-
PDF
- Split View
-
Views
-
Cite
Cite
Chand Desai, Joy Purdy, The Neural Receptor Protein Tyrosine Phosphatase DPTP69D Is Required During Periods of Axon Outgrowth in Drosophila, Genetics, Volume 164, Issue 2, 1 June 2003, Pages 575–588, https://doi.org/10.1093/genetics/164.2.575
Close - Share Icon Share
Abstract
We have isolated and characterized a series of 18 chemically induced alleles of Ptp69D ranging in strength from viable to worse than null, which represent unique tools for probing the structure, function, and signaling pathway of DPTP69D. Three alleles are strongly temperature sensitive and were used to define the developmental periods requiring DPTP69D function; adult health requires DPTP69D during the mid- to late-pupal stage, eclosion requires DPTP69D during the early to mid-larval stage, and larval survival requires DPTP69D during embryogenesis. Mutations predicted to abolish the phosphatase activity of the membrane proximal D1 domain severely reduce but do not abolish DPTP69D function. Six alleles appear null; only 20% of null homozygotes pupate and <5% eclose, only to fall into the food and drown. One allele, Ptp69D7, confers axon and viability defects more severe than those of the null phenotype. Sequence analysis predicts that Ptp69D7 encodes a mutant protein that may bind but not release substrate. Like mutations in the protein tyrosine phosphatase gene Dlar, strong Ptp69D alleles cause the ISNb nerve to bypass its muscle targets. Genetic analysis reveals that the bypass defect in Dlar and Ptp69D mutants is dependent upon DPTP99A function, consistent with the hypothesis that DPTP69D and DLAR both counteract DPTP99A, allowing ISNb axons to enter their target muscle field.
AXONS navigate through a highly varied and dynamic embryonic environment to reach their synaptic targets. Along the way, differentially expressed proteins provide positional information that helps guide developing axons along appropriate pathways (Tessier-Lavigne and Goodman 1996). Growth cones at the tip of elongating axons express a large variety of receptor molecules that bind to these guidance cues. Binding of receptor to ligand in turn regulates the actin-based growth cone cytoskeleton via a variety of signal transduction mechanisms.
Tyrosine phosphorylation, one such signaling pathway, is emerging as a key regulator of actin polymerization during axon guidance. For example, Drosophila Ena, which promotes actin polymerization (Krause et al. 2002), is a substrate of the receptor tyrosine phosphatase DLAR in vitro and the cytoplasmic tyrosine kinase Abl in vitro and in vivo (Comer et al. 1998; Wills et al. 1999). Embryonically, Ena, DLAR, and Abl are all expressed in axons, suggesting that signal transduction involving tyrosine phosphorylation can affect axon guidance by modulating actin polymerization through Ena (Bennett and Hoffmann 1992; Gertler et al. 1995; Krueger et al. 1996; Comer et al. 1998).
Tyrosine phosphorylation also regulates axon guidance through the small GTPases Rac, Rho, and Cdc42. These G proteins regulate actin filament polymerization, branching, and bundling in part by regulating the actin depolymerizing protein Cofilin and the actin filament capping complex Arp2/3 (Meyer and Feldman 2002). Several lines of evidence indicate that Rac, Rho, and Cdc42 function in tyrosine-kinase-stimulated axon outgrowth. In Drosophila, Abl and DLAR interact genetically with Trio, a guanine nucleotide exchange factor for Rac, during embryonic motor axon guidance (Bateman et al. 2000; Liebl et al. 2000). In PC12 cells, nerve growth factor stimulation of the Trk tyrosine kinase induces neurite formation in a Rac- and Cdc42-dependent manner (Daniels et al. 1998). In cerebellar granule cells, stimulation of neurite outgrowth by L1, a neural cell adhesion molecule (CAM), requires the Src cytoplasmic tyrosine kinase, activation of the mitogen-activated protein kinase pathway, and Rac (Schmid et al. 2000). Finally, Ena, Abl, and DLAR, among several other tyrosine kinases and phosphatases, have been shown to be directly involved in axon guidance in Drosophila, and many of their vertebrate homologs also function in this process (Callahan et al. 1995; Desai et al. 1997b; Bashaw et al. 2000).
Although the mechanisms by which tyrosine kinases affect axon guidance are relatively well studied, much less is known about their enzymatic antagonists, the receptor-like protein tyrosine phosphatases (RPTPs). RPTPs comprise a diverse protein family and are expressed in the developing nervous systems of many animals (Van Vactor et al. 1993; Yan et al. 1993; Desai et al. 1994; Gershon et al. 1998; Ledig et al. 1999). Extracellularly, RPTPs resemble CAMs, often containing multiple immunoglobulin (Ig) domains and/or fibronectin (Fn) type III repeats. Intracellularly, RPTPs consist of one or two phosphatase domains. The ability of RPTPs to reverse the catalytic action of tyrosine kinases implicates both protein families in many of the same processes, including cell proliferation, adhesion, and migration. In Drosophila five RPTPs, DLAR, DPTP10, DPTP52F, DPTP69D, and DPTP99A, facilitate motor and central nervous system (CNS) axon guidance during embryogenesis (Krueger et al. 1996; Desai et al. 1997a; Sun et al. 2000; Schindelholz et al. 2001). DLAR and DPTP69D are also required for retinal axon guidance during late larval and pupal stages (Garrity et al. 1999; Clandinin et al. 2001; Maurel-Zaffran et al. 2001). To date, the molecular mechanisms that mediate these effects remain unclear, although the interactions among Dlar, Trio, Abl, and Ena indicate that regulation of actin polymerization is a key outcome.
