3L)78Cb1/+ DNA was half of that in wild type.
Complementation of DNAprim mutations by a UAS-DNAprim transgene: A 1.8-kb DNA fragment containing the DNAprim open reading frame was generated by PCR using the Advantage-HF2 kit (CLONTECH, Palo Alto, CA) with wild-type genomic DNA prepared as described above as the template and the following two primers: (1) 5′-GGCGCGCCATTACTATAG CAATTCCCGCGAC-3′ and (2) 5′-GGCGCGCCCGTGCTAC TTGTGCCGGAGGTG-3′. The DNA fragment was ligated into the vector PCR2.1 (Invitrogen, San Diego) to generate PCR2. 1-DNAprim and its DNA sequence was determined. The DNAprim gene was then isolated as a 1.8-kb AscI fragment from PCR2.1-DNAprim and ligated into the AscI site of the P-element transformation vector pUAST-XA (Huang and Fischer-Vize 1996) to generate pUAS-DNAprim. One second chromosome-linked P-element transformant was generated with pUAS-DNAprim using standard procedures (Spradling 1986) with w1118 flies as the host. Flies homozygous for l(3)j10B2 that also carry one copy each of the Act5C-Gal4 and UAS-DNAprim transgenes and thus express the cloned DNAprim gene ubiquitously were viable and apparently wild type except that they were nearly sterile. Expression of the UAS-DNAprim transgene using the Act5C-Gal4 driver resulted in a few viable l(3)j10B2/S(faf)240 escapers that died after 1 day.
Phenotypic analysis of eyes: Scanning electron micrographs (Huang et al. 1995) and 1-μm plastic sections of adult eyes (Tomlinson and Ready 1987a) were prepared as described previously. Cobalt sulfide staining of pupal eyes was performed using a modified version of the procedure described in Wolff and Ready (1991) as follows. Retina/brain complexes of 40-60-hr pupae grown at 22° were dissected in 0.1 m NaPO4, washed in H2O, and then fixed in 2% glutaraldehyde in 0.1 m NaPO4 for 10 min. After washing in 0.1 m NaPO4, the retinas were incubated in 4% Co(NO3)2 for 10-15 min, and then rinsed thoroughly in H2O. With a fine tungsten needle, the outsides of the retinas were scraped clean of precipitates and peripodial membrane. After washing again in H2O, mouth hooks and brains were removed and the retinas were mounted in 80% glycerol. All eye preparations were photographed with a Zeiss Axioplan microscope.
BrdU incorporation experiments: Eye disc/brain complexes were dissected in Drosophila Schneider’s medium at room temperature and then incubated for 2 hr in Schneider’s medium with 75 μg/ml bromodeoxyuridine (BrdU; Boehringer Mannheim, Indianapolis) in a small petri dish. All of the following manipulations were also carried out in small petri dishes. Disc complexes were washed 10 min in fresh Schneider’s, twice for 10 min in PBS, and then fixed in Carnoy’s fixative (60% ethanol, 10% glacial acetic acid, 30% chloroform) for 30 min. The fixed tissue was rehydrated with an ethanol series (5 min each in 70, 50, and 30%), then permeabilized in PBS + 0.6% Triton X-100 for 30 min. The DNA was denatured by a 30-min incubation in PBS + 0.3% Triton X-100, 2N HCl, and then washed twice for 10 min in PBS + 0.3% Triton X-100. The following manipulations were carried out in 18 μl of solution in 96-well microtiter plates. The disc complexes were incubated in primary antibody (anti-BrdU; Becton Dickinson, San Jose, CA) diluted 1:100 in PBS + 0.3% Triton X-100 and 10% goat serum overnight at 4°. After two 10-min washes in PBS + 0.3% Triton X-100, the tissues were incubated for 2 hr at room temperature in secondary antibody (HRP-conjugated goat anti-mouse; Vector, Burlingame, CA) diluted 1:50 in PBS + 0.3% Triton X-100 and 10% goat serum. After two 10-min washes in PBS + 0.3% Triton X-100, the signal was developed in HRP-staining solution (1× PBS, 0.1% Triton X-100, 0.5 mg/ml diaminobenzidine, 0.003% H2O2) for ∼1 min. The tissues were washed twice for 5 min in PBS + 0.3% Triton X-100, transferred to a petri dish containing 50% ethanol for 10 min, and then to 100% ethanol twice for 10 min. Finally, the eye discs were dissected away from the complex and mounted in DPX (Fluka, Buchs, Switzerland).
