-
PDF
- Split View
-
Views
-
Cite
Cite
Scott L Page, Kim S McKim, Benjamin Deneen, Tajia L Van Hook, R Scott Hawley, Genetic Studies of mei-P26 Reveal a Link Between the Processes That Control Germ Cell Proliferation in Both Sexes and Those That Control Meiotic Exchange in Drosophila, Genetics, Volume 155, Issue 4, 1 August 2000, Pages 1757–1772, https://doi.org/10.1093/genetics/155.4.1757
Close - Share Icon Share
Abstract
We present the cloning and characterization of mei-P26, a novel P-element-induced exchange-defective female meiotic mutant in Drosophila melanogaster. Meiotic exchange in females homozygous for mei-P261 is reduced in a polar fashion, such that distal chromosomal regions are the most severely affected. Additional alleles generated by duplication of the P element reveal that mei-P26 is also necessary for germline differentiation in both females and males. To further assess the role of mei-P26 in germline differentiation, we tested double mutant combinations of mei-P26 and bag-of-marbles (bam), a gene necessary for the control of germline differentiation and proliferation in both sexes. A null mutation at the bam locus was found to act as a dominant enhancer of mei-P26 in both males and females. Interestingly, meiotic exchange in mei-P261; bamΔ86/+ females is also severely decreased in comparison to mei-P261 homozygotes, indicating that bam affects the meiotic phenotype as well. These data suggest that the pathways controlling germline differentiation and meiotic exchange are related and that factors involved in the mitotic divisions of the germline may regulate meiotic recombination.
MEIOSIS is the component of gametogenesis responsible for the segregation of homologous chromosomes into haploid gametes. In most organisms, including humans and Drosophila melanogaster females, the proper segregation of homologs during meiosis is facilitated by the formation of reciprocal genetic exchanges along the lengths of the chromosomes. Chromosomes with no exchange or with abnormally positioned exchanges are segregated with decreased fidelity (Koehler et al. 1996; Lamb et al. 1996). Although organisms have evolved systems to ensure the segregation of nonexchange chromosomes, these systems can usually effectively handle only one or two pairs of nonexchange homologs (Hawley and Theurkauf 1993), so the proper regulation of recombination is essential for reliably producing euploid offspring.
In Drosophila oogenesis, meiotic recombination occurs in the context of a cyst of 16 interconnected germline cells. Germline cyst development begins with the asymmetric division of a germline stem cell (GSC) to produce a GSC daughter and a cystoblast (for a review of germline cyst development, see de Cuevas et al. 1997). The cystoblast then undergoes four mitotic divisions with incomplete cytokinesis. The residual connections between the cells, called ring canals, are inherited by one of the daughters after each division, such that after the four mitoses, two cells each retain four ring canals each. As the gonial mitotic divisions progress, a structure called a fusome is constructed within the cyst. Fusome formation begins with the asymmetric inheritance of spectrosome material into the cystoblast (Deng and Lin 1997). The fusome eventually takes on a branched structure, entering each cell of the cyst through the ring canals (de Cuevas and Spradling 1998). Although the process of oocyte selection is unknown, one of the two cells with four ring canals becomes the oocyte and progresses through meiosis. The remaining cells within the cyst take on a nurse cell fate. The result is a cyst containing one oocyte and 15 nurse cells.
A similar process of cyst development occurs in males (reviewed by Lindsley and Tokuyasu 1980), where GSCs divide to produce a daughter GSC and a primary spermatogonial cell. The primary spermatogonial cell then divides four times to form a cyst of 16 interconnected primary spermatocytes. In contrast to oogenesis, no meiotic recombination occurs, and all 16 primary spermatocytes within the cyst go through both meiotic divisions synchronously to produce 64 spermatids, which then differentiate into mature spermatozoa.
During meiotic prophase in females, chromosomes condense and pair to form bivalents. Synapsis of homologous chromosomes culminates in the formation of synaptonemal complex (SC) along the lengths of the chromosomes (Carpenter 1975a; 1979b). Mutations in the Drosophila c(3)G gene cause both a complete absence of SC formation and a lack of exchange (Smith and King 1968; Hall 1972). SC formation appears to be normal in mei-W68 and mei-P22, although meiotic recombination and gene conversion are abolished (McKim et al. 1998). Moreover, cloning of the mei-W68 locus showed that it encodes a homolog of the yeast gene SP011, which is responsible for the first step of recombination, double-strand break formation, in budding yeast (McKim and Hayashi-Hagihara 1999). Thus, SC is apparently necessary for, but not dependent upon, the initiation of meiotic recombination in Drosophila.
In wild-type Drosophila females, meiotic exchanges are distributed in a nonrandom pattern along the euchromatin of each chromosome arm (with the exception of the small fourth chromosome; Lindsley and Sandler 1977). Crossing over is generally reduced in the centromeric and telomeric regions and entirely absent in the heterochromatin. Meiotic exchange can be assayed genetically in crosses with marked chromosomes, or cytologically by the analysis of the timing and number of recombination nodules (Carpenter 1975b; 1979a,b).
Studies of meiotic mutants in Drosophila have demonstrated that multiple levels of control of recombination exist. Because c(3)G, mei-W68, and mei-P22 completely suppress exchange and gene conversion, they are known as recombination-null mutants (McKim et al. 1998). A greater number of Drosophila mutants decrease exchange without eliminating it. These mutants were initially classified on the basis of their effects on the distribution of exchanges. Carpenter and Sandler (1974) expected that mutants that affect the process of exchange itself would produce a uniform decrease in the frequency of exchanges without altering their distribution along a chromosome arm. Mutants that affect the distribution of exchanges were also identified. These mutants, when compared to wild type, bear similar patterns of residual exchanges. These patterns are characterized by a preference for proximal exchanges and an increasing suppression of exchange that increases toward the distal tips of the chromosomes. The term “polar” was used to describe this distribution. Mutants that redistribute exchanges in a polar fashion were suggested to correspond to genes necessary for a set of preconditions required for any given chromosomal interval to be competent to form a crossover and thus were called “precondition mutants” (Sandler et al. 1968; Carpenter and Sandler 1974). Differential probabilities of certain chromosomal regions for meeting the necessary preconditions for exchange were thought to lead to the nonrandom pattern of crossover distribution observed for each chromosome arm in wild type. Precondition mutants were expected to be defective for setting up the preconditions for exchange, thus resulting in the abnormal crossover distribution (Carpenter and Sandler 1974).
A new exchange-defective female meiotic mutant, mei-P26, has been cloned and characterized. mei-P26 causes a polar decrease in recombination, similar to many other exchange-defective mutants. The mei-P26 gene encodes a novel member of the RING finger B-box coiled coil (RBCC) family of proteins. Phenotypic analyses of additional loss-of-function alleles of this locus show that this gene also affects both male and female gametogenesis. Furthermore, a null mutation in bag-of-marbles (bam), which affects both male and female gametogenesis, acts as a dominant enhancer of mei-P26 in both males and females. The effects of mei-P26 mutations on germline differentiation and meiosis, as well as the genetic interaction of mei-P26 with bam, suggest that mei-P26 either has multiple roles in the germline, or that the proper regulation of germline cyst formation is required for the normal levels of meiotic exchange.
MATERIALS AND METHODS
Drosophila stocks and culture: The genetic markers and chromosomes used in this study are described in Lindsley and Zimm (1992), FlyBase (1999), and Sekelsky et al. (1999). Flies were reared on standard cornmeal-molasses-dextrose medium at 25°. For egg counts, eggs were collected on grapejuice plates (Ashburner 1989) as described by Wieschaus and Nüsslein-Volhard (1986).
Molecular techniques: Plasmid rescue was performed as described previously (Ashburner 1989). Library screening was done essentially as described by Sambrook et al. (1989) using Church hybridization and wash buffers (Church and Gilbert 1984). An ABI 377XL (Applied Biosystems, Foster City, CA) was used to perform DNA sequencing.
Genetic analyses: The assay for measuring X and 4th chromosome nondisjunction in females crossed to YSX · YL, v f B/0; C(4)RM, ci eyR/0 males is described by Sekelsky et al. (1999). The calculation of XX ↔ 44 nonhomologous disjunction frequency is described by Hawley et al. (1993).
Recombination along the left arm of chromosome 2 was scored among the progeny of females of the genotype X/X; net ho dp Sp b pr cn/+ + + + + + + crossed to +/Y; net ho dp b pr cn/net ho dp b pr cn males. For most crosses, only female progeny were scored, because the presence of w on one or both of the X chromosomes would obscure the scoring of pr and cn in male progeny, because of the epistasis of w to these two eye color markers. Exchange rank frequencies, the frequencies of tetrads bearing zero, one, two, or three exchanges, were calculated as described by Weinstein (1936).
A screen for additional alleles of mei-P26: New alleles of mei-P26 were generated by exposing the P{lacW} element responsible for mei-P261 to a source of P transposase and isolating imprecise excisions and local hops of the P element. Non-Stubble (Sb) female progeny of mei-P261/y+Y; TMS, Sb Δ2-3/+; spapol crossed to FM7w; +/+; spapol were selected on the basis of loss or change in eye color generated by the mini-white+ (w+mC) gene encoded by P{lacW}. For each of the isolated X chromosomes, denoted by an asterisk, y w mei-P26*/FM7w; spapol females were crossed to FM7w/y+Y; spapol males to establish stocks. Stocks in which the eye color phenotype failed to segregate with the X chromosome (i.e., jumps to the autosomes) were discarded. To assay X chromosome nondisjunction, homozygous y w mei-P26*; spapol females were selected from the stocks and crossed to FM7w/y+Y; spapol males. The normal progeny from this cross are yellow females and yellow+ males. Exceptional progeny resulting from diplo-X or nullo-X ova are recovered as yellow+ (y w mei-P26*/y w mei-P26*/y+Y) females and yellow (FM7w/0) males.
Germline transformation: The transformation construct P{w+mC; N1} was constructed by inserting a restriction fragment containing ~13 kb of genomic DNA (see Figure 2) into the vector pW8 (Klemenz et al. 1987). pP{w+mC; N1} DNA was purified (QIAGEN, Valencia, CA) and resuspended in injection buffer at a concentration of 1 μg/μl with the transposase source pTurboΔ2-3 (FlyBase 1999) at 0.3 μg/μl and microinjected into w1118 embryos (Spradling and Rubin 1982; Spradling 1986).