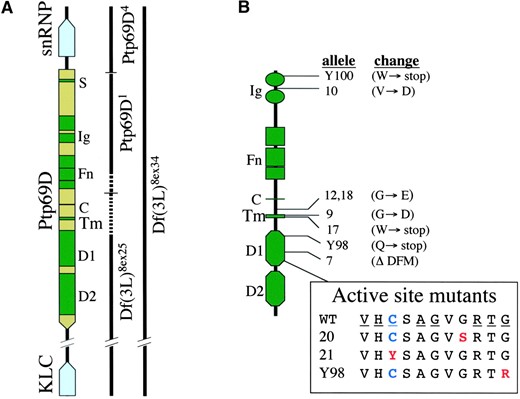
To further elucidate RPTP signaling pathways, we have undertaken a detailed genetic structure-function analysis of DPTP69D. DPTP69D contains two Ig and 3 Fn domains extracellularly and two phosphatase domains intracellularly (Figure 1B). DPTP69D is cleaved at an extracellular RDKR site ∼90 residues from the trans-membrane segment, but the cleavage products remain associated (P. Snow, personal communication). DPT-P69D is particularly interesting because unlike Ptp10D, Ptp99A, and Dlar, Ptp69D is an essential gene. Mutations in Ptp69D confer mild motor axon defects in the embryo and highly penetrant retinal axon defects in the larva (Desai et al. 1996; Garrity et al. 1999; Newsome et al. 2000), while Ptp10D and Ptp99A mutants appear wild type. Interestingly, Ptp69D Ptp99A and Ptp10D;Ptp69D double mutants display severe and highly penetrant defects in embryonic motor and central axons, respectively (Desai et al. 1996; Sun et al. 2000). In this article, we describe the isolation and characterization of 18 new alleles of Ptp69D. Utilizing these alleles, we demonstrate that the membrane-proximal phosphatase domain is critical for function and that DPTP69D is required during developmental stages coinciding with periods of axon outgrowth. This panel of alleles will also serve as genetic tools useful for delineating the DPTP69D signal transduction pathway.
—Ptp69D genomic region, protein, and mutants. (A) The rectangles represent sequences transcribed into RNA, with the pointed end showing the direction of transcription: green for Ptp69D and blue for KLC and snRNP. The dark green boxes signify sequences that encode motifs within the DPTP69D protein: S, signal sequence; Ig, immunoglobulin-like domains; Fn, fibronectin type III repeats; C, cleavage site; Tm, transmembrane domain; D1 and D2, the first and second PTP enzymatic domains. The lines to the right of the boxes indicate sequences removed in the indicated Ptp69D deletion alleles. Breakpoints occur within the dotted segments of the lines. (B) Cartoon showing DPTP69D domains and mutation sites. The lines link the mutation site to the Ptp69D allele and associated coding change. WT, the sequence of amino acid residues comprising the D1 active site, with the catalytic cysteine indicated in blue. The consensus sequence residues are underlined. The changes found in the three active site mutants are indicated in red.
MATERIALS AND METHODS
Mutagenesis: About 100 3- to 5-day-old Oregon R or w; 11F3 males were places into bottles containing paper saturated with a solution of 3% sugar, 4% green food color, and 25 mm EMS (Sigma, St. Louis). After 6 hr, the flies were examined and those with green abdomens were returned to the bottle. After 24 hr, the greenest males were placed on fresh food and allowed to recover. Twenty-four hours later, nongreen males, usually between 10 and 30 in number, were mated en mass with 50–100 virgin w; TM3/TM6B females. Males and females were removed after 4 and 8 days, respectively. F1 males were individually mated to w; Df(3L)8ex34/TM6B or w; Ptp69D1/TM6B virgins. Vials producing no unbalanced F2 progeny were tested further.
Genetics: Strains that failed to complement all four deletion alleles were considered to harbor new Ptp69D alleles, propagated as balanced stocks, and backcrossed by recombining the markers ru h th st cu sr e ca onto the mutant chromosomes. Single w; ru h Ptp69D* th st cu sr e ca/TM6B males were then mated with w1118 females and the markers were removed by recombination. This procedure resulted in two recombination events within 12% genetic distance of Ptp69D on the left and two events within 4% to the right, removing most or all nonselected mutations induced during mutagenesis. The Ptp69D transgene used to test for extraneous lethal mutations is described in Desai et al. (1996).