RESULTS
Mutations in a gene at 77B6/7 are suppressors of faf mutations: To identify genes in the faf pathway, we performed an EMS mutagenesis screen for dominant suppressors of the faf mutant phenotype in the eye (Figure 1). We screened ∼15,000 flies and isolated 13 suppressor mutations [S(faf)], five of which were strong and were analyzed further (Table 1 and Figure 2). One of these is an allele of UbcD1, which was shown previously to be a strong suppressor of faf mutations (Wu et al. 1999). Of the remaining 4 S(faf) mutations, only S(faf)240 has an essential function; S(faf)240 results in a mutant phenotype when homozygous or in trans to a deficiency chromosome in otherwise wild-type flies (see Table 1 footnote).
The S(faf)240 allele was mapped meiotically between th and cu (materials and methods). Subsequent physical mapping positioned S(faf)240 within 77A1-77C; flies that are S(faf)240/Df(3L)rdgC-co2 (77A1-77D1) die early during development, whereas S(faf)240/Df(3L)ri-79c (77C-78A) flies are wild type. Lethal P{w+} insertions within 77A1-77C were tested for complementation of S(faf)240, and the insertion P{lacW}j10B2 (l(3)j10B2) in 77B6/7 (Spradling et al. 1999) failed to complement the lethality of S(faf)240. In addition, l(3)j10B2 and Df(3L)rdgC-co2 are also dominant suppressors of faf (Figure 2G). Three EMS-induced lethal mutations in the l(3)77ABe gene and In(3L)78Cb1 (78B5-9/78C8-9) were reported to be allelic to l(3)j10B2 (Lukinova et al. 1999) and all of them are also allelic to S(faf)240 (Table 2).
As l(3)j10B2 does not complement the lethality of S(faf)240 and both of these chromosomes as well as a deficiency that uncovers each lethal mutation suppress faf, the mutant gene causing the lethality is most likely also causing the suppression of faf mutations. To test whether the lethality of l(3)j10B2 is due to the P{lacW} insertion, the P element was mobilized and 100 chromosomes from which it had hopped out were isolated (materials and methods). Of these 100 chromosomes, 57 of them, presumably precise excisions of the P element, resulted in viable flies with no obvious mutant phenotypes when homozygous or in trans to S(faf)240. Thus, the P-element insertion was the cause of the lethal mutation allelic to S(faf)240 on the chromosome containing P{lacW}j10B2. The other 43 hop-out chromosomes, presumably imprecise excisions, fell into two groups: when homozygous or in trans to S(faf)240, 40 chromosomes were lethal and 3 were viable, resulting in flies with rough eyes. Presumably, the lethal chromosomes are deletions of most or all of the gene disrupted by P{lacW}j10B2, and the viable chromosomes are smaller deletions that disrupt the gene partially (see below). Two of the lethal imprecise excision chromosomes and three of the viable ones were kept for further analysis (Table 2).
—Cross scheme for suppressor of faf mutagenesis screen. All of the markers are described in Lindsley and Zimm (1992). The asterisk (*) indicates a mutagenized chromosome. The faf FO8 allele is a strong mutant; its eye phenotype when homozygous or in trans to a deficiency is nearly indistinguishable from that of the null allele faf BX4 (Fischer-Vize et al. 1992). The DNA lesion of faf FO8 results in a single amino acid change; the second His of Faf’s catalytic domain is changed to Tyr (Chen and Fischer 2000). This His residue is required for wild-type Faf function, but may not be absolutely essential (Huang et al. 1995).