Cytology: Ovaries were dissected in phosphate-buffered saline (PBS) or modified Robb's medium (Theurkauf 1994) and fixed in 4% formaldehyde, 100 mm sodium cacodylate, pH 7.2, 100 mm sucrose, 40 mm sodium acetate, 10 mm EGTA for 5 min. They were then rinsed in PBS containing 0.1% Tween 20 and stained with 0.5 μg/ml DAPI (4′,6-diamidino-2-phenylindole) in PBS for 5 min. After three washes with PBS + 0.1% Tween 20, the ovarioles were teased apart and mounted in 90% glycerol containing p-phenylenediamine at a concentration of 1 mg/ml. Testes were dissected in testis buffer (Ashburner 1989) and viewed unfixed using differential interference contrast (DIC) optics. All microscopy and photography was performed on a Zeiss Axioplan microscope equipped for fluorescence and DIC microscopy. Color photographs were scanned into an Apple Power Macintosh 7500 computer using a Nikon LS-1000 35-mm film scanner, converted to grayscale using Adobe Photoshop 5.0.2, and printed using a Tektronix Phaser 440 dye sublimation printer.
RESULTS
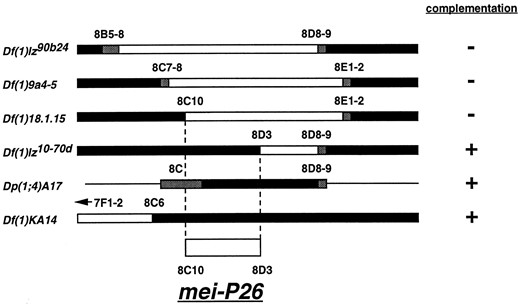
Molecular characterization of the mei-P26 interval in region 8C: The first allele of mei-P26, denoted mei-P261, was isolated during a P-element screen for mutants that affect female meiosis in D. melanogaster (Sekelsky et al. 1999). In situ hybridization of a P-element probe to mei-P261 larval salivary gland polytene chromosomes revealed the presence of a single P{lacW} insertion in region 8 of the Drosophila polytene map (Sekelsky et al. 1999). Three deficiencies in this region, Df(1)lz90b24, Df(1)9a4-5, and Df(1)18.1.15, failed to complement mei-P261 (see below). A fourth deficiency, Df(1)lz10-70d, complemented mei-P261 and positioned the gene in region 8C10-8D3 (Figure 1).
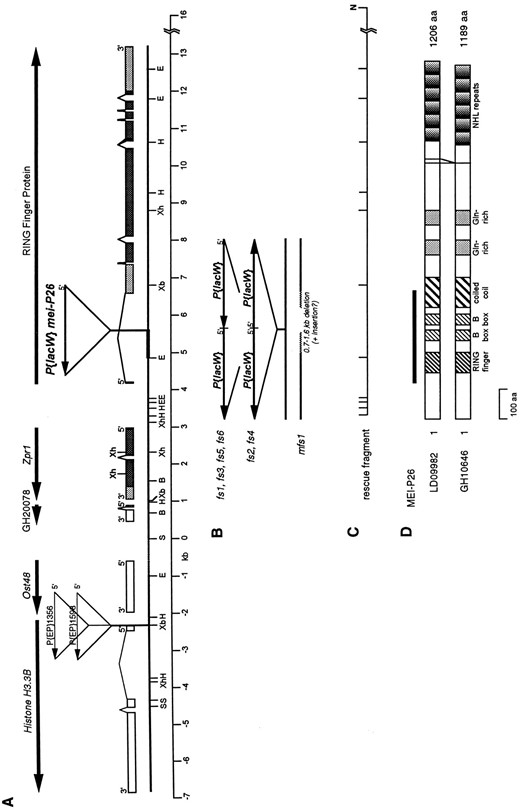
We have characterized the region surrounding the P{lacW} insertion in mei-P261. An EcoRI fragment containing ~740 base pairs (bp) of DNA flanking the 3′ end of the P{lacW} insertion was isolated by plasmid rescue. Two overlapping lambda clones representing the mei-P26 locus were identified by using the rescue fragment as a probe to screen a Drosophila genomic library. Portions of these clones were then subcloned into pBluescript and sequenced. Sequencing and restriction mapping allowed the construction of a map representing the genomic region surrounding the mei-P261 insertion (Figure 2A). cDNA library screening using the genomic subclones and BLAST searches (Altschul et al. 1997) using their sequences led to the identification of five transcription units in the region. Two were previously cloned Drosophila genes; one was found based on homology to a mammalian gene; and two, one being the mei-P26 gene, were discovered based on homology to Drosophila expressed sequence tags (ESTs). We will first describe the neighboring genes and then in the following section we will describe the mei-P26 transcription unit.
The 5′ ends of the genes encoding the oligosaccharyl-transferase 48-kD subunit (Ost48) and histone H3.3B (His3.3B) are located ~6 and 8 kb from the 3′ end of the P element, respectively (Figure 2A). These genes were previously cloned and characterized and are known to be tightly linked (Akhmanova et al. 1995; Stagljar et al. 1995). Two P{EP} insertions have been identified by the Berkeley Drosophila Genome Project (BDGP) close to the His3.3B transcription start site, in the 5′ flanking sequence adjacent to the His3.3B transcription start site (Rørth et al. 1998; E. J. Rehm and G. M. Rubin, unpublished results). These have both been mapped to 8C15-16 by in situ hybridization (T. Laverty, personal communication), which agrees well with the deficiency mapping of mei-P26.
The mei-P261 P element was found to be 2.6 kb upstream of a region showing strong homology to the mouse and human ZPR1 genes (Galcheva-Gargova et al. 1996, 1998). In mammalian cells, the ZPR1 protein binds intracellularly to the epidermal growth factor receptor (EGFR), and, upon ligand binding to the EGFR, ZPR1 accumulates in the nucleus, where it appears to regulate nucleolar function in proliferating cells by localizing to the nucleolus and binding to translation elongation factor 1α (Galcheva-Gargova et al. 1996, 1998; Gangwani et al. 1998). A total of 13 cDNAs for this gene were isolated by screening a Drosophila ovary library (Stroumbakis et al. 1994) using a genomic fragment containing this region. Sequencing of the longest of these indicates that this gene is expected to encode a protein of 457 amino acids with 49% identity and 67% similarity to human ZPR1. The ZPR1 protein has a bipartite structure that is conserved throughout eukaryotes and consists of two repeats of a zinc finger and an associated sequence (Galcheva-Gargova et al. 1996, 1998; Gangwani et al. 1998). Comparison of the genomic and cDNA sequences for the Drosophila homolog indicates a single intron located approximately in the middle of the protein coding region, separating the two repeats (Figure 2A).
A BLAST search of the BDGP EST database using sequence located between the Zpr1 and Ost48 genes revealed a high scoring match to one EST, GH20078 (Berkeley Drosophila Genome Project/HHMI EST
Deficiency and duplication mapping of mei-P26 positions the mutation between polytene map positions 8C10 and 8D3 on the X chromosome. Open boxes represent chromosomal regions that are deleted in the deficiencies indicated, solid boxes represent chromosomal regions that are present, and shaded boxes represent ambiguities from the mapping of deficiency and duplication breakpoints.
Project, unpublished results). This 378-bp cDNA appears to represent a spliced transcript oriented in the same direction as Zpr1 and Ost48. The longest open reading frame in this cDNA is 77 amino acids long and is predicted to encode an 8.4-kD polypeptide with no homologies to any known proteins.
The mei-P26 transcription unit: Sequence flanking the 5′ end of the mei-P261 P element was found to match a 5′ EST in BLAST searches of the BDGP database. The cDNA corresponding to the EST, LD09982, was sequenced and the genomic structure of the gene was determined (Figure 2A). The P element was found to be located within the 2.4-kb first intron of this gene, and P-element excision data and transgenic rescue experiments (shown below) confirm that this transcription unit does indeed correspond to the mei-P26 gene.
A total of three probable full-length (GH10646, LD09982, and LD30261) and two partial cDNAs (HL02723 and LD34505) for this gene have been identified among ESTs from the BDGP (Berkeley Drosophila Genome Project/HHMI EST Project, unpublished results). One of these, GH10646, was sequenced as part of the BDGP (GenBank accession no. AF145661), and we sequenced a second, LD09982. The full-length sequencing of these cDNAs revealed two sequence differences between GH10646 and LD09982. These sequence differences can both be explained by alternative splicing. Analysis of the LD09982 sequence predicts a novel 131-kD protein of 1206 amino acids. Interestingly, the GH10646 sequence contains a 51-bp in-frame deletion as the result of one of the alternative splice sites, and is expected to encode a slightly smaller protein of 1189 amino acids and 129.5 kD.
The proteins encoded by both of these cDNAs are expected to contain a RING finger, followed by two B-box motifs and a region predicted to form a coiled coil structure. RING fingers and B-boxes are cysteinerich zinc-binding protein motifs thought to be involved in interactions with proteins or nucleic acids (Borden et al. 1995, 1996; Cao et al. 1997, 1998). Two glutaminerich stretches reside near the middle of the protein, which may also form coiled coils. The C-terminal region of the protein bears six copies of a repeated motif called an NHL repeat (Slack and Ruvkun 1998). NHL repeats are similar to WD repeats in length and predicted structure and may be involved in protein-protein interactions (Fridell et al. 1995; El-Husseini and Vincent 1999). The 17-amino acid region missing from the 1189-amino acid version of the protein is located between the glutamine-rich domains and the NHL repeats and does not overlap with any of the recognizable motifs (Figure 2D). The portion of MEI-P26 containing the RBCC domain is most similar to the TIF1 (transcriptional intermediary factor 1) family of nuclear receptor mediator proteins (TIF1α, TIF1β, TIF1γ; Le Douarin et al. 1995, 1996; Venturini et al. 1999), with ~27% identity and 47% similarity. However, the C-terminal portions of the TIF1 proteins each contain PHD finger and bromodomain motifs, while MEI-P26 contains six NHL repeats.
Isolation of more severe alleles: Several lines of evidence indicate that the original mei-P261 allele is a hypomorph (see below). To isolate stronger alleles of mei-P26, the P element was mobilized using Δ2-3 transposase and putative excisions or local hops were selected on the basis of eye color. Full or partial excisions, including imprecise excisions, of P{lacW} that deleted the mini-white+ gene were expected to result in a total loss of mini-white+ expression (white eyes). Local hops of the P element were expected to produce changes in the mini-white+ expression level, resulting in a change in eye color, due to a change in P{lacW} copy number or position effects. For each X chromosome recovered, a stock was established and X chromosome nondisjunction was assayed in females homozygous for the X chromosome.