Immunohistochemistry: Twenty-four-hour embryo collections were fixed and prepared for immunohistochemistry as
Trans-heterozygous viability at 25°
| . | Df(3L)8ex34 . | Ptp69D1 . | Df(3L)8ex25 . | Ptp69D4 . | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Allele . | A . | E . | P . | A . | E . | P . | A . | E . | P . | A . | E . | P . |
| Ptp69D1 | 0 | 0 | 8 | NA | NA | NA | 0 | 7 | 28 | NA | NA | NA |
| Ptp69D4 | 0a | 0a | 20a | NA | NA | NA | 0 | 12 | 48 | NA | NA | NA |
| Ptp69D7 | 0 | 1 | 7 | 0 | 0 | 9 | 0 | 0 | 5 | 0 | 5 | 21 |
| Ptp69Dy100 | 0 | 0 | 9 | 0 | 1 | 11 | 0 | 0 | 10 | 0 | 5 | 24 |
| Ptp69Dy98 | 0 | 0 | 11 | 0 | 0 | 14 | 0 | 2 | 18 | 0 | 4 | 23 |
| Ptp69D11 | 0 | 2 | 10 | 0 | 0 | 7 | 0 | 2 | 21 | 0 | 5 | 17 |
| Ptp69D21 | 0 | 0 | 6 | 0 | 2 | 10 | 0 | 4 | 14 | 0 | 8 | 30 |
| Ptp69Dy125 | 0 | 4 | 24 | 0 | 1 | 13 | 0 | 2 | 12 | 0 | 3 | 21 |
| Ptp69D16 | 0 | 0 | 1 | 0 | 0 | 13 | 0 | 0 | 0 | 0 | 3 | 27 |
| Ptp69D6 | 0 | 1 | 14 | 0 | 2 | 13 | 0 | 5 | 18 | 0 | 15 | 29 |
| Ptp69D8 | 0 | 5 | 10 | 0 | 2 | 17 | 0 | 9 | 24 | 0 | 6 | 23 |
| Ptp69D14 | 0 | 1 | 15 | 0 | 3 | 18 | 0 | 7 | 22 | 0 | 7 | 24 |
| Ptp69D17 | 0 | 2 | 18 | 0 | 4 | 11 | 0 | 5 | 18 | 0 | 13 | 29 |
| Ptp69D15 | 0 | 0 | 8 | 0 | 4 | 16 | 2 | 12 | 33 | 0 | 15 | 32 |
| Ptp69D13 | 0 | 2 | 25 | 0 | 4 | 18 | 0 | 9 | 31 | 0 | 10 | 29 |
| Ptp69D22 | 0 | 3 | 35 | 1 | 5 | 27 | 0 | 1 | 25 | 0 | 8 | 42 |
| Ptp69D9 | 0 | 6 | 24 | 0 | 23 | 41 | 1 | 19 | 34 | 0 | 9 | 25 |
| Ptp69D5 | 0 | 7 | 27 | 0 | 3 | 17 | 0 | 25 | 47 | 1 | 27 | 48 |
| Ptp69D19 | 0 | 4 | 19 | 0 | 10 | 36 | 0 | 15 | 46 | 1 | 25 | 52 |
| Ptp69D12 | 0 | 4 | 44 | 0 | 5 | 26 | 0 | 21 | 61 | 0 | 10 | 36 |
| Ptp69D18 | 0 | 7 | 36 | 0 | 7 | 27 | 0 | 13 | 49 | 0 | 35 | 56 |
| Ptp69D10 | 13 | 69 | 81 | 10 | 88 | 96 | 12 | 60 | 68 | 35 | 70 | 78 |
| Ptp69D20 | 34 | 72 | 85 | 20 | 60 | 67 | 65 | 85 | 88 | 41 | 89 | 100 |
| . | Df(3L)8ex34 . | Ptp69D1 . | Df(3L)8ex25 . | Ptp69D4 . | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Allele . | A . | E . | P . | A . | E . | P . | A . | E . | P . | A . | E . | P . |
| Ptp69D1 | 0 | 0 | 8 | NA | NA | NA | 0 | 7 | 28 | NA | NA | NA |
| Ptp69D4 | 0a | 0a | 20a | NA | NA | NA | 0 | 12 | 48 | NA | NA | NA |
| Ptp69D7 | 0 | 1 | 7 | 0 | 0 | 9 | 0 | 0 | 5 | 0 | 5 | 21 |
| Ptp69Dy100 | 0 | 0 | 9 | 0 | 1 | 11 | 0 | 0 | 10 | 0 | 5 | 24 |
| Ptp69Dy98 | 0 | 0 | 11 | 0 | 0 | 14 | 0 | 2 | 18 | 0 | 4 | 23 |
| Ptp69D11 | 0 | 2 | 10 | 0 | 0 | 7 | 0 | 2 | 21 | 0 | 5 | 17 |
| Ptp69D21 | 0 | 0 | 6 | 0 | 2 | 10 | 0 | 4 | 14 | 0 | 8 | 30 |
| Ptp69Dy125 | 0 | 4 | 24 | 0 | 1 | 13 | 0 | 2 | 12 | 0 | 3 | 21 |
| Ptp69D16 | 0 | 0 | 1 | 0 | 0 | 13 | 0 | 0 | 0 | 0 | 3 | 27 |
| Ptp69D6 | 0 | 1 | 14 | 0 | 2 | 13 | 0 | 5 | 18 | 0 | 15 | 29 |
| Ptp69D8 | 0 | 5 | 10 | 0 | 2 | 17 | 0 | 9 | 24 | 0 | 6 | 23 |
| Ptp69D14 | 0 | 1 | 15 | 0 | 3 | 18 | 0 | 7 | 22 | 0 | 7 | 24 |
| Ptp69D17 | 0 | 2 | 18 | 0 | 4 | 11 | 0 | 5 | 18 | 0 | 13 | 29 |
| Ptp69D15 | 0 | 0 | 8 | 0 | 4 | 16 | 2 | 12 | 33 | 0 | 15 | 32 |
| Ptp69D13 | 0 | 2 | 25 | 0 | 4 | 18 | 0 | 9 | 31 | 0 | 10 | 29 |
| Ptp69D22 | 0 | 3 | 35 | 1 | 5 | 27 | 0 | 1 | 25 | 0 | 8 | 42 |
| Ptp69D9 | 0 | 6 | 24 | 0 | 23 | 41 | 1 | 19 | 34 | 0 | 9 | 25 |
| Ptp69D5 | 0 | 7 | 27 | 0 | 3 | 17 | 0 | 25 | 47 | 1 | 27 | 48 |
| Ptp69D19 | 0 | 4 | 19 | 0 | 10 | 36 | 0 | 15 | 46 | 1 | 25 | 52 |
| Ptp69D12 | 0 | 4 | 44 | 0 | 5 | 26 | 0 | 21 | 61 | 0 | 10 | 36 |
| Ptp69D18 | 0 | 7 | 36 | 0 | 7 | 27 | 0 | 13 | 49 | 0 | 35 | 56 |
| Ptp69D10 | 13 | 69 | 81 | 10 | 88 | 96 | 12 | 60 | 68 | 35 | 70 | 78 |
| Ptp69D20 | 34 | 72 | 85 | 20 | 60 | 67 | 65 | 85 | 88 | 41 | 89 | 100 |
The adult survival (A), eclosion (E), and pupation (P) rates (in percent) of mutant progeny from crosses of Ptp69D alleles (left column) with deletion alleles (top row) are indicated. Alleles are ranked from strongest to weakest, starting at the top. More than 200 progeny were counted for each cross. NA, not applicable (some crosses were not performed because of shared second-site mutations).