Suppressors of faf
| Suppressors . | Location . | Comments . |
|---|---|---|
| S(faf)86 | 25D-30C | Homozygotes wild type |
| S(faf)184 | 88D2/3 | Allelic to UbcD1 |
| S(faf)196 | near th | Wild type in trans to Df |
| S(faf)240 | 77B6/7 | Allelic to DNAprim |
| S(faf)244 | Chromosome 3 | No recombination with th cu sr e chromosome |
| Suppressors . | Location . | Comments . |
|---|---|---|
| S(faf)86 | 25D-30C | Homozygotes wild type |
| S(faf)184 | 88D2/3 | Allelic to UbcD1 |
| S(faf)196 | near th | Wild type in trans to Df |
| S(faf)240 | 77B6/7 | Allelic to DNAprim |
| S(faf)244 | Chromosome 3 | No recombination with th cu sr e chromosome |
The S(faf)86 and S(faf)196 mutations were localized imprecisely because it was discovered during the mapping process that these mutations are likely to be in genes without essential functions (the mutants are homozygous viable), which rendered them difficult to map physically. The S(faf)244 mutation could not be mapped meiotically as no recombination could be observed between this third chromosome and the mapping chromosome. The S(faf) mutations are not dominant suppressors of the sterility of fafFO8 females.
Suppressors of faf
| Suppressors . | Location . | Comments . |
|---|---|---|
| S(faf)86 | 25D-30C | Homozygotes wild type |
| S(faf)184 | 88D2/3 | Allelic to UbcD1 |
| S(faf)196 | near th | Wild type in trans to Df |
| S(faf)240 | 77B6/7 | Allelic to DNAprim |
| S(faf)244 | Chromosome 3 | No recombination with th cu sr e chromosome |
| Suppressors . | Location . | Comments . |
|---|---|---|
| S(faf)86 | 25D-30C | Homozygotes wild type |
| S(faf)184 | 88D2/3 | Allelic to UbcD1 |
| S(faf)196 | near th | Wild type in trans to Df |
| S(faf)240 | 77B6/7 | Allelic to DNAprim |
| S(faf)244 | Chromosome 3 | No recombination with th cu sr e chromosome |
The S(faf)86 and S(faf)196 mutations were localized imprecisely because it was discovered during the mapping process that these mutations are likely to be in genes without essential functions (the mutants are homozygous viable), which rendered them difficult to map physically. The S(faf)244 mutation could not be mapped meiotically as no recombination could be observed between this third chromosome and the mapping chromosome. The S(faf) mutations are not dominant suppressors of the sterility of fafFO8 females.
The S(faf) gene at 77B6/7 is DNAprim: To isolate the suppressor of faf locus at 77B6/7, genomic DNA adjacent to P{lacW}j10B2 was isolated by plasmid rescue (materials and methods). The DNA sequence of the cloned genomic DNA immediately adjacent to the P element was determined and compared to the Drosophila genomic and expressed sequenced tag DNA sequence databases (materials and methods). This analysis revealed that the P element is inserted in the 5′-untranslated region of the DNAprim gene. To test if the S(faf)240 and the l(3)77ABe mutations are also in the DNAprim gene, the DNA sequences of their DNAprim genes were determined (materials and methods). All of the DNAprim alleles in these mutants have suffered DNA lesions likely to result in loss of function (Figure 3). In addition, it was determined in genomic DNA blot experiments that one breakpoint of In(3L)78Cb1 is within 2.6 kb of the start of transcription of DNAprim (data not shown; see materials and methods). Finally, a DNAprim transgene expressed ubiquitiously using the Gal4/UAS system (Brand and Perrimon 1993) rescues the lethality of l(3)j10B2 homozygotes (see materials and methods). Thus, we conclude that mutations in DNAprim are suppressors of faf. We refer to S(faf)240, l(3)j10B2, and the three l(3)77ABe alleles as DNAprim alleles from here on (Table 2).