The mei-P26 locus. (A) Molecular map of the genomic region near mei-P26. The horizontal line shows a partial restriction map of the region surrounding the mei-P261 P{lacW} insertion. The sequence on the left, including the His3.3B and Ost48 genes, was previously published (accession nos. X81207 and X81999; Akhmanova et al. 1995; Stagljar et al. 1995). The dashed line signifies ~1 kb of unsequenced genomic DNA. The thickened line indicates the fragment isolated by plasmid rescue. Boxes above the line indicate exons for each gene except Ost48, for which the genomic structure was not determined. The positioning of Ost48 on this map is approximate. For Zpr1 and the RING finger gene, protein coding sequence is shaded dark gray, and untranslated sequence is shaded light gray. The 5′ to 3′ orientations of the genes and P elements are shown by arrows. B, BamHI; E, EcoRI; H, HindIII; S, SalI; Xb, XbaI; Xh, XhoI. (B) Mutations in new mei-P26 alleles. The mutations in the mei-P26fs1, mei-P26fs2, mei-P26fs3, mei-P26fs4, mei-P26fs5, mei-P26fs6, and mei-P26mfs1 alleles are shown corresponding to the map in A. (C) Restriction fragment used for transformation rescue construct P{w+mC; N1} is shown corresponding to the map in A. N, NotI (site from lambda vector). (D) Schematic representation of the proteins encoded by mei-P26 cDNAs LD09982 and GH10646, showing the 17-amino acid difference in the C-terminal region resulting from alternative splicing of the transcript. The line above the proteins represents the region of similarity to the TIF1 family.
Of 44 white-eyed excisions isolated, 37 completely reverted the mei-P261 phenotype, while 3 retained an X nondisjunction frequency of 2–4%, and 3 additional excisions retained a nondisjunction phenotype similar to mei-P261 (8–17% X nondisjunction). PCR analysis of these latter excisions showed small internal deletions of P{lacW} sequence, while deletions of most or all of the P{lacW} sequence were evident in the revertants (data not shown). These results demonstrate that the P-element insertion is indeed responsible for the mutant phenotype. One excision chromosome was a recessive lethal, apparently due to a deletion of the P{lacW} insert and flanking DNA on both sides. This chromosome fails to complement mei-P261, but its lethality is likely due to the deletion of one or more essential genes near mei-P26, rather than mei-P26 itself, since the lethality is not rescued by a transgenic construct that contains the wild-type mei-P26 transcription unit (described below).
A total of 20 X chromosomes were recovered with a change in eye color, representing putative local hops of the P element. Homozygotes for 6 of these produced no progeny in multiple crosses. All heteroallelic combinations of these chromosomes were also sterile in females. While these chromosomes are fertile when placed over mei-P261, the fertility of these flies is reduced, and all 6 of these failed to complement the mei-P261 nondisjunction phenotype. mei-P26fs/mei-P261 females showed higher levels of nondisjunction than mei-P261 homozygotes assayed contemporaneously (data not shown). The noncomplementation with mei-P261, and with each other, suggested that these chromosomes bear novel alleles of mei-P26, and they were given the allele designations fs1, fs2, fs3, fs4, fs5, and fs6.
PCR analysis of the P{lacW} in these alleles revealed that in each case two partial or full copies of P{lacW} were now present in the first intron of the RING finger gene. In four cases, there had been a tandem duplication of the P element, and in two of these, an inverted duplication was found (Figure 2B). The insertion of a P element into the first intron of this gene may compromise its expression, leading to a meiotic phenotype, as in the original mei-P261 allele. Addition of a second copy of the P element may further compromise expression, leading to the more severe female sterility phenotype.
A male sterile allele of mei-P26: Unexpectedly, a male and female sterile allele, mei-P26mfs1, was also recovered in the screen for additional mei-P26 mutants. Crosses using males bearing this chromosome paired with virgin females produced no progeny. This chromosome also failed to complement mei-P261, producing ~30% X nondisjunction when in heteroallelic combination with mei-P261. A duplication of the mei-P26 region on chromosome 4, Dp(1;4)A17 (Santamaria and Randsholt 1995), rescued the infertility, suggesting that a wild-type copy of the mei-P26 gene could complement the male sterility. mei-P26mfs1/Y; Dp(1;4)A17/spapol males were used to generate females homozygous for the male sterile chromosome. Homozygotes for this chromosome were completely sterile, although homozygote females bearing Dp(1;4)A17 were fertile.
Although this allele displays an increase in eye color relative to the original mei-P261 allele, PCR analysis of the P{lacW} indicated a deletion of the P-element ends and 0.7–1.6 kb of DNA flanking both sides of the original insertion site (Figure 2B). This may be in addition to a more complex rearrangement, however, possibly involving the insertion of sequence, since the deleted region could not be spanned by PCR from primers flanking the deleted sequence. P{lacW} sequence may remain, as evidenced by mini-white+ expression, which maps to the same interval of the X chromosome as mei-P26 (ct-m). Male sterility was an unexpected phenotype, which is not seen in any of the other mei-P26 alleles.
A single rescue construct rescues all three phenotypes: To confirm that the mutations in the RING finger gene were indeed responsible for the phenotypes observed in the mei-P26 alleles, a construct containing ~13 kb of genomic DNA, including this gene, 650–850 bp of 5′ flanking sequence, and ~3 kb of 3′ flanking sequence, in the P-element transformation vector pW8 was microinjected into embryos. Four independent transgenic lines bearing this construct were established. In tests for phenotypic rescue, the construct restored levels of nondisjunction and recombination to wild-type levels (Table 1). The slight deviation of the rescued phenotype from wild type may reflect either differences in genetic background, or incomplete rescue by the transgene due to noninclusion of all regulatory sequences or a position effect on expression. Heterozygosity for the rescue construct also rescued the sterility phenotype of mei-P26fs1 and both the male and female sterility of mei-P26mfs1.
Analysis of nondisjunction in mei-P26: Data for X and 4th chromosome segregation in mei-P261 females are shown in Table 2. Homozygous mei-P261 females displayed 17.3% X chromosome nondisjunction and 8.4% 4th chromosome nondisjunction. These nondisjunction frequencies were compared to those for females hemizygous for mei-P261 using deficiencies that fail to complement this gene. Females of the genotype mei-P261/Df(1)18.1.15 show an increase in nondisjunction frequency in comparison to mei-P261 homozygotes (Table 2) and a concomitant decrease in fertility. Df(1)9a4-5 and Df(1)lz90b24 showed an even more profound effect on fertility when heterozygous with mei-P261. From these data we may conclude that the original mei-P261 allele is hypomorphic. Too few progeny are produced by mei-P261/Df(1)9a4-5 or mei-P261/Df(1)lz90b24 females to accurately measure nondisjunction or recombination.
Testing of X and 4th chromosome nondisjunction in mei-P26fs1/mei-P261 females revealed a more severe phenotype than is seen in mei-P261 homozygotes. Nondisjunction of both the X and 4th chromosome is increased
Rescue of mei-P26 phenotypes with P{w+mC; N1} rescue construct on chromosome 3
| Genotype . | Phenotype assayed . | − rescue (+/+) (%) . | + rescue (P{w+mC; N1}/+) (%) . |
|---|---|---|---|
| y w mei-P26/y w mei-P26 | X nondisjunction | 14.1 | 0.74 |
| 4 nondisjunction | 7.5 | 1.11 | |
| y w mei-P26 m f/y w mei-P26 | m-f map distance | 1.7 | 16.0 |
| y w mei-P26fs1 m f/y w mei-P26 | m-f map distance | 1.3 | 14.1 |
| y w mei-P26fs1/y w mei-P26u | Fertility | Sterile | Fertile |
| Ovary morphology | Tumors | No tumors | |
| y w mei-P26mfs1/y w mei-P26mfs1 | Fertility | Sterile | Fertile |
| Ovary morphology | Tumors | No tumors | |
| y w mei-P26mfs1/y+ Y | Fertility | Sterile | Fertile |
| Genotype . | Phenotype assayed . | − rescue (+/+) (%) . | + rescue (P{w+mC; N1}/+) (%) . |
|---|---|---|---|
| y w mei-P26/y w mei-P26 | X nondisjunction | 14.1 | 0.74 |
| 4 nondisjunction | 7.5 | 1.11 | |
| y w mei-P26 m f/y w mei-P26 | m-f map distance | 1.7 | 16.0 |
| y w mei-P26fs1 m f/y w mei-P26 | m-f map distance | 1.3 | 14.1 |
| y w mei-P26fs1/y w mei-P26u | Fertility | Sterile | Fertile |
| Ovary morphology | Tumors | No tumors | |
| y w mei-P26mfs1/y w mei-P26mfs1 | Fertility | Sterile | Fertile |
| Ovary morphology | Tumors | No tumors | |
| y w mei-P26mfs1/y+ Y | Fertility | Sterile | Fertile |
Rescue of mei-P26 phenotypes with P{w+mC; N1} rescue construct on chromosome 3
| Genotype . | Phenotype assayed . | − rescue (+/+) (%) . | + rescue (P{w+mC; N1}/+) (%) . |
|---|---|---|---|
| y w mei-P26/y w mei-P26 | X nondisjunction | 14.1 | 0.74 |
| 4 nondisjunction | 7.5 | 1.11 | |
| y w mei-P26 m f/y w mei-P26 | m-f map distance | 1.7 | 16.0 |
| y w mei-P26fs1 m f/y w mei-P26 | m-f map distance | 1.3 | 14.1 |
| y w mei-P26fs1/y w mei-P26u | Fertility | Sterile | Fertile |
| Ovary morphology | Tumors | No tumors | |
| y w mei-P26mfs1/y w mei-P26mfs1 | Fertility | Sterile | Fertile |
| Ovary morphology | Tumors | No tumors | |
| y w mei-P26mfs1/y+ Y | Fertility | Sterile | Fertile |
| Genotype . | Phenotype assayed . | − rescue (+/+) (%) . | + rescue (P{w+mC; N1}/+) (%) . |
|---|---|---|---|
| y w mei-P26/y w mei-P26 | X nondisjunction | 14.1 | 0.74 |
| 4 nondisjunction | 7.5 | 1.11 | |
| y w mei-P26 m f/y w mei-P26 | m-f map distance | 1.7 | 16.0 |
| y w mei-P26fs1 m f/y w mei-P26 | m-f map distance | 1.3 | 14.1 |
| y w mei-P26fs1/y w mei-P26u | Fertility | Sterile | Fertile |
| Ovary morphology | Tumors | No tumors | |
| y w mei-P26mfs1/y w mei-P26mfs1 | Fertility | Sterile | Fertile |
| Ovary morphology | Tumors | No tumors | |
| y w mei-P26mfs1/y+ Y | Fertility | Sterile | Fertile |
(Table 2). However, X chromosome segregation is more greatly affected in mei-P261/Df than in mei-P261/mei-P26fs1, suggesting that even the mei-P26fs1 allele may not be fully null.
At least the majority of the misbehavior of the X chromosome in mei-P26 mutants appears to be the result of nondisjunction rather than chromosome loss, as shown by comparing the frequencies of nullo-X and diplo-X exceptions. Chromosome loss would be expected to produce an excess of nullo-X exceptions. However, mei-P26 mutants have approximately equal frequencies of nul-lo-X and diplo-X exceptions, showing that nondisjunction is the primary cause of aberrant X chromosome segregation.