Data are from w; Ptp69D4 P[snRNP-1.5]/TM6B × w; Df(3L)8ex34/TM6B. P[snRNP-1.5] contains genomic sequences encoding the snRNP69D gene, which is deleted in both Ptp69D4 and Df(3L)8ex34.
Trans-heterozygous viability at 25°
| . | Df(3L)8ex34 . | Ptp69D1 . | Df(3L)8ex25 . | Ptp69D4 . | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Allele . | A . | E . | P . | A . | E . | P . | A . | E . | P . | A . | E . | P . |
| Ptp69D1 | 0 | 0 | 8 | NA | NA | NA | 0 | 7 | 28 | NA | NA | NA |
| Ptp69D4 | 0a | 0a | 20a | NA | NA | NA | 0 | 12 | 48 | NA | NA | NA |
| Ptp69D7 | 0 | 1 | 7 | 0 | 0 | 9 | 0 | 0 | 5 | 0 | 5 | 21 |
| Ptp69Dy100 | 0 | 0 | 9 | 0 | 1 | 11 | 0 | 0 | 10 | 0 | 5 | 24 |
| Ptp69Dy98 | 0 | 0 | 11 | 0 | 0 | 14 | 0 | 2 | 18 | 0 | 4 | 23 |
| Ptp69D11 | 0 | 2 | 10 | 0 | 0 | 7 | 0 | 2 | 21 | 0 | 5 | 17 |
| Ptp69D21 | 0 | 0 | 6 | 0 | 2 | 10 | 0 | 4 | 14 | 0 | 8 | 30 |
| Ptp69Dy125 | 0 | 4 | 24 | 0 | 1 | 13 | 0 | 2 | 12 | 0 | 3 | 21 |
| Ptp69D16 | 0 | 0 | 1 | 0 | 0 | 13 | 0 | 0 | 0 | 0 | 3 | 27 |
| Ptp69D6 | 0 | 1 | 14 | 0 | 2 | 13 | 0 | 5 | 18 | 0 | 15 | 29 |
| Ptp69D8 | 0 | 5 | 10 | 0 | 2 | 17 | 0 | 9 | 24 | 0 | 6 | 23 |
| Ptp69D14 | 0 | 1 | 15 | 0 | 3 | 18 | 0 | 7 | 22 | 0 | 7 | 24 |
| Ptp69D17 | 0 | 2 | 18 | 0 | 4 | 11 | 0 | 5 | 18 | 0 | 13 | 29 |
| Ptp69D15 | 0 | 0 | 8 | 0 | 4 | 16 | 2 | 12 | 33 | 0 | 15 | 32 |
| Ptp69D13 | 0 | 2 | 25 | 0 | 4 | 18 | 0 | 9 | 31 | 0 | 10 | 29 |
| Ptp69D22 | 0 | 3 | 35 | 1 | 5 | 27 | 0 | 1 | 25 | 0 | 8 | 42 |
| Ptp69D9 | 0 | 6 | 24 | 0 | 23 | 41 | 1 | 19 | 34 | 0 | 9 | 25 |
| Ptp69D5 | 0 | 7 | 27 | 0 | 3 | 17 | 0 | 25 | 47 | 1 | 27 | 48 |
| Ptp69D19 | 0 | 4 | 19 | 0 | 10 | 36 | 0 | 15 | 46 | 1 | 25 | 52 |
| Ptp69D12 | 0 | 4 | 44 | 0 | 5 | 26 | 0 | 21 | 61 | 0 | 10 | 36 |
| Ptp69D18 | 0 | 7 | 36 | 0 | 7 | 27 | 0 | 13 | 49 | 0 | 35 | 56 |
| Ptp69D10 | 13 | 69 | 81 | 10 | 88 | 96 | 12 | 60 | 68 | 35 | 70 | 78 |
| Ptp69D20 | 34 | 72 | 85 | 20 | 60 | 67 | 65 | 85 | 88 | 41 | 89 | 100 |
| . | Df(3L)8ex34 . | Ptp69D1 . | Df(3L)8ex25 . | Ptp69D4 . | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Allele . | A . | E . | P . | A . | E . | P . | A . | E . | P . | A . | E . | P . |
| Ptp69D1 | 0 | 0 | 8 | NA | NA | NA | 0 | 7 | 28 | NA | NA | NA |
| Ptp69D4 | 0a | 0a | 20a | NA | NA | NA | 0 | 12 | 48 | NA | NA | NA |
| Ptp69D7 | 0 | 1 | 7 | 0 | 0 | 9 | 0 | 0 | 5 | 0 | 5 | 21 |
| Ptp69Dy100 | 0 | 0 | 9 | 0 | 1 | 11 | 0 | 0 | 10 | 0 | 5 | 24 |
| Ptp69Dy98 | 0 | 0 | 11 | 0 | 0 | 14 | 0 | 2 | 18 | 0 | 4 | 23 |
| Ptp69D11 | 0 | 2 | 10 | 0 | 0 | 7 | 0 | 2 | 21 | 0 | 5 | 17 |
| Ptp69D21 | 0 | 0 | 6 | 0 | 2 | 10 | 0 | 4 | 14 | 0 | 8 | 30 |
| Ptp69Dy125 | 0 | 4 | 24 | 0 | 1 | 13 | 0 | 2 | 12 | 0 | 3 | 21 |
| Ptp69D16 | 0 | 0 | 1 | 0 | 0 | 13 | 0 | 0 | 0 | 0 | 3 | 27 |
| Ptp69D6 | 0 | 1 | 14 | 0 | 2 | 13 | 0 | 5 | 18 | 0 | 15 | 29 |
| Ptp69D8 | 0 | 5 | 10 | 0 | 2 | 17 | 0 | 9 | 24 | 0 | 6 | 23 |