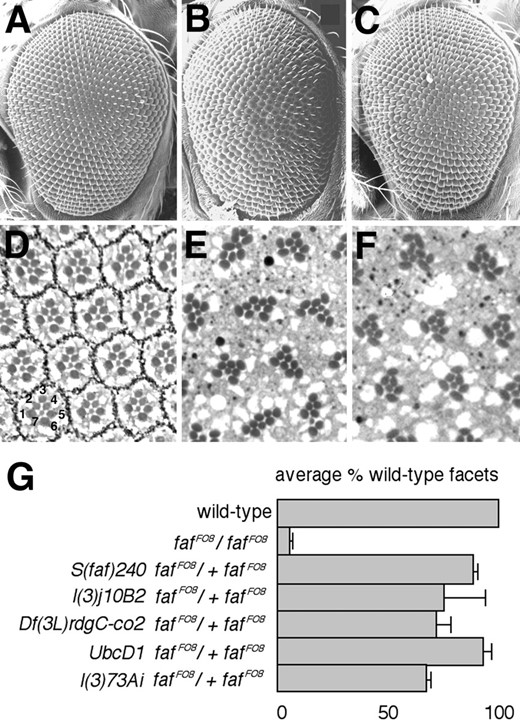
—Suppression of the faf mutant eye phenotype. Scanning electron micrographs (A-C) and tangential sections (D-F) of Drosophila adult eyes are shown. Anterior is to the left in A-C. The genotypes are as follows: (A and D) wild type; (B and E) faf FO8; and (C and F) S(faf)240 faf FO8/+ faf FO8. The numbers in D indicate photoreceptor cells R1-R7. The central R8 cell is not visible in this apical plane of section. (E) Each facet contains more than the normal number of photoreceptors. (G) A histogram showing the dominant suppressor of faf activity of three different DNAprim mutant alleles. For comparison, the extent to which UbcD1 and l(3)73Ai suppress faf FO8 is also shown (Huang et al. 1995; Wu et al. 1999). The average fraction of wild-type facets was calculated by scoring the number of wild-type and mutant facets in at least 100 facets in at least three different flies of each genotype. The error bars indicate standard deviations, which represent the variation between individuals of a given genotype.
Weak DNAprim mutants have eye defects: Eight of the 11 DNAprim alleles, including In(3L)78Cb1, are early lethal in trans to Dfr(3L)rdgC-co2 or S(faf)240 (Table 2). As In(3L)78Cb1 breaks within the coding region of DNAprim, the other 7 alleles are also likely to be null mutants. A weaker allele, DNAprimT2, is pupal lethal in trans to Dfr(3L)rdgC-co2 or S(faf)240. There is cell death in the eye anterior of late-stage pupae of these genotypes (Figure 4A). The weakest alleles are the homozygous viable group of imprecise excisions, exemplified by DNAprimj10B2Δ13-15 (Table 2). When homozygous or in trans to Df(3L)rdgC-co2, the DNAprimj10B2Δ13-15 alleles result in slightly roughened external eyes (Figure 4B). Patterning defects, either in the accessory cells (cone and pigment cells) of pupal eyes (Figure 4C) or in adult photoreceptors (Figure 4D), could not be detected in these flies. Thus, the external rough eyes are most likely due to subtle defects in cone cells, pigment cells, and/or bristles in the final stages of eye development; absent bristles and bristle misplacements are apparent in the adult eyes (Figure 4B).