Two lines of evidence suggest that the X chromosome nondisjunction induced by mei-P26 mutations is due to the failed segregation of nonexchange tetrads. First, as noted by Baker and Hall (1976), the frequency of X chromosome nondisjunction in recombination-defective mutants is correlated with the cube of E0, the frequency of nonexchange tetrads. E0 is calculated from recombination data using the method of Weinstein (1936).
X and 4th chromosome segregation in mei-P26 mutants
| Gamete types . | Maternal genotype . | |||
|---|---|---|---|---|
| Maternal . | Paternal . | mei-P261 . | mei-P261/mei-P26fs1 . | mei-P261/Df(1)18.1.15 . |
| Normal | ||||
| X; 4 | XY; 44 | 658 | 124 | 155 |
| X; 4 | 0; 44 | 884 | 139 | 173 |
| X nondisjunction | ||||
| 0; 4 | XY; 44 | 49 | 14 | 34 |
| XX; 4 | 0; 44 | 63 | 15 | 25 |
| 4 nondisjunction | ||||
| X; 0 | XY; 44 | 15 | 9 | 0 |
| X; 0 | 0; 44 | 14 | 14 | 2 |
| X; 44 | XY; 0 | 11 | 8 | 10 |
| X; 44 | 0; 0 | 11 | 10 | 0 |
| X; 4 nondisjunction | ||||
| 0; 0 | XY; 44 | 30 | 5 | 9 |
| XX; 44 | 0; 0 | 9 | 4 | 10 |
| 0; 44 | XY; 0 | 7 | 4 | 1 |
| XX; 0 | 0; 44 | 9 | 2 | 0 |
| Total progeny | 1760 | 348 | 419 | |
| Adjusted total | 1927 | 392 | 498 | |
| % nullo-X | 8.9 | 11.7 | 17.7 | |
| % diplo-X | 8.4 | 10.7 | 14.1 | |
| Total % X nondisjunction | 17.3 | 22.4 | 31.7 | |
| % nullo-4 | 5.6 | 9.4 | 4.0 | |
| % diplo-4 | 2.8 | 8.7 | 6.4 | |
| Total % 4 nondisjunction | 8.4 | 18.1 | 10.4 | |
| % nonhomologous disjunction | 0 | 0 | 0 | |
| Gamete types . | Maternal genotype . | |||
|---|---|---|---|---|
| Maternal . | Paternal . | mei-P261 . | mei-P261/mei-P26fs1 . | mei-P261/Df(1)18.1.15 . |
| Normal | ||||
| X; 4 | XY; 44 | 658 | 124 | 155 |
| X; 4 | 0; 44 | 884 | 139 | 173 |
| X nondisjunction | ||||
| 0; 4 | XY; 44 | 49 | 14 | 34 |
| XX; 4 | 0; 44 | 63 | 15 | 25 |
| 4 nondisjunction | ||||
| X; 0 | XY; 44 | 15 | 9 | 0 |
| X; 0 | 0; 44 | 14 | 14 | 2 |
| X; 44 | XY; 0 | 11 | 8 | 10 |
| X; 44 | 0; 0 | 11 | 10 | 0 |
| X; 4 nondisjunction | ||||
| 0; 0 | XY; 44 | 30 | 5 | 9 |
| XX; 44 | 0; 0 | 9 | 4 | 10 |
| 0; 44 | XY; 0 | 7 | 4 | 1 |
| XX; 0 | 0; 44 | 9 | 2 | 0 |
| Total progeny | 1760 | 348 | 419 | |
| Adjusted total | 1927 | 392 | 498 | |
| % nullo-X | 8.9 | 11.7 | 17.7 | |
| % diplo-X | 8.4 | 10.7 | 14.1 | |
| Total % X nondisjunction | 17.3 | 22.4 | 31.7 | |
| % nullo-4 | 5.6 | 9.4 | 4.0 | |
| % diplo-4 | 2.8 | 8.7 | 6.4 | |
| Total % 4 nondisjunction | 8.4 | 18.1 | 10.4 | |
| % nonhomologous disjunction | 0 | 0 | 0 | |
X and 4th chromosome segregation in mei-P26 mutants
| Gamete types . | Maternal genotype . | |||
|---|---|---|---|---|
| Maternal . | Paternal . | mei-P261 . | mei-P261/mei-P26fs1 . | mei-P261/Df(1)18.1.15 . |
| Normal | ||||
| X; 4 | XY; 44 | 658 | 124 | 155 |
| X; 4 | 0; 44 | 884 | 139 | 173 |
| X nondisjunction | ||||
| 0; 4 | XY; 44 | 49 | 14 | 34 |
| XX; 4 | 0; 44 | 63 | 15 | 25 |
| 4 nondisjunction | ||||
| X; 0 | XY; 44 | 15 | 9 | 0 |
| X; 0 | 0; 44 | 14 | 14 | 2 |
| X; 44 | XY; 0 | 11 | 8 | 10 |
| X; 44 | 0; 0 | 11 | 10 | 0 |
| X; 4 nondisjunction | ||||
| 0; 0 | XY; 44 | 30 | 5 | 9 |
| XX; 44 | 0; 0 | 9 | 4 | 10 |
| 0; 44 | XY; 0 | 7 | 4 | 1 |
| XX; 0 | 0; 44 | 9 | 2 | 0 |
| Total progeny | 1760 | 348 | 419 | |
| Adjusted total | 1927 | 392 | 498 | |
| % nullo-X | 8.9 | 11.7 | 17.7 | |
| % diplo-X | 8.4 | 10.7 | 14.1 | |
| Total % X nondisjunction | 17.3 | 22.4 | 31.7 | |
| % nullo-4 | 5.6 | 9.4 | 4.0 | |
| % diplo-4 | 2.8 | 8.7 | 6.4 | |
| Total % 4 nondisjunction | 8.4 | 18.1 | 10.4 | |
| % nonhomologous disjunction | 0 | 0 | 0 | |
| Gamete types . | Maternal genotype . | |||
|---|---|---|---|---|
| Maternal . | Paternal . | mei-P261 . | mei-P261/mei-P26fs1 . | mei-P261/Df(1)18.1.15 . |
| Normal | ||||
| X; 4 | XY; 44 | 658 | 124 | 155 |
| X; 4 | 0; 44 | 884 | 139 | 173 |
| X nondisjunction | ||||
| 0; 4 | XY; 44 | 49 | 14 | 34 |
| XX; 4 | 0; 44 | 63 | 15 | 25 |
| 4 nondisjunction | ||||
| X; 0 | XY; 44 | 15 | 9 | 0 |
| X; 0 | 0; 44 | 14 | 14 | 2 |
| X; 44 | XY; 0 | 11 | 8 | 10 |
| X; 44 | 0; 0 | 11 | 10 | 0 |
| X; 4 nondisjunction | ||||
| 0; 0 | XY; 44 | 30 | 5 | 9 |
| XX; 44 | 0; 0 | 9 | 4 | 10 |
| 0; 44 | XY; 0 | 7 | 4 | 1 |
| XX; 0 | 0; 44 | 9 | 2 | 0 |
| Total progeny | 1760 | 348 | 419 | |
| Adjusted total | 1927 | 392 | 498 | |
| % nullo-X | 8.9 | 11.7 | 17.7 | |
| % diplo-X | 8.4 | 10.7 | 14.1 | |
| Total % X nondisjunction | 17.3 | 22.4 | 31.7 | |
| % nullo-4 | 5.6 | 9.4 | 4.0 | |
| % diplo-4 | 2.8 | 8.7 | 6.4 | |
| Total % 4 nondisjunction | 8.4 | 18.1 | 10.4 | |
| % nonhomologous disjunction | 0 | 0 | 0 | |
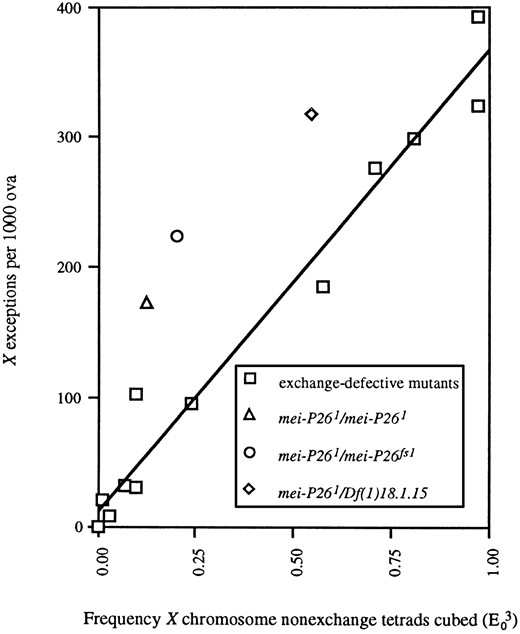
The cube of E0, when plotted against X chromosome exceptions per 1000 ova, reveals a linear relationship (Baker and Hall 1976). In recombination-defective mutants, X nondisjunction was found to be increased in oocytes that are also exceptional for one of the major autosomes (Baker and Carpenter 1972; Hall 1972; Parry 1973), which led to the hypothesis that the correlation with reflects a necessity for three chromosome arms (the X and the two arms of either chromosome 2 or 3) to be nonexchange in oocytes where the X chromosome fails to segregate properly (Baker and Hall 1976). The nondisjunction presumably results from failures of the achiasmate segregation system, resulting in XXA ↔ A segregations (Hawley and Theurkauf 1993). Baker and Hall (1976) demonstrated that the relationship between and X nondisjunction frequency is evident using recombination data measured on either chromosome X or 2 to estimate E0. We therefore used our recombination data for the left arm of chromosome 2 in mei-P26 mutant genotypes (Table 3; discussed below) to estimate E0, and found that the X nondisjunction frequency is also proportional to in mei-P26 mutants (Figure 3).
The second line of evidence that the X chromosome nondisjunction induced by mei-P26 mutations is due to the failed segregation of nonexchange tetrads comes from a genetic cross in which y w mei-P261 m f · y+/y pn cv mei-P261 females were crossed to YSX · YL, v f B/0; C(4)RM, ci eyR/0 males (data not shown). In this cross, recombination can be monitored in the ova leading to diplo-X exceptions, since these will be recovered as progeny resulting from fertilization with nullo-X sperm. If recombination occurs in tetrads leading to the diplo-X exceptions, one-fourth of the progeny would be expected to be homozygous for pn, owing to recombination between the centromere and this distal marker. However, 0/74 diplo-X exceptions recovered from this cross were homozygous for pn, indicating that most, if not all, of the nondisjunction in mei-P26 results from the missegregation of nonexchange tetrads.