| Ptp69D14 | 0 | 1 | 15 | 0 | 3 | 18 | 0 | 7 | 22 | 0 | 7 | 24 |
| Ptp69D17 | 0 | 2 | 18 | 0 | 4 | 11 | 0 | 5 | 18 | 0 | 13 | 29 |
| Ptp69D15 | 0 | 0 | 8 | 0 | 4 | 16 | 2 | 12 | 33 | 0 | 15 | 32 |
| Ptp69D13 | 0 | 2 | 25 | 0 | 4 | 18 | 0 | 9 | 31 | 0 | 10 | 29 |
| Ptp69D22 | 0 | 3 | 35 | 1 | 5 | 27 | 0 | 1 | 25 | 0 | 8 | 42 |
| Ptp69D9 | 0 | 6 | 24 | 0 | 23 | 41 | 1 | 19 | 34 | 0 | 9 | 25 |
| Ptp69D5 | 0 | 7 | 27 | 0 | 3 | 17 | 0 | 25 | 47 | 1 | 27 | 48 |
| Ptp69D19 | 0 | 4 | 19 | 0 | 10 | 36 | 0 | 15 | 46 | 1 | 25 | 52 |
| Ptp69D12 | 0 | 4 | 44 | 0 | 5 | 26 | 0 | 21 | 61 | 0 | 10 | 36 |
| Ptp69D18 | 0 | 7 | 36 | 0 | 7 | 27 | 0 | 13 | 49 | 0 | 35 | 56 |
| Ptp69D10 | 13 | 69 | 81 | 10 | 88 | 96 | 12 | 60 | 68 | 35 | 70 | 78 |
| Ptp69D20 | 34 | 72 | 85 | 20 | 60 | 67 | 65 | 85 | 88 | 41 | 89 | 100 |
The adult survival (A), eclosion (E), and pupation (P) rates (in percent) of mutant progeny from crosses of Ptp69D alleles (left column) with deletion alleles (top row) are indicated. Alleles are ranked from strongest to weakest, starting at the top. More than 200 progeny were counted for each cross. NA, not applicable (some crosses were not performed because of shared second-site mutations).
Data are from w; Ptp69D4 P[snRNP-1.5]/TM6B × w; Df(3L)8ex34/TM6B. P[snRNP-1.5] contains genomic sequences encoding the snRNP69D gene, which is deleted in both Ptp69D4 and Df(3L)8ex34.
described (Patel 1994). Antibody binding was visualized using horseradish-peroxidase-conjugated goat anti-mouse or anti-rabbit antibodies and diaminobenzidine immunohistochemistry. Ptp69D mutant embryos were identified by their failure to stain with anti-DPTP69D mAbs 3F11, 3F12, or 5A6 (Desai et al. 1996), or anti-β-galactosidase sera (Jackson Laboratories, Bar Harbor, ME; expressed by TM6B T8-lacZ balancer chromosome, gift of Peter Kolodziej); sorted; restained with mAb 1D4 to visualize motor axons; cleared in 70% glycerol::30% PBS; and dissected. Embryos were examined on an Olympus AX-70 microscope using differential interference contrast optics and photographed using a Spot-cooled CCD camera.
Temperature-shift experiments: Four vials containing 15–20 w; Ptp69D1/TM6B virgin females were crossed to 10–15 w; Ptp69D10/TM6B males at 25°. These flies were transferred to new food every 12 hr, and dead flies were replaced to maintain progeny counts. When F1 flies began to eclose from the first group of four vials, the entire set was transferred to 18°. Flies were counted and removed daily from each group, until no new flies eclosed in a 24-hr period. At that time, hatched and unhatched pupae were counted. Similar experiments were carried out with crosses of w; Df(3L)8ex34/TM6B females with w; Ptp69D12/TM6B or w; Ptp69D18/TM6B, except that these crosses were first maintained at 18° and the parents were transferred to new food every 24 hr.