The cell cycle is abnormal in DNAprim mutants: To understand the basis for the genetic interactions observed between faf and DNAprim, we asked whether the cell cycle is altered in these mutants. DNAprim mutants would be expected to have difficulty during S phase when DNA is replicated. If S phase is slower in these mutants, then more cells would be expected to be in S phase at any given time. During larval eye disc development, undifferentiated cells anterior to the morphogenetic furrow divide asynchronously and then arrest in G1 within the morphogenetic furrow prior to differentiation. No cell division occurs again until later in the eye disc, when cells that will give rise to three of the eight photoreceptor types and all of the other cell types undergo a single mitosis (Wolff and Ready 1993). By assaying BrdU incorporation with anti-BrdU antibodies (materials and methods), we visualized cells in S phase in larval eye discs of flies with a pupal lethal combination of DNAprim mutations, as well as wild-type and faf - eye discs. We found that BrdU incorporation in the faf - eye discs resembled wild type (Figure 5, A and B). By contrast, in the DNAprim mutants, more cells were in S phase at a given time (Figure 5C). As faf mutants do not have defects in S phase, the genetic interactions between DNAprim and faf most likely reflect indirect interactions between the functions of the two proteins.
Mutant DNAprim alleles
| Allele name (synonym) . | How induced . | Phenotype in trans to Df(3L)rdgC-co2 or DNAprimSfaf240 . | References . |
|---|---|---|---|
| DNAprimSfaf240 (S(faf)240) | EM | Early lethal | This article |
| DNAprimj10B2 (l(3)j10B2) | P{lacW}j10B2 | Early lethal | Spradling et al. (1999) |
| DNAprimJ1 (l(3)77ABeJ1) | EMS | Early lethal | Lukinova et al. (1999) |
| DNAprimL5 (l(3)77ABeL5) | EMS | Early lethal | Lukinova et al. (1999) |
| DNAprimT2 (l(3)77ABeT2) | EMS | Pupal lethal | Lukinova et al. (1999) |
| DNAprimj10B2Δ11 | P{lacW} hop-out | Early lethal | This article |
| DNAprimj10B2Δ12 | P{lacW} hop-out | Early lethal | This article |
| DNAprimj10B2Δ13 | P{lacW} hop-out | Viable; rough eyes | This article |
| DNAprimj10B2Δ14 | P{lacW} hop-out | Viable; rough eyes | This article |
| DNAprimj10B2Δ15 | P{lacW} hop-out | Viable; rough eyes | This article |
| In(3L)78Cb1 | X rays | Early lethal | Russell et al. (1996) |
| Allele name (synonym) . | How induced . | Phenotype in trans to Df(3L)rdgC-co2 or DNAprimSfaf240 . | References . |
|---|---|---|---|
| DNAprimSfaf240 (S(faf)240) | EM | Early lethal | This article |
| DNAprimj10B2 (l(3)j10B2) | P{lacW}j10B2 | Early lethal | Spradling et al. (1999) |
| DNAprimJ1 (l(3)77ABeJ1) | EMS | Early lethal | Lukinova et al. (1999) |
| DNAprimL5 (l(3)77ABeL5) | EMS | Early lethal | Lukinova et al. (1999) |
| DNAprimT2 (l(3)77ABeT2) | EMS | Pupal lethal | Lukinova et al. (1999) |
| DNAprimj10B2Δ11 | P{lacW} hop-out | Early lethal | This article |
| DNAprimj10B2Δ12 | P{lacW} hop-out | Early lethal | This article |
| DNAprimj10B2Δ13 | P{lacW} hop-out | Viable; rough eyes | This article |
| DNAprimj10B2Δ14 | P{lacW} hop-out | Viable; rough eyes | This article |
| DNAprimj10B2Δ15 | P{lacW} hop-out | Viable; rough eyes | This article |
| In(3L)78Cb1 | X rays | Early lethal | Russell et al. (1996) |
All known DNAprim mutant alleles are listed. The allele DNAprimJ9 is not listed here because by DNA sequence analysis we found that it is the same as DNAprimJ1 (Lukinova et al. 1999).