The levels of chromosome 4 nondisjunction increase as the frequency of X nondisjunction increases (Table 2). In the genotypes tested, from 26.7 to 62.5% of 4th chromosome exceptions in mei-P26 mutant females occurred in oocytes that were simultaneously exceptional for the X chromosome. Moreover, the number of simultaneous X; 4 exceptions is about two to four times greater than would be expected if X and 4th chromosome segregation were independent. Missegregation of the always achiasmate 4th chromosome would be expected from an achiasmate segregation mutant, but in such mutants (e.g., Axs), the chromosome pairs predominantly segregate to opposite poles, a process known an excess as nonhomologous disjunction (e.g., XX ↔ 44). In no genotype mutant for mei-P26 did we observe of nonhomologous disjunction (XX ↔ 44; Table 2). The frequency of XX and 44 segregation to the same pole was equal to or greater than the frequency of XX and 44 segregation to opposite poles in mei-P26 mutants. Thus, when simultaneously nondisjunctional, the X and 4th chromosome segregate randomly to the poles.
Most recombination-defective meiotic mutants in Drosophila also show an excess of simultaneous X and 4th chromosome nondisjunction with no preference for nonhomologous disjunction. These observations led to the hypothesis that nonexchange X chromosomes disrupt 4th chromosome segregation through physical association of the chromosomes (Baker and Carpenter 1972; Hall 1972). Alternatively, association of the chromosome pairs in the presence of the narrow meiosis I spindle may disrupt the segregation of other chromosomes, resulting in nondisjunction at random with respect to the spindle poles (Hawley and Theurkauf 1993).
Mutants in the mei-P26 gene reduce meiotic exchange in a polar manner: Analyses of recombination along the X chromosome and the left arm of chromosome 2 revealed a polar recombination defect similar to many other previously identified recombination-defective meiotic mutants in Drosophila (Baker and Hall 1976; Sekelsky et al. 1999). As shown in Table 3, testing of chromosome 2 recombination in mei-P261 homozygotes reveals a reduction in recombination frequency and an abnormal distribution of exchanges. Recombination is strongly decreased in distal 2L, but shows frequencies greater than wild type in the proximal region of the chromosome, near the centromeric heterochromatin. The recombination analysis of mei-P26fs1/mei-P261 females agrees with the nondisjunction data in that the decrease in recombination is more severe in these females than in mei-P261 homozygotes (Table 3). As expected, the largest decrease in recombination is observed in mei-P261/Df(1)18.1.15 females, in which the genetic map distance is reduced to one-third of that observed in mei-P261 homozygotes and one-fifth of wild type (Table 3). The recombination distribution in mei-P26fs1/mei-P261 and mei-P261/Df(1)18.1.15 females is polar, as in mei-P261 homozygotes, but is more exaggerated.
Comparison of overall map distances shows that the frequency of recovered crossovers is reduced by 30 to 80%, depending on the mei-P26 mutant genotype. However, analysis of recombination frequencies allows us to assess only one of the four products of a meiotic tetrad. Using the methods of Weinstein (1936), exchange rank frequencies, the frequencies of meiotic tetrads with zero, one, two, or more crossovers, can be calculated from recombination data. This demonstrates that mei-P26 increases E0, the frequency of tetrads in which no exchange occurred (Table 3). E0 increases to ~0.5 in mei-P261 homozygotes and to ~0.59 in mei-P26fs1/mei-P261, compared to ~0.2 in wild type. Similarly, E0 in mei-P261/Df (1)18.1.15 females is increased to ~0.8, indicating an elevated frequency of tetrads in which no exchange has occurred, while E2 and E3 tetrads are not
Results of crosses of females of the genotype X/X; net ho dp Sp b pr cn/+ + + + + + + carrying the indicated X chromosomes by +/Y; net ho dp b pr cn / net ho dp b pr cn malesa
| . | Maternal genotype . | |||||||
|---|---|---|---|---|---|---|---|---|
| Progeny typeb . | +/+c . | y w/+ c . | mei-P261 . | mei-P261/+ d . | mei-P261/mei-P26fs1 . | mei-P26fs1/+ . | mei-P261/Dfe . | mei-P261; bamΔ86/+f . |
| Noncrossover | 995 | 1512 | 773 | 819 | 363 | 2188 | 169 | 96 |
| Single crossover | ||||||||
| 1 | 228 | 299 | 48 | 158 | 13 | 450 | 1 | 1 |
| 2 | 369 | 598 | 122 | 386 | 36 | 982 | 9 | 4 |
| 3 | 56 | 54 | 70 | 77 | 21 | 140 | 2 | 1 |
| 4 | 7 | 15 | 24 | 15 | 26 | 26 | 5 | 1 |
| Double crossover | ||||||||
| 1,2 | 20 | 7 | 2 | 9 | 1 | 27 | — | — |
| 1,3 | 3 | 1 | 4 | 4 | — | 22 | — | — |
| 1,4 | 1 | 3 | 2 | 1 | — | 4 | — | — |
| 2,3 | 7 | 6 | 6 | 8 | 2 | 25 | — | — |
| 2,4 | 2 | 3 | 5 | 7 | 1 | 4 | — | — |
| 3,4 | − | — | 2 | 1 | 1 | — | — | — |
| Triple crossover | − | — | 1 | 1 | — | 2 | — | — |
| Quadruple crossover | — | — | — | — | — | — | — | — |
| Total progeny | 1712 | 2500 | 1061 | 1486 | 464 | 3891 | 186 | 103 |
| Map distances | ||||||||
| 1 (net-dp) | 14.7 | 12.4 | 5.4 | 11.6 | 3.0 | 13.0 | 0.5 | 1.0 |
| 2 (dp-b) | 24.2 | 24.6 | 12.8 | 27.7 | 8.6 | 27.2 | 4.8 | 3.9 |
| 3 (b-pr) | 3.9 | 2.4 | 7.9 | 6.1 | 5.2 | 4.9 | 1.1 | 1.0 |
| 4 (pr-cn) | 0.6 | 0.8 | 3.1 | 1.6 | 6.0 | 0.9 | 2.7 | 1.0 |
| Total | 43.4 | 40.3 | 29.2 | 47.0 | 22.8 | 46.0 | 9.1 | 6.8 |
| Map relative to | ||||||||
| control | ||||||||
| 1 (net-dp) | 1 | 0.844 | 0.367 | 0.789 | 0.204 | 0.884 | 0.034 | 0.068 |
| 2 (dp-b) | 1 | 1.017 | 0.529 | 1.145 | 0.355 | 1.124 | 0.198 | 0.161 |
| 3 (b-pr) | 1 | 0.615 | 2.026 | 1.564 | 1.333 | 1.256 | 0.282 | 0.462 |
| 4 (pr-cn) | 1 | 1.333 | 5.167 | 2.667 | 10.000 | 1.500 | 4.500 | 3.000 |
| Total | 1 | 0.929 | 0.673 | 1.083 | 0.525 | 1.060 | 0.210 | 0.196 |
| Exchange rank | ||||||||
| frequencies | ||||||||
| E0 | 0.209 | 0.226 | 0.499 | 0.143 | 0.586 | 0.167 | 0.817 | 0.864 |
| E1 | 0.714 | 0.742 | 0.426 | 0.779 | 0.371 | 0.751 | 0.183 | 0.136 |
| E2 | 0.077 | 0.032 | 0.068 | 0.073 | 0.043 | 0.078 | 0 | 0 |
| E3 | 0 | 0 | 0.008 | 0.005 | 0 | 0.004 | 0 | 0 |
| E4 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| . | Maternal genotype . | |||||||
|---|---|---|---|---|---|---|---|---|
| Progeny typeb . | +/+c . | y w/+ c . | mei-P261 . | mei-P261/+ d . | mei-P261/mei-P26fs1 . | mei-P26fs1/+ . | mei-P261/Dfe . | mei-P261; bamΔ86/+f . |
| Noncrossover | 995 | 1512 | 773 | 819 | 363 | 2188 | 169 | 96 |
| Single crossover | ||||||||
| 1 | 228 | 299 | 48 | 158 | 13 | 450 | 1 | 1 |
| 2 | 369 | 598 | 122 | 386 | 36 | 982 | 9 | 4 |
| 3 | 56 | 54 | 70 | 77 | 21 | 140 | 2 | 1 |
| 4 | 7 | 15 | 24 | 15 | 26 | 26 | 5 | 1 |
| Double crossover | ||||||||
| 1,2 | 20 | 7 | 2 | 9 | 1 | 27 | — | — |
| 1,3 | 3 | 1 | 4 | 4 | — | 22 | — | — |
| 1,4 | 1 | 3 | 2 | 1 | — | 4 | — | — |
| 2,3 | 7 | 6 | 6 | 8 | 2 | 25 | — | — |
| 2,4 | 2 | 3 | 5 | 7 | 1 | 4 | — | — |
| 3,4 | − | — | 2 | 1 | 1 | — | — | — |
| Triple crossover | − | — | 1 | 1 | — | 2 | — | — |
| Quadruple crossover | — | — | — | — | — | — | — | — |
| Total progeny | 1712 | 2500 | 1061 | 1486 | 464 | 3891 | 186 | 103 |
| Map distances | ||||||||
| 1 (net-dp) | 14.7 | 12.4 | 5.4 | 11.6 | 3.0 | 13.0 | 0.5 | 1.0 |
| 2 (dp-b) | 24.2 | 24.6 | 12.8 | 27.7 | 8.6 | 27.2 | 4.8 | 3.9 |
| 3 (b-pr) | 3.9 | 2.4 | 7.9 | 6.1 | 5.2 | 4.9 | 1.1 | 1.0 |
| 4 (pr-cn) | 0.6 | 0.8 | 3.1 | 1.6 | 6.0 | 0.9 | 2.7 | 1.0 |
| Total | 43.4 | 40.3 | 29.2 | 47.0 | 22.8 | 46.0 | 9.1 | 6.8 |
| Map relative to | ||||||||
| control | ||||||||
| 1 (net-dp) | 1 | 0.844 | 0.367 | 0.789 | 0.204 | 0.884 | 0.034 | 0.068 |
| 2 (dp-b) | 1 | 1.017 | 0.529 | 1.145 | 0.355 | 1.124 | 0.198 | 0.161 |
| 3 (b-pr) | 1 | 0.615 | 2.026 | 1.564 | 1.333 | 1.256 | 0.282 | 0.462 |
| 4 (pr-cn) | 1 | 1.333 | 5.167 | 2.667 | 10.000 | 1.500 | 4.500 | 3.000 |
| Total | 1 | 0.929 | 0.673 | 1.083 | 0.525 | 1.060 | 0.210 | 0.196 |
| Exchange rank | ||||||||
| frequencies | ||||||||
| E0 | 0.209 | 0.226 | 0.499 | 0.143 | 0.586 | 0.167 | 0.817 | 0.864 |
| E1 | 0.714 | 0.742 | 0.426 | 0.779 | 0.371 | 0.751 | 0.183 | 0.136 |
| E2 | 0.077 | 0.032 | 0.068 | 0.073 | 0.043 | 0.078 | 0 | 0 |
| E3 | 0 | 0 | 0.008 | 0.005 | 0 | 0.004 | 0 | 0 |
| E4 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
Female progeny were scored in the indicated crosses (see materials and methods) except for the +/+ control and y w mei-P261; bamΔ86 ry e/ + + + crosses, in which all progeny were scored. In the y w mei-P261; bamΔ86ry e/ + + + cross, eye color phenotype was not scored in male progeny, with the assumption that few b-pr or pr-cn recombinants were present. For this reason, the map distances for these intervals may have been underestimated in this cross.