Molecular biology: Genomic DNA from single embryos: Stage 16 embryos were bleach dechorionated, rinsed, picked individually into siliconized microtiter plate wells (Sigmacote), devitellinized in 5 μl of 1:1 heptane/methanol, and washed twice with 10 μl 50 mm KCl, 10 mm Tris pH 8.3, and 0.05 mm EDTA. After removing the fluid, embryos were pulverized with siliconized toothpicks, resuspended in 4 μl 50 mm KCl, 10 mm Tris pH 8.3, 0.5% Tween, 0.5% NP-40, and transferred to a tube containing 4 μl “last buffer” (50 mm KCl, 10 mm Tris pH 8.3). Each well was rinsed with 4 μl last buffer and this was added to the appropriate tube, which was stored on dry ice. Embryo lysates were digested with 8 μl last buffer
Trans-heterozygous viability at 18°
| . | Df(3L)8ex34 . | Ptp69D1 . | Df(3L)8ex25 . | Ptp69D4 . | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Allele . | A . | E . | P . | A . | E . | P . | A . | E . | P . | A . | E . | P . |
| Ptp69D1 | 0 | 1 | 20 | NA | NA | NA | 0 | 9 | 46 | NA | NA | NA |
| Ptp69D4 | 0a | 0a | 44a | NA | NA | NA | 0 | 10 | 39 | NA | NA | NA |
| Ptp69D11 | 0 | 2 | 21 | 0 | 4 | 15 | 0 | 4 | 24 | 1 | 6 | 25 |
| Ptp69D7 | 0 | 1 | 14 | 1 | 1 | 13 | 0 | 6 | 34 | 1 | 6 | 34 |
| Ptp69Dy100 | 0 | 2 | 19 | 0 | 5 | 14 | 0 | 9 | 25 | 1 | 6 | 39 |
| Ptp69D8 | 0 | 3 | 19 | 0 | 4 | 19 | 0 | 7 | 36 | 0 | 10 | 29 |
| Ptp69Dy98 | 0 | 0 | 25 | 0 | 6 | 30 | 0 | 10 | 36 | 0 | 7 | 36 |
| Ptp69D16 | 0 | 0 | 0 | 0 | 2 | 28 | 0 | 0 | 0 | 0 | 7 | 52 |
| Ptp69D15 | 1 | 5 | 18 | 0 | 4 | 22 | 0 | 5 | 34 | 0 | 14 | 41 |
| Ptp69D14 | 0 | 2 | 22 | 1 | 3 | 22 | 2 | 9 | 34 | 0 | 12 | 44 |
| Ptp69Dy125 | 0 | 6 | 33 | 0 | 0 | 12 | 1 | 3 | 25 | 1 | 12 | 39 |
| Ptp69D21 | 0 | 4 | 32 | 0 | 2 | 17 | 0 | 13 | 43 | 0 | 18 | 35 |
| Ptp69D17 | 0 | 9 | 16 | 0 | 6 | 31 | 0 | 8 | 34 | 0 | 23 | 50 |
| Ptp69D13 | 1 | 3 | 36 | 0 | 9 | 43 | 1 | 14 | 38 | 3 | 14 | 43 |
| Ptp69D5 | 0 | 8 | 19 | 0 | 16 | 42 | 0 | 18 | 37 | 0 | 27 | 48 |
| Ptp69D9 | 0 | 17 | 26 | 0 | 13 | 26 | 1 | 26 | 51 | 0 | 18 | 39 |
| Ptp69D6 | 0 | 16 | 30 | 0 | 13 | 29 | 1 | 23 | 55 | 0 | 23 | 46 |
| Ptp69D22 | 1 | 19 | 74 | 0 | 11 | 59 | 0 | 13 | 63 | 0 | 38 | 76 |
| Ptp69D19 | 1 | 20 | 47 | 0 | 18 | 59 | 2 | 22 | 67 | 3 | 40 | 76 |
| Ptp69D12 | 73 | 84 | 86 | 63 | 63 | 62 | 78 | 78 | 79 | 68 | 67 | 69 |
| Ptp69D20 | 54 | 78 | 97 | 54 | 61 | 68 | 92 | 100 | 92 | 77 | 79 | 86 |
| Ptp69D18 | 98 | 99 | 108 | 80 | 80 | 82 | 81 | 81 | 81 | 84 | 84 | 88 |
| Ptp69D10 | 99 | 101 | 128 | 109 | 116 | 106 | 107 | 107 | 106 | 111 | 112 | 104 |
| . | Df(3L)8ex34 . | Ptp69D1 . | Df(3L)8ex25 . | Ptp69D4 . | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Allele . | A . | E . | P . | A . | E . | P . | A . | E . | P . | A . | E . | P . |
| Ptp69D1 | 0 | 1 | 20 | NA | NA | NA | 0 | 9 | 46 | NA | NA | NA |
| Ptp69D4 | 0a | 0a | 44a | NA | NA | NA | 0 | 10 | 39 | NA | NA | NA |
| Ptp69D11 | 0 | 2 | 21 | 0 | 4 | 15 | 0 | 4 | 24 | 1 | 6 | 25 |
| Ptp69D7 | 0 | 1 | 14 | 1 | 1 | 13 | 0 | 6 | 34 | 1 | 6 | 34 |
| Ptp69Dy100 | 0 | 2 | 19 | 0 | 5 | 14 | 0 | 9 | 25 | 1 | 6 | 39 |
| Ptp69D8 | 0 | 3 | 19 | 0 | 4 | 19 | 0 | 7 | 36 | 0 | 10 | 29 |
| Ptp69Dy98 | 0 | 0 | 25 | 0 | 6 | 30 | 0 | 10 | 36 | 0 | 7 | 36 |
| Ptp69D16 | 0 | 0 | 0 | 0 | 2 | 28 | 0 | 0 | 0 | 0 | 7 | 52 |
| Ptp69D15 | 1 | 5 | 18 | 0 | 4 | 22 | 0 | 5 | 34 | 0 | 14 | 41 |
| Ptp69D14 | 0 | 2 | 22 | 1 | 3 | 22 | 2 | 9 | 34 | 0 | 12 | 44 |
| Ptp69Dy125 | 0 | 6 | 33 | 0 | 0 | 12 | 1 | 3 | 25 | 1 | 12 | 39 |
| Ptp69D21 | 0 | 4 | 32 | 0 | 2 | 17 | 0 | 13 | 43 | 0 | 18 | 35 |
| Ptp69D17 | 0 | 9 | 16 | 0 | 6 | 31 | 0 | 8 | 34 | 0 | 23 | 50 |
| Ptp69D13 | 1 | 3 | 36 | 0 | 9 | 43 | 1 | 14 | 38 | 3 | 14 | 43 |
| Ptp69D5 | 0 | 8 | 19 | 0 | 16 | 42 | 0 | 18 | 37 | 0 | 27 | 48 |
| Ptp69D9 | 0 | 17 | 26 | 0 | 13 | 26 | 1 | 26 | 51 | 0 | 18 | 39 |
| Ptp69D6 | 0 | 16 | 30 | 0 | 13 | 29 | 1 | 23 | 55 | 0 | 23 | 46 |
| Ptp69D22 | 1 | 19 | 74 | 0 | 11 | 59 | 0 | 13 | 63 | 0 | 38 | 76 |
| Ptp69D19 | 1 | 20 | 47 | 0 | 18 | 59 | 2 | 22 | 67 | 3 | 40 | 76 |
| Ptp69D12 | 73 | 84 | 86 | 63 | 63 | 62 | 78 | 78 | 79 | 68 | 67 | 69 |
| Ptp69D20 | 54 | 78 | 97 | 54 | 61 | 68 | 92 | 100 | 92 | 77 | 79 | 86 |
| Ptp69D18 | 98 | 99 | 108 | 80 | 80 | 82 | 81 | 81 | 81 | 84 | 84 | 88 |
| Ptp69D10 | 99 | 101 | 128 | 109 | 116 | 106 | 107 | 107 | 106 | 111 | 112 | 104 |
The adult survival (A), eclosion (E), and pupation (P) rates (in percent) of mutant progeny from crosses of Ptp69D alleles (left column) with deletion alleles (top row) are indicated. Alleles are ranked from strongest to weakest, starting at the top. More than 200 progeny were counted for each cross. NA, not applicable.