Mutant DNAprim alleles
| Allele name (synonym) . | How induced . | Phenotype in trans to Df(3L)rdgC-co2 or DNAprimSfaf240 . | References . |
|---|---|---|---|
| DNAprimSfaf240 (S(faf)240) | EM | Early lethal | This article |
| DNAprimj10B2 (l(3)j10B2) | P{lacW}j10B2 | Early lethal | Spradling et al. (1999) |
| DNAprimJ1 (l(3)77ABeJ1) | EMS | Early lethal | Lukinova et al. (1999) |
| DNAprimL5 (l(3)77ABeL5) | EMS | Early lethal | Lukinova et al. (1999) |
| DNAprimT2 (l(3)77ABeT2) | EMS | Pupal lethal | Lukinova et al. (1999) |
| DNAprimj10B2Δ11 | P{lacW} hop-out | Early lethal | This article |
| DNAprimj10B2Δ12 | P{lacW} hop-out | Early lethal | This article |
| DNAprimj10B2Δ13 | P{lacW} hop-out | Viable; rough eyes | This article |
| DNAprimj10B2Δ14 | P{lacW} hop-out | Viable; rough eyes | This article |
| DNAprimj10B2Δ15 | P{lacW} hop-out | Viable; rough eyes | This article |
| In(3L)78Cb1 | X rays | Early lethal | Russell et al. (1996) |
| Allele name (synonym) . | How induced . | Phenotype in trans to Df(3L)rdgC-co2 or DNAprimSfaf240 . | References . |
|---|---|---|---|
| DNAprimSfaf240 (S(faf)240) | EM | Early lethal | This article |
| DNAprimj10B2 (l(3)j10B2) | P{lacW}j10B2 | Early lethal | Spradling et al. (1999) |
| DNAprimJ1 (l(3)77ABeJ1) | EMS | Early lethal | Lukinova et al. (1999) |
| DNAprimL5 (l(3)77ABeL5) | EMS | Early lethal | Lukinova et al. (1999) |
| DNAprimT2 (l(3)77ABeT2) | EMS | Pupal lethal | Lukinova et al. (1999) |
| DNAprimj10B2Δ11 | P{lacW} hop-out | Early lethal | This article |
| DNAprimj10B2Δ12 | P{lacW} hop-out | Early lethal | This article |
| DNAprimj10B2Δ13 | P{lacW} hop-out | Viable; rough eyes | This article |
| DNAprimj10B2Δ14 | P{lacW} hop-out | Viable; rough eyes | This article |
| DNAprimj10B2Δ15 | P{lacW} hop-out | Viable; rough eyes | This article |
| In(3L)78Cb1 | X rays | Early lethal | Russell et al. (1996) |
All known DNAprim mutant alleles are listed. The allele DNAprimJ9 is not listed here because by DNA sequence analysis we found that it is the same as DNAprimJ1 (Lukinova et al. 1999).
—Mutations in EMS-induced DNAprim alleles. Amino acid sequence changes, derived from DNA sequence determination, in the putative DNA primase proteins produced by four different EMS-induced mutant alleles are shown. Two of the mutations are single nucleotide changes; T2 is GAG → AAG and L5 is CAG → TAG. Two of the mutations are deletions; J1 is an in-frame deletion of 27 bp and Sfaf240 is a deletion of 101 bp that results in a frameshift. The mutations in J1, L5, and Sfaf240 (strong alleles) are all in regions highly conserved in the yeast, human, and mouse genes (Huikeshoven and Cotterill 1999). T2 is a weak allele, and E138 is somewhat conserved (D in human and mouse, F in yeast; Huikeshoven and Cotterill 1999). In addition to the amino acid sequence we derived shown here, three translated amino acid sequences for Drosophila DNA primase are available: genomic clone AC017679 and cDNA clone GM13640 from the Drosophila Genome Project, and also the sequence derived by Huikeshoven and Cotterill (1999). Our translated sequence matches exactly with that derived from AC017679, except that AC017679 is missing 1 bp, which is present in the GM13640 sequence, that would interrupt the reading frame. We assume that this is an error in the AC017679 sequence. Our translated sequence differs from the translation of GM13640 in two amino acids: Y42 in our sequence is F in GM13640 and S49 in our sequence is G in GM13640. There are numerous differences between our sequence and that in Huikeshoven and Cotterill (1999). The GenBank accession number for the DNA sequence we report here is AF291873.