Region 1 is net-dp; region 2 is dp-b; region 3 is b-pr; region 4 is pr-cn.
It should be noted that the control recombination levels differ from the standard Drosophila genetic map (Lindsley and Zimm 1992), although they are similar to previously published control values (Baker and Carpenter 1972).
A possible semidominant effect on recombination was suggested by Sekelsky et al. (1999) using a comparison to standard map distances. However, comparison of recombination in mei-P261/+ females vs. wild-type controls indicates no such effect.
Df(1)18.1.15 was used in this experiment.
bamΔ86 is carried on chromosome 3.
Results of crosses of females of the genotype X/X; net ho dp Sp b pr cn/+ + + + + + + carrying the indicated X chromosomes by +/Y; net ho dp b pr cn / net ho dp b pr cn malesa
| . | Maternal genotype . | |||||||
|---|---|---|---|---|---|---|---|---|
| Progeny typeb . | +/+c . | y w/+ c . | mei-P261 . | mei-P261/+ d . | mei-P261/mei-P26fs1 . | mei-P26fs1/+ . | mei-P261/Dfe . | mei-P261; bamΔ86/+f . |
| Noncrossover | 995 | 1512 | 773 | 819 | 363 | 2188 | 169 | 96 |
| Single crossover | ||||||||
| 1 | 228 | 299 | 48 | 158 | 13 | 450 | 1 | 1 |
| 2 | 369 | 598 | 122 | 386 | 36 | 982 | 9 | 4 |
| 3 | 56 | 54 | 70 | 77 | 21 | 140 | 2 | 1 |
| 4 | 7 | 15 | 24 | 15 | 26 | 26 | 5 | 1 |
| Double crossover | ||||||||
| 1,2 | 20 | 7 | 2 | 9 | 1 | 27 | — | — |
| 1,3 | 3 | 1 | 4 | 4 | — | 22 | — | — |
| 1,4 | 1 | 3 | 2 | 1 | — | 4 | — | — |
| 2,3 | 7 | 6 | 6 | 8 | 2 | 25 | — | — |
| 2,4 | 2 | 3 | 5 | 7 | 1 | 4 | — | — |
| 3,4 | − | — | 2 | 1 | 1 | — | — | — |
| Triple crossover | − | — | 1 | 1 | — | 2 | — | — |
| Quadruple crossover | — | — | — | — | — | — | — | — |
| Total progeny | 1712 | 2500 | 1061 | 1486 | 464 | 3891 | 186 | 103 |
| Map distances | ||||||||
| 1 (net-dp) | 14.7 | 12.4 | 5.4 | 11.6 | 3.0 | 13.0 | 0.5 | 1.0 |
| 2 (dp-b) | 24.2 | 24.6 | 12.8 | 27.7 | 8.6 | 27.2 | 4.8 | 3.9 |
| 3 (b-pr) | 3.9 | 2.4 | 7.9 | 6.1 | 5.2 | 4.9 | 1.1 | 1.0 |
| 4 (pr-cn) | 0.6 | 0.8 | 3.1 | 1.6 | 6.0 | 0.9 | 2.7 | 1.0 |
| Total | 43.4 | 40.3 | 29.2 | 47.0 | 22.8 | 46.0 | 9.1 | 6.8 |
| Map relative to | ||||||||
| control | ||||||||
| 1 (net-dp) | 1 | 0.844 | 0.367 | 0.789 | 0.204 | 0.884 | 0.034 | 0.068 |
| 2 (dp-b) | 1 | 1.017 | 0.529 | 1.145 | 0.355 | 1.124 | 0.198 | 0.161 |
| 3 (b-pr) | 1 | 0.615 | 2.026 | 1.564 | 1.333 | 1.256 | 0.282 | 0.462 |
| 4 (pr-cn) | 1 | 1.333 | 5.167 | 2.667 | 10.000 | 1.500 | 4.500 | 3.000 |
| Total | 1 | 0.929 | 0.673 | 1.083 | 0.525 | 1.060 | 0.210 | 0.196 |
| Exchange rank | ||||||||
| frequencies | ||||||||
| E0 | 0.209 | 0.226 | 0.499 | 0.143 | 0.586 | 0.167 | 0.817 | 0.864 |
| E1 | 0.714 | 0.742 | 0.426 | 0.779 | 0.371 | 0.751 | 0.183 | 0.136 |
| E2 | 0.077 | 0.032 | 0.068 | 0.073 | 0.043 | 0.078 | 0 | 0 |
| E3 | 0 | 0 | 0.008 | 0.005 | 0 | 0.004 | 0 | 0 |
| E4 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| . | Maternal genotype . | |||||||
|---|---|---|---|---|---|---|---|---|
| Progeny typeb . | +/+c . | y w/+ c . | mei-P261 . | mei-P261/+ d . | mei-P261/mei-P26fs1 . | mei-P26fs1/+ . | mei-P261/Dfe . | mei-P261; bamΔ86/+f . |
| Noncrossover | 995 | 1512 | 773 | 819 | 363 | 2188 | 169 | 96 |
| Single crossover | ||||||||
| 1 | 228 | 299 | 48 | 158 | 13 | 450 | 1 | 1 |
| 2 | 369 | 598 | 122 | 386 | 36 | 982 | 9 | 4 |
| 3 | 56 | 54 | 70 | 77 | 21 | 140 | 2 | 1 |
| 4 | 7 | 15 | 24 | 15 | 26 | 26 | 5 | 1 |
| Double crossover | ||||||||
| 1,2 | 20 | 7 | 2 | 9 | 1 | 27 | — | — |
| 1,3 | 3 | 1 | 4 | 4 | — | 22 | — | — |
| 1,4 | 1 | 3 | 2 | 1 | — | 4 | — | — |
| 2,3 | 7 | 6 | 6 | 8 | 2 | 25 | — | — |
| 2,4 | 2 | 3 | 5 | 7 | 1 | 4 | — | — |
| 3,4 | − | — | 2 | 1 | 1 | — | — | — |
| Triple crossover | − | — | 1 | 1 | — | 2 | — | — |
| Quadruple crossover | — | — | — | — | — | — | — | — |
| Total progeny | 1712 | 2500 | 1061 | 1486 | 464 | 3891 | 186 | 103 |
| Map distances | ||||||||
| 1 (net-dp) | 14.7 | 12.4 | 5.4 | 11.6 | 3.0 | 13.0 | 0.5 | 1.0 |
| 2 (dp-b) | 24.2 | 24.6 | 12.8 | 27.7 | 8.6 | 27.2 | 4.8 | 3.9 |
| 3 (b-pr) | 3.9 | 2.4 | 7.9 | 6.1 | 5.2 | 4.9 | 1.1 | 1.0 |
| 4 (pr-cn) | 0.6 | 0.8 | 3.1 | 1.6 | 6.0 | 0.9 | 2.7 | 1.0 |
| Total | 43.4 | 40.3 | 29.2 | 47.0 | 22.8 | 46.0 | 9.1 | 6.8 |
| Map relative to | ||||||||
| control | ||||||||
| 1 (net-dp) | 1 | 0.844 | 0.367 | 0.789 | 0.204 | 0.884 | 0.034 | 0.068 |
| 2 (dp-b) | 1 | 1.017 | 0.529 | 1.145 | 0.355 | 1.124 | 0.198 | 0.161 |
| 3 (b-pr) | 1 | 0.615 | 2.026 | 1.564 | 1.333 | 1.256 | 0.282 | 0.462 |
| 4 (pr-cn) | 1 | 1.333 | 5.167 | 2.667 | 10.000 | 1.500 | 4.500 | 3.000 |
| Total | 1 | 0.929 | 0.673 | 1.083 | 0.525 | 1.060 | 0.210 | 0.196 |
| Exchange rank | ||||||||
| frequencies | ||||||||
| E0 | 0.209 | 0.226 | 0.499 | 0.143 | 0.586 | 0.167 | 0.817 | 0.864 |
| E1 | 0.714 | 0.742 | 0.426 | 0.779 | 0.371 | 0.751 | 0.183 | 0.136 |
| E2 | 0.077 | 0.032 | 0.068 | 0.073 | 0.043 | 0.078 | 0 | 0 |
| E3 | 0 | 0 | 0.008 | 0.005 | 0 | 0.004 | 0 | 0 |
| E4 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
Female progeny were scored in the indicated crosses (see materials and methods) except for the +/+ control and y w mei-P261; bamΔ86 ry e/ + + + crosses, in which all progeny were scored. In the y w mei-P261; bamΔ86ry e/ + + + cross, eye color phenotype was not scored in male progeny, with the assumption that few b-pr or pr-cn recombinants were present. For this reason, the map distances for these intervals may have been underestimated in this cross.
Region 1 is net-dp; region 2 is dp-b; region 3 is b-pr; region 4 is pr-cn.
It should be noted that the control recombination levels differ from the standard Drosophila genetic map (Lindsley and Zimm 1992), although they are similar to previously published control values (Baker and Carpenter 1972).
A possible semidominant effect on recombination was suggested by Sekelsky et al. (1999) using a comparison to standard map distances. However, comparison of recombination in mei-P261/+ females vs. wild-type controls indicates no such effect.
Df(1)18.1.15 was used in this experiment.
bamΔ86 is carried on chromosome 3.
observed at all. These data are consistent with mei-P261 and mei-P26fs1 being hypomorphic alleles of differing severity.
Exchange in females carrying mutations in mei-P26 is decreased overall and altered in a polar fashion, where distal and medial exchanges are more strongly suppressed than those near the centromere. In more severe alleles of mei-P26, recombination is more severely affected, and the calculated frequency of nonexchange tetrads is increased. The alleles of mei-P26 form an allelic series as follows: mei-P261 < mei-P26fs1 (and other fs alleles) < mei-P26mfs1, in which the effects on male and
A linear correlation exists between X nondisjunction frequency and in exchange-defective mutants and mei-P26. This graph is based on Figure 4 of Baker and Hall (1976). Data for the exchange-defective mutants are from Baker and Hall (1976) and references therein. Data for mei-P26 are from Tables 2 and 3 (this study). While there is some deviation in the frequency of X nondisjunction in mei-P26 mutants from that expected based on the other exchange-defective mutants, we believe that this is attributable to differences in genetic background, since the slope of the linear relationship between and X nondisjunction differs very little between mei-P26 (slope = 320.2) and the exchange-defective mutants (slope = 348.6).
female germline differentiation (see below) and meiotic recombination increase in severity.