Data are from w; Ptp69D4 P[snRNP-1.5]/TM6B × w; Df(3L)8ex34/TM6B.
Trans-heterozygous viability at 18°
| . | Df(3L)8ex34 . | Ptp69D1 . | Df(3L)8ex25 . | Ptp69D4 . | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Allele . | A . | E . | P . | A . | E . | P . | A . | E . | P . | A . | E . | P . |
| Ptp69D1 | 0 | 1 | 20 | NA | NA | NA | 0 | 9 | 46 | NA | NA | NA |
| Ptp69D4 | 0a | 0a | 44a | NA | NA | NA | 0 | 10 | 39 | NA | NA | NA |
| Ptp69D11 | 0 | 2 | 21 | 0 | 4 | 15 | 0 | 4 | 24 | 1 | 6 | 25 |
| Ptp69D7 | 0 | 1 | 14 | 1 | 1 | 13 | 0 | 6 | 34 | 1 | 6 | 34 |
| Ptp69Dy100 | 0 | 2 | 19 | 0 | 5 | 14 | 0 | 9 | 25 | 1 | 6 | 39 |
| Ptp69D8 | 0 | 3 | 19 | 0 | 4 | 19 | 0 | 7 | 36 | 0 | 10 | 29 |
| Ptp69Dy98 | 0 | 0 | 25 | 0 | 6 | 30 | 0 | 10 | 36 | 0 | 7 | 36 |
| Ptp69D16 | 0 | 0 | 0 | 0 | 2 | 28 | 0 | 0 | 0 | 0 | 7 | 52 |
| Ptp69D15 | 1 | 5 | 18 | 0 | 4 | 22 | 0 | 5 | 34 | 0 | 14 | 41 |
| Ptp69D14 | 0 | 2 | 22 | 1 | 3 | 22 | 2 | 9 | 34 | 0 | 12 | 44 |
| Ptp69Dy125 | 0 | 6 | 33 | 0 | 0 | 12 | 1 | 3 | 25 | 1 | 12 | 39 |
| Ptp69D21 | 0 | 4 | 32 | 0 | 2 | 17 | 0 | 13 | 43 | 0 | 18 | 35 |
| Ptp69D17 | 0 | 9 | 16 | 0 | 6 | 31 | 0 | 8 | 34 | 0 | 23 | 50 |
| Ptp69D13 | 1 | 3 | 36 | 0 | 9 | 43 | 1 | 14 | 38 | 3 | 14 | 43 |
| Ptp69D5 | 0 | 8 | 19 | 0 | 16 | 42 | 0 | 18 | 37 | 0 | 27 | 48 |
| Ptp69D9 | 0 | 17 | 26 | 0 | 13 | 26 | 1 | 26 | 51 | 0 | 18 | 39 |
| Ptp69D6 | 0 | 16 | 30 | 0 | 13 | 29 | 1 | 23 | 55 | 0 | 23 | 46 |
| Ptp69D22 | 1 | 19 | 74 | 0 | 11 | 59 | 0 | 13 | 63 | 0 | 38 | 76 |
| Ptp69D19 | 1 | 20 | 47 | 0 | 18 | 59 | 2 | 22 | 67 | 3 | 40 | 76 |
| Ptp69D12 | 73 | 84 | 86 | 63 | 63 | 62 | 78 | 78 | 79 | 68 | 67 | 69 |
| Ptp69D20 | 54 | 78 | 97 | 54 | 61 | 68 | 92 | 100 | 92 | 77 | 79 | 86 |
| Ptp69D18 | 98 | 99 | 108 | 80 | 80 | 82 | 81 | 81 | 81 | 84 | 84 | 88 |
| Ptp69D10 | 99 | 101 | 128 | 109 | 116 | 106 | 107 | 107 | 106 | 111 | 112 | 104 |
| . | Df(3L)8ex34 . | Ptp69D1 . | Df(3L)8ex25 . | Ptp69D4 . | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Allele . | A . | E . | P . | A . | E . | P . | A . | E . | P . | A . | E . | P . |
| Ptp69D1 | 0 | 1 | 20 | NA | NA | NA | 0 | 9 | 46 | NA | NA | NA |
| Ptp69D4 | 0a | 0a | 44a | NA | NA | NA | 0 | 10 | 39 | NA | NA | NA |
| Ptp69D11 | 0 | 2 | 21 | 0 | 4 | 15 | 0 | 4 | 24 | 1 | 6 | 25 |
| Ptp69D7 | 0 | 1 | 14 | 1 | 1 | 13 | 0 | 6 | 34 | 1 | 6 | 34 |
| Ptp69Dy100 | 0 | 2 | 19 | 0 | 5 | 14 | 0 | 9 | 25 | 1 | 6 | 39 |
| Ptp69D8 | 0 | 3 | 19 | 0 | 4 | 19 | 0 | 7 | 36 | 0 | 10 | 29 |
| Ptp69Dy98 | 0 | 0 | 25 | 0 | 6 | 30 | 0 | 10 | 36 | 0 | 7 | 36 |
| Ptp69D16 | 0 | 0 | 0 | 0 | 2 | 28 | 0 | 0 | 0 | 0 | 7 | 52 |
| Ptp69D15 | 1 | 5 | 18 | 0 | 4 | 22 | 0 | 5 | 34 | 0 | 14 | 41 |
| Ptp69D14 | 0 | 2 | 22 | 1 | 3 | 22 | 2 | 9 | 34 | 0 | 12 | 44 |
| Ptp69Dy125 | 0 | 6 | 33 | 0 | 0 | 12 | 1 | 3 | 25 | 1 | 12 | 39 |
| Ptp69D21 | 0 | 4 | 32 | 0 | 2 | 17 | 0 | 13 | 43 | 0 | 18 | 35 |
| Ptp69D17 | 0 | 9 | 16 | 0 | 6 | 31 | 0 | 8 | 34 | 0 | 23 | 50 |
| Ptp69D13 | 1 | 3 | 36 | 0 | 9 | 43 | 1 | 14 | 38 | 3 | 14 | 43 |
| Ptp69D5 | 0 | 8 | 19 | 0 | 16 | 42 | 0 | 18 | 37 | 0 | 27 | 48 |
| Ptp69D9 | 0 | 17 | 26 | 0 | 13 | 26 | 1 | 26 | 51 | 0 | 18 | 39 |
| Ptp69D6 | 0 | 16 | 30 | 0 | 13 | 29 | 1 | 23 | 55 | 0 | 23 | 46 |
| Ptp69D22 | 1 | 19 | 74 | 0 | 11 | 59 | 0 | 13 | 63 | 0 | 38 | 76 |