DISCUSSION
In a screen for dominant suppressors of the faf mutant eye phenotype, we identified mutations in the DNAprim gene, which encodes the large subunit (60 kD) of DNA primase, a component of DNA polymerase α. The precise function of the 60-kD primase subunit is unknown. The locations of the DNA lesions in several DNAprim mutants have been determined; this information could help to define functional domains of DNAprim protein. In addition, we find that Drosophila eye development is particularly sensitive to DNAprim activity and that in the developing eyes of flies with reduced levels of DNAprim activity, S phase of the cell cycle is slower relative to wild type.
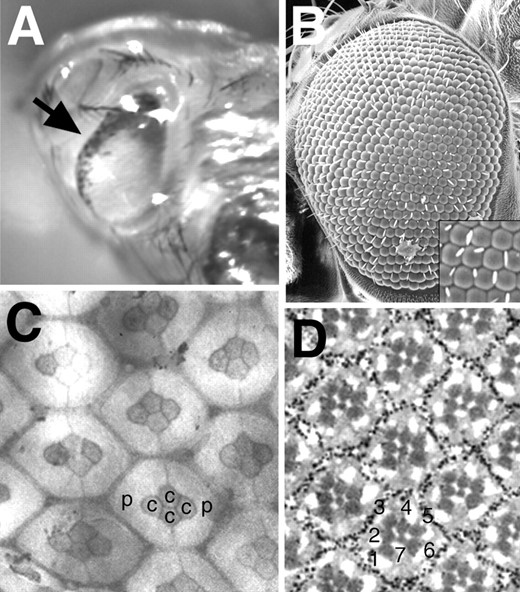
—Eye phenotypes of weak DNAprim mutants. (A) The eye of a late-stage pupa of the genotype DNAprimT2/DNAprimSfaf240 is shown. The arrow points to a region of cell death (black dots) at the eye anterior. (B) A scanning electron micrograph showing the slightly rough eye of a DNAprimj10B2Δ14/DNAprimSfaf240 adult. The inset is a magnification of a portion of the micrograph, showing that bristles, which should be present at alternate corners of the hexagons, are often missing or misplaced. (C) A pupal eye disc of the genotype DNAprimj10B2Δ13/DNAprimSfaf240 stained with cobalt sulfide is shown. The primary pigment cells (p), the cone cells (c), and the general shape of the hexagonal lattice surrounding them appear normally arranged. (D) A tangential section through an adult eye of the genotype DNAprimj10B2Δ13/Df(3L)rdgC-co2 is shown. The numbers indicate the photoreceptors R1-R7, which appear normally arranged.
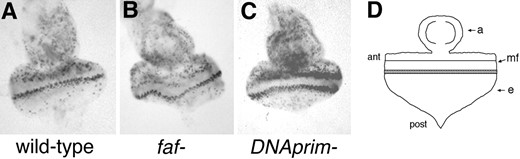
—BrdU incorporation in larval eye discs. Cells in S phase during a 2-hr period in third instar larval eye discs were visualized by BrdU incorporation (materials and methods). The genotypes of the discs are as follows: (A) wild type (w1118); (B) faf FO8/faf BX4; (C) DNAprimT2/DNAprimS240. The black dots are the cells that have incorporated BrdU. (D) A diagram of a third instar larval eye-antennal disc is shown; ant, anterior; post, posterior; mf, morphogenetic furrow; a, antennal portion of disc; e, eye portion of disc. In C, more cells have incorporated BrdU both anterior to the furrow and in the band posterior to the furrow. The discs shown are representative of all of the discs stained (∼20 of each genotype).