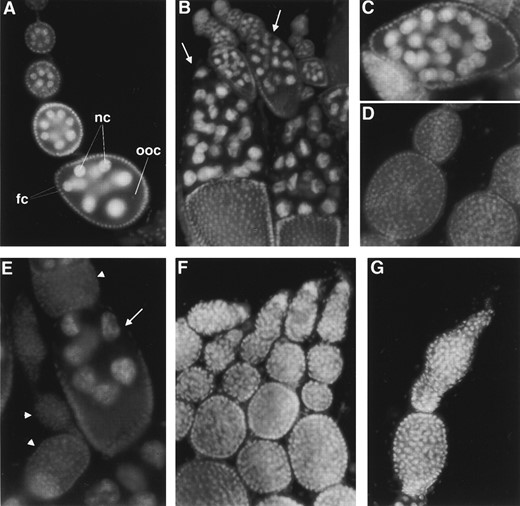
Cytological studies of female sterility: To investigate the cause of the sterility in the female sterile alleles, ovaries from homozygous mei-P26fs1 females were analyzed using fluorescence microscopy after DAPI staining. In wild-type ovaries, the ovarioles contain a series of developing egg chambers, each comprised of 15 nurse cells and an oocyte (Figure 4A). In contrast, ovarioles from mei-P26fs1 ovaries often contained chamberscysts filled with large numbers of small, apparently undifferentiated cells (Figure 4D). This phenotype resembled the “tumorous ovary” phenotype observed in a group of female sterile mutants including bam, bgcn, otu, etc. (Gateff 1981; Schüpbach 1985; King et al. 1986; Perrimon et al. 1986; Salz et al. 1987; McKearin and Spradling 1990; Oliver et al. 1990). A total of 84.4% of the chambers (n = 495) in mei-P26fs1 females were tumorous. The remaining chambers more closely resembled wild type, in that nurse cells were present. Although some of these appeared to be normal, many contained abnormal numbers of nurse cells (Figure 4). In agreement with these findings, mei-P26fs1 females do lay a reduced number of eggs (0.07 eggs per female per hour, compared to 1.87 eggs per female per hour for wild type).
For comparison, ovaries from mei-P261 homozygotes were also analyzed. While tumorous chambers were very rare (0.7%, n = 305), the other abnormalities in oogenesis observed for mei-P26fs1 were also present in mei-P261 ovaries (Figure 4B). The most prominent of these were the defects in nurse cell number, with chambers often observed to have either too many or too few nurse cells. In mei-P261/Df females, a more severe phenotype was observed, with a greater frequency of tumorous egg chambers (data not shown).
Ovaries from homozygotes for mei-P26mfs1, the male and female sterile allele of mei-P26, were completely tumorous (Figure 4E). No normal-looking egg chambers were seen in ovaries homozygous for this allele. Instead, 100% of the egg chambers present (n = 384) resembled the tumorous chambers observed at a lesser frequency in mei-P26fs1 homozygotes, and mei-P26mfs1 homozygotes do not lay eggs. The ovarian defects in mei-P26 mutants appear to affect the differentiation and proliferation of female germline cells, which leads to sterility. The severity of these defects correlates with the severity of the meiotic phenotype and infertility in mei-P261 homozygotes and mei-P26fs1/mei-P261 heterozygotes.
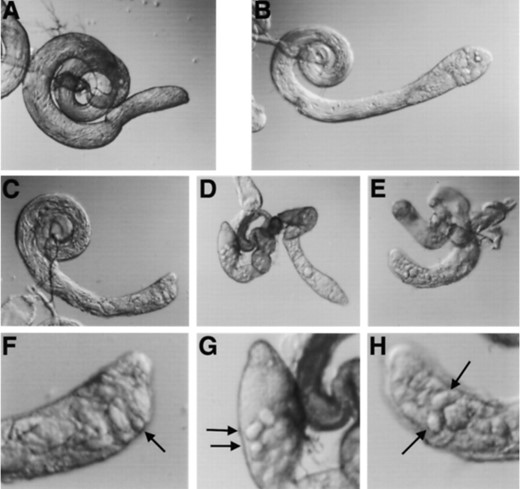
Effects of mei-P26mfs1 on male fertility: Testes from mei-P26mfs1/y+Y males were examined to determine the cause of the male sterility. In contrast to testes from mei-P261/y+Y or mei-P26fs1/y+Y males, no mature motile spermatozoa were present, although somewhat disorganized immature elongated spermatid bundles are produced (Figure 5C). In addition, the testes contained highly refractile cysts that most likely represent degenerating cysts of spermatogonial cells (Figure 5F). Although mei-P261/y+Y males produce motile spermatozoa and are fertile, these refractile cysts are also often observed in testes of this genotype (Figure 5B). Cysts of this morphology are often seen in testes mutant for bam, another gene that affects male and female gametogenesis (Figure 5, D and G). mei-P26mfs1 represents a more severe class of allele than mei-P26fs1, since it affects both male and female gametogenesis more drastically than any other mei-P26 allele.
mei-P26 shows multiple genetic interactions with a null mutation at the bam locus: The product of the bam gene is important in both male and female germlines for both the specification of cystoblast fate and the cessation of the germline mitotic divisions before entry into the meiotic cell cycle (McKearin and Spradling 1990;
Ovarian phenotypes in mei-P26 mutants. Nuclei were visualized by DAPI staining. (A) Wild-type ovarioles are composed of a series of egg chambers that each contain 15 nurse cells (nc) and an oocyte (ooc), surrounded by a layer of follicle cells (fc). (B) Homozygous mei-P261 ovarioles also show defects, most notably in nurse cell number. Arrows indicate egg chambers with >15 nurse cells. (C) Abnormal egg chamber from a mei-P26fs1 homozygote, with 31 nurse cells. (D) Tumorous egg chambers in ovarioles from mei-P26fs1 homozygote females contain large numbers of undifferentiated cells. (E) mei-P261; bamΔ86/+ ovarioles show a higher frequency of tumorous chambers (arrowheads) than do mei-P261 homozygotes and egg chambers with abnormal numbers of nurse cells (arrow indicates a chamber with 7 nurse cells). (F) Ovarioles from mei-P26mfs1 homozygotes show only tumorous chambers. (G) Ovariole from a bamΔ86 homozygote, showing tumorous phenotype.
McKearin and Ohlstein 1995; Gönczy et al. 1997; Ohlstein and McKearin 1997). Since mutations in the mei-P26 gene produce an ovary phenotype with similarities to that of bam and similarities exist in the effect of mei-P26 in testes, we tested double mutant combinations of mei-P261 and bamΔ86, a null allele (McKearin and Ohlstein 1995), to determine whether these mutants could interact genetically.
Although heterozygotes for the bamΔ86 allele are fertile, males of the genotype mei-P261/Y; bamΔ86/+ were found to be completely sterile. Analysis of testes from males of this genotype showed a phenotype almost identical to bamΔ86 homozygote males (Figure 5, E and H). Similarly, mei-P261; bamΔ86/+ females also showed a severe decrease in fertility compared to mei-P261 homozygotes, due to an increase in the formation of ovarian tumors and other egg chamber defects (Figure 4D). In mei-P261; bamΔ86/+ females, 28.9% of chambers were tumorous, compared to 0.7% in mei-P261; +/+ females. A normal level of fertility was observed for mei-P261/+; bamΔ86/+ females. Finally, recombination, as measured in the few progeny produced by mei-P261; bamΔ86/+ females, was also severely decreased (Table 3). These data show that mei-P26 interacts with bam in the regulation of germline differentiation and that defects in proper differentiation of the germline cyst may disturb the proper distribution of meiotic recombination events.
DISCUSSION
Our genetic analysis of mei-P26 shows it to be a novel exchange-defective female meiotic mutant in Drosophila, which is also essential for proper germline differentiation in both males and females. Furthermore, the effects of mei-P26 on meiotic recombination and germline differentiation in both sexes are exacerbated by heterozygosity for bam, a gene already known to function in the germline to control the differentiation and proliferation of germline cells (McKearin and Spradling 1990; McKearin and Ohlstein 1995; Gönczy et al. 1997; Ohlstein and McKearin 1997). These results indicate that some of the processes that underlie proper germline cyst development may also be required for the normal levels and distribution of meiotic crossovers.
The role of mei-P26 in female germline development: mei-P26-induced defects in female germline differentiation appear to occur during the early mitotic divisions in cyst formation, and hypomorphic alleles of mei-P26,
Testis morphology in mei-P26 mutants as visualized by DIC microscopy. (A) Wild-type testis. (B) mei-P261/y+Y testis contains highly refractile degenerating cysts (arrow). (C) mei-P26mfs1/y+Y testes support the production of elongated spermatid bundles, but not motile spermatids. (D) No elongated spermatids are present in testes from bamΔ86 homozygotes. (E) Spermatid differentiation also fails to progress in mei-P261/y+Y; bamΔ86/+ testes (note morphological similarity to D). (F) Magnified view of anterior tip of the mei-P26mfs1/y+Y testes from C (above). Arrow, degenerating cyst. (G) Magnified view of anterior tip of the bamΔ86 homozygote testes from D (above). Arrows, degenerating cysts. (H) Magnified view of anterior tip of the mei-P261/y+Y; bamΔ86/+ testes from E (above). Arrows, degenerating cysts.
such as mei-P261, produce egg chambers with abnormal numbers of nurse cells. Cysts with an increased number of nurse cells may be due to an additional round of mitosis occurring in some or all of the cystocytes (Hawkins et al. 1996). Alternatively, a defect in follicle cell packaging of normal cysts may result in chambers containing an excess of cells (Ruhola et al. 1991; Goode et al. 1992; Siegel et al. 1993). However, we did not observe egg chambers containing two oocytes, which would be expected if two 16-cell cysts were packaged together. Similarly, cysts with too few nurse cells could also result from packaging of cysts that have abnormally broken their intercellular connections to form smaller clusters of cells (Yue and Spradling 1992). Cysts containing 7 nurse cells and an oocyte may also result from the cystocytes undergoing only three divisions (Lilly and Spradling 1996; de Cuevas et al. 1997).
Allelic combinations of mei-P26 that alter the number of nurse cells also reduce the number of normal eggs produced, but they do not cause complete sterility. In addition to the defects in nurse cell number, a tumorous ovary phenotype is more frequent in severe alleles. In females homo- or hemizygous for these mutations, the egg chambers become filled with hundreds of small cells, and no oocyte develops. This phenotype is similar to that exhibited by mutants in the bam and benign gonial cell neoplasm (bgcn) genes (Gateff 1981; McKearin and Spradling 1990).