| Ptp69D19 | 1 | 20 | 47 | 0 | 18 | 59 | 2 | 22 | 67 | 3 | 40 | 76 |
| Ptp69D12 | 73 | 84 | 86 | 63 | 63 | 62 | 78 | 78 | 79 | 68 | 67 | 69 |
| Ptp69D20 | 54 | 78 | 97 | 54 | 61 | 68 | 92 | 100 | 92 | 77 | 79 | 86 |
| Ptp69D18 | 98 | 99 | 108 | 80 | 80 | 82 | 81 | 81 | 81 | 84 | 84 | 88 |
| Ptp69D10 | 99 | 101 | 128 | 109 | 116 | 106 | 107 | 107 | 106 | 111 | 112 | 104 |
The adult survival (A), eclosion (E), and pupation (P) rates (in percent) of mutant progeny from crosses of Ptp69D alleles (left column) with deletion alleles (top row) are indicated. Alleles are ranked from strongest to weakest, starting at the top. More than 200 progeny were counted for each cross. NA, not applicable.
Data are from w; Ptp69D4 P[snRNP-1.5]/TM6B × w; Df(3L)8ex34/TM6B.
containing 0.25 mg/ml Proteinase K (Sigma) for 1 hr at 50°, incubated at 94° for 10 min, and stored at –20°.
Embryo genotyping: DNA from individual embryos was used in tandem PCR reactions designed to detect the presence of the TM6B-T8 lacZ balancer. One microliter of lysate was added to 19 μl standard PCR buffer (Herculase; Stratagene, La Jolla, CA) containing 250 nm each lacZ primer (GCA GGC TTC TGC TTC AAT CAG CGT GCC and GTT TGC CGT CTG AAT TTG ACC TGA GCG). PCR reactions that lacked the ∼500-bp lacZ PCR product were diluted 1:100 and reamplified. Lysates that lacked the lacZ PCR product, but generated a Ptp69D PCR product, were considered to be mutant.
Sequencing: A total of 1–2 μl lysate from mutant embryos was used in a series of PCR reactions designed to amplify overlapping ∼1-kb fragments of the Ptp69D gene. These primary PCR reaction products were purified using the Ultra-Clean DNA purification kit (CLP), diluted 1:100, and reamplified using nested Ptp69D primers. Purified secondary PCR product (0.5 μl) was used in a BigDye (ABI, Columbia, MD) sequencing reaction containing 4 μl SeqSaver (Sigma) in a volume of 20 μl. Sequencing samples were purified by ethanol precipitation and submitted to the sequencing core at Vanderbilt University Medical Center. Sequences were aligned to a GenBank sequence of Ptp69D using Vector (Burlingame, CA) NTI software. Mutations were confirmed by forward and reverse primers in the same embryo and by sequencing the same region of the gene in a second mutant embryo.
Ranking of alleles: Alleles are ranked in Tables 1,2,3 according to penetrance and expressivity of their viability defects with the strongest alleles at the top. The rankings were determined by comparing the sum of the expected percentages of healthy free-walking adults, total adults, and total pupae produced by each genotype. The ranking in Table 5 was determined by comparing the sum of the rates of total and severe ISNb defects, ISN defects, SNa defects, and SNc defects.
RESULTS
Existing Ptp69D alleles: Previous genetic studies of Ptp69D utilized four deletion alleles derived from the imprecise excision of two P elements (Desai et al. 1996). Two derive from 59A, an insertion 3′ to Ptp69D in the nearby kinesis light chain (KLC) gene (Figure 1A; Gindhart et al. 1998). These are Df(3L)8ex25, which lacks sequences that encode most of the cytoplasmic domains, and Df(3L)8ex34, which lacks the entire Ptp69D gene and many other essential complementation