Why do DNAprim mutations suppress the faf mutant eye phenotype? To answer this question, we must consider the origins of the ectopic photoreceptors in faf mutant eyes and also the time and place in the larval eye disc in which faf normally functions.
The ommatidia of faf mutant eyes contain one or more photoreceptors in addition to the normal eight. The extra photoreceptors originate from the mystery cells, which are present in early facet preclusters (Fischer-Vize et al. 1992). The precursors to the first five photoreceptors (R8, R2/5, and R3/4) and one or more mystery cells emerge posterior to the morphogenetic furrow as preclusters of six or more cells, each surrounded by undifferentiated cells (Tomlinson and Ready 1987b; Wolff and Ready 1991). Normally, the mystery cells detach from the preclusters and go on to divide in the wave of mitosis posterior to the furrow, giving rise to the remainder of the photoreceptors (R1, R6, and R7) and to the accessory cells (cone, pigment, and bristle cells). In faf mutants, the mystery cells remain within the preclusters and become extra R3/4-like cells (Fischer-Vize et al. 1992) Although faf is required for inhibition of mystery cell neurogenesis, mosaic analysis as well as faf transgene complementation experiments indicate that faf is not required within the mystery cells nor within any photoreceptor precursors in the precluster (Fischer-Vize et al. 1992; Huang and Fischer-Vize 1996).
Normally, faf is expressed at high levels anterior to the morphogenetic furrow, where cells are undergoing mitosis, and then its expression shuts off within the furrow, just prior to precluster formation (Fischer-Vize et al. 1992). Although faf normally functions anterior to the furrow, later expression of faf in a subgroup of cells within the furrow is sufficient to rescue the faf mutant eye phenotype (Huang and Fischer-Vize 1996). The subgroup of cells that require faf expression are those cells that express the rough gene; these cells surround those that express the proneural gene atonal, which are destined to become the precluster cells (Dokucu et al. 1996). Thus, faf activity in the dividing cells anterior to the furrow facilitates a cell communication pathway that may occur anterior to or within the furrow. Perhaps when DNA primase levels are halved in heterozygous mutants, the cells anterior to the furrow remain in S phase longer, which changes subtly the output of the signaling pathway that faf modulates in such a way that the requirement for faf is largely overcome.
Mutations in cell cycle proteins have been isolated previously in genetic screens for mutants affecting the Sevenless receptor tyrosine kinase signaling pathway in eye development. For example, mutations in the Drosophila homolog of Saccharomyces cerevisiae cell cycle regulator cdc37 were identified as dominant enhancers of weak sevenless mutants (Cutforth and Rubin 1994). Also, mutations in the peanut gene, which encodes a protein similar to yeast bud neck filaments and functions in cytokinesis, were identified as dominant enhancers of the phenotype of weak seven in absentia mutants (Neufeld and Rubin 1994); seven in absentia is a nuclear protein that functions downstream of Sevenless to target a transcriptional repressor for proteolysis (Carthew and Rubin 1990; Li et al. 1997; Zhang et al. 1998). In these cases, the genetic interactions may also be an indirect consequence of cells that are receiving an inductive signal remaining longer in a particular phase of the cycle than they would normally. The results presented here underscore the connection between the cell cycle and signaling pathways that govern cell determination. A less likely possibility is that the Cdc37, Peanut, and/or the 60-kD subunit of DNA primase are bifunctional proteins, with distinct functions in the cell cycle and in signal transduction.
Acknowledgement
We thank the University of Texas at Austin DNA Analysis Facility for DNA sequence analysis; John Mendenhall for performing the scanning electron microscopy; Erin Overstreet for the pupal eye disc preparations; and Nina Lukinova, Mark Fortini, Kevin Moses, and the Bloomington Drosophila Stock Center for flies. The first two authors contributed equally to this work.
Footnotes
Communicating editor: K. Anderson
LITERATURE CITED