The most severe mei-P26 allele affects germline differentiation in both males and females. In mei-P26mfs1, ovaries consist entirely of tumorous egg chambers and, in males carrying this allele, spermatid differentiation progresses only to the point of producing elongated spermatid bundles, and mature spermatozoa are not produced. The mei-P26mfs1 allele thus bears some similarities to mutants in the bam and bgcn genes, both of which cause a tumorous phenotype in ovaries and arrest of spermatogenesis (Gateff 1981; McKearin and Spradling 1990). However, spermatid differentiation is arrested at an earlier stage in bam and bgcn mutants. Ovary morphology differs somewhat, in that mei-P26 mutant ovarioles consist of a series of defined chambers, while bam and bgcn ovarioles often appear as distended germaria with few distinct chambers. Nevertheless, the interaction of mei-P26 with bam suggests that the similarities in the phenotypes in these mutants are not coincidental (see below).
Certain mutations in the Sex-lethal (Sxl), ovarian tumor (otu), and ovo genes also cause sterility in females due to the formation of tumorous egg chambers (Schüpbach 1985; King et al. 1986; Perrimon et al. 1986; Salz et al. 1987; Oliver et al. 1990). Interestingly, mild to severe defects in meiotic recombination frequencies are observed in females bearing heteroallelic combinations of female sterile alleles for these loci (Cook 1993) and in females in which the effects of Sxl mutations have been partially suppressed using genetic modifiers (Bopp et al. 1999). These results further suggest that the processes that control exchange position and determine germline cyst formation may be coordinately controlled.
mei-P26 encodes a RING finger protein: The structure of the MEI-P26 protein suggests a few possibilities for the role of mei-P26. mei-P26 is predicted to encode a member of the RBCC family of proteins, which contain, in their N-terminal regions, a RING finger motif followed by one or two copies of a second cysteine-rich motif called the B-box and a coiled coil region (Reddy et al. 1992; Saurin et al. 1996). The RING finger and B-box motifs are believed to mediate physical interactions with other proteins (Borden et al. 1995, 1996; Cao et al. 1997, 1998). RBCC proteins are only a subset of the large number of known RING finger proteins, which have diverse roles in oncogenesis, transcriptional regulation, signal transduction, and development (reviewed by Saurin et al. 1996).
Several RBCC proteins, such as PML and the TIF1 family, are known to regulate transcription by binding to nuclear hormone receptors as coactivators or corepressors (Le Douarin et al. 1995; vom Baur et al. 1996; Zhong et al. 1999). While assembling factors for transcriptional regulation is one role for RBCC proteins, certain other RBCC proteins appear to function in capacities such as signal transduction (Rpt1, Patarca et al. 1988; Staf50, Tissot and Mechti 1995), or by forming ribonucleoprotein complexes (SS-A/Ro, Keech et al. 1995). Although a growing number of RBCC proteins have been identified, only a handful also contain NHL repeats, named after the proteins NCL-1, HT2A, and LIN-41 (Slack and Ruvkun 1998). The NHL repeat has been shown to be involved in protein-protein interactions in the RBCC proteins HT2A and BERP (Fridell et al. 1995; El-Husseini and Vincent 1999).
The RBCC-NHL proteins are mostly of unknown function (the predicted proteins KIAA0517, Nagase et al. 1998; F54G8.4 and F26F4.7, Wilson et al. 1994), or have not been extensively characterized at a molecular level (NCL-1, Frank and Roth 1998; HT2A, Fridell et al. 1995; LIN-41, Slack and Ruvkun 1998; BERP, El-Husseini and Vincent 1999; and MEI-P26, this study). RBCC-NHL proteins, which contain both RBCC and NHL domains, include at least two potential protein-protein interaction motifs, so a strong possibility is that these also may participate in the formation of multiprotein complexes. One possible candidate for a MEI-P26 partner protein is Bam.
The interaction of mei-P26 and bam: We have demonstrated a genetic interaction between mei-P26 and bam. Heterozygosity for a null mutant in bam enhances the phenotype of mei-P26, causing sterility in males, an increase in tumor formation in females, and a decrease in meiotic exchange. On the basis of the characterization of the bam and mei-P26 gene products, we can speculate on the nature of the interaction between these two genes.
First, MEI-P26 may act as a transcriptional or translational regulator that controls bam expression. A variety of ovarian defects like those observed in mei-P26, including tumorous chambers and cysts with abnormal numbers of nurse cells, are also seen in mutants for the Drosophila Rbp9 gene (Kim-Ha et al. 1999). The Rbp9 gene was shown to encode an RNA binding protein that binds specifically to the bam transcript and may act to regulate Bam expression in the germarium. Similarly, misregulation of bam expression in the encore (enc) mutant may underlie the effect of enc on nurse cell number (Hawkins et al. 1996). However, since heterozygosity for bam exacerbates the meiotic phenotype of mei-P26, this model predicts that the meiotic defects are due to the misregulation of bam, rather than through other genes possibly regulated by mei-P26.
Second, mei-P26 could be required for the proper localization or function of Bam. The product of the bam gene is expressed in the cytoplasm of cystoblasts and early germline cysts in females, where it is required for cystoblast differentiation. Bam protein also associates with the fusome, a large organelle mostly comprised of cytoskeletal and vacuolar components, which is present in early germline cysts (McKearin and Ohlstein 1995). According to this model, MEI-P26 may physically interact with Bam, possibly through the RBCC, NHL, or other motifs in the MEI-P26 protein. Alternatively, this regulation may be indirect, requiring other proteins. For example, MEI-P26 may regulate the bgcn gene product, which is required for Bam function (Lavoie et al. 1999). Again, this suggests that the effects on meiotic exchange are mediated by bam.
The evidence presented here does not allow us to determine the relative positions of bam and mei-P26 in a pathway. Therefore, bam may be required for mei-P26 function, which in turn would be required for proper germline cyst development and meiotic recombination. In this third model, MEI-P26 may physically interact with Bam and/or other proteins in the cytoplasm, possibly as a component of the fusome, from which MEI-P26 may facilitate normal germline development and meiotic exchange. Alternatively, MEI-P26 might be indirectly controlled by Bam as a downstream effector. While the relationship between mei-P26 and bam has not been fully elucidated, these models are intriguing, as they all suggest a role for bam in a pathway ensuring proper meiotic exchange.
A speculative model for precondition mutants: mei-P26 appears to behave as expected for a female meiotic precondition mutant. This group of Drosophila mutants presents a phenotype in which the total frequency of meiotic exchange is often reduced, although to differing levels, and residual exchanges are abnormally distributed in a polar fashion, with reduced frequencies in the distal parts of the chromosome arms (for a review, see Lindsley and Sandler 1977). In mei-P26, exchange is decreased overall and the distribution is polar. In more severe alleles, recombination is more severely affected, and E0 is increased. To provide a first step toward explaining the various components of the mei-P26 phenotype and the connection with germline differentiation, we propose the following rather speculative model for precondition mutants in general.
In many organisms, telomeres have been proposed as sites responsible for initiating at least part of the pairing interactions between homologous chromosomes. Evidence for the clustering of telomeres during meiotic prophase has been gathered through cytological studies in many species (reviewed by Dernburg et al. 1995). In these studies, chromosomes in meiotic prophase were observed to form what has been described as a “bouquet” configuration, where the telomeres are positioned together in a small portion of the nuclear volume. The telomeres are often clustered near a region of the nuclear envelope adjacent to the position of the cytoplasmic centrosome. It is thought that the clustering of telomeres may facilitate homolog pairing in meiosis.
The existence of a bouquet configuration has been demonstrated recently in Saccharomyces cerevisiae (Trelles-Sticken et al. 1999), and telomeres are responsible for a delay in meiotic progression observed in haploid yeast strains that are disomic for a single chromosome pair (Rockmill and Roeder 1998). Furthermore, the delay associated with the presence of telomeres required an intact NDJ1 gene (Chua and Roeder 1997; Conrad et al. 1997; Rockmill and Roeder 1998). In a wild-type background, deletions of NDJ1 interfere with synapsis and alter the distribution of recombination events. Ndj1p localizes to telomeres and is believed to be necessary for telomeric pairing (Chua and Roeder 1997; Conrad et al. 1997; Rockmill and Roeder 1998). Defects in synapsis and a decrease in the frequency of recombination are observed in the presence of mutations in yeast KAR3, which encodes a kinesin-like protein (Bascom-Slack and Dawson 1997). This observation suggests that Kar3p may be involved in chromosome movement that is necessary for proper homolog alignment during meiotic prophase.
This clustering of telomeres has not been demonstrated in Drosophila oocytes, but the observations by electron microscopy may not have detected transient telomere clustering at the leptotene or zygotene stage of meiosis (Carpenter 1975a,b, 1979a,b; Dernburg et al. 1995). Thus, perhaps the association of telomeres is important for homologous chromosome pairing in Drosophila. The mechanism by which telomere clustering occurs may be similar to that in other organisms. In particular, the movement of the chromosomes may require associations with the cytoskeleton, resulting in the clustering of telomeres near centrosomes. Within germline cysts, the fusome acts to orient the spindles in the cystocyte divisions by interacting with one centrosome at each mitosis (Deng and Lin 1997). Although the fusome begins to break down after the cessation of the mitotic divisions, remnants of this structure may remain in the oocyte during meiotic prophase (germarium regions 2 and 3; Lin et al. 1994; de Cuevas and Spradling 1998). Therefore, the fusome may position a determinant, possibly in the form of a cytoskeletal element, necessary for telomere clustering at a point on or near the nuclear membrane. Perhaps Bam, as a component of the fusome, is indirectly necessary for telomere clustering by marking the site of clustering on the nuclear membrane. The polar recombination defect observed in mei-P26 could therefore be the indirect result of abnormal telomeric clustering due to a disruption of Bam function.
Since proximal euchromatic regions may rely on the pairing of centric heterochromatin, and perhaps not require telomeric clustering, disruption of telomeric clustering may primarily impact the frequency of exchange in the distal regions of the chromosomes. Thus, the specific disruption of telomeric interactions may result in a distribution of residual exchanges like that seen in recombination precondition mutants. Further work will be necessary to determine whether the meiotic defects in precondition mutants are the result of abnormalities in telomeric clustering.
Acknowledgement
We thank Dennis McKearin and Pedro Santamaria for providing fly stocks and Annette Lentz and Ken Shaw for assistance with embryo microinjections. We also thank Jeff Sekelsky and members of the Hawley lab for helpful discussions and critical reading of this manuscript. S.L.P. was supported by a fellowship from the Cancer Research Fund of the Damon Runyon-Walter Winchell Foundation (DRG-1476). This work was supported by a grant to R.S.H. from the National Institutes of Health.
Footnotes
Communicating editor: S. Henikoff
LITERATURE CITED
Author notes
Present address: Molecular Biology Institute, University of California, Los Angeles, CA 90096.
Present address: Department of Anatomy, University of California San Francisco School of Medicine, San Francisco, CA 94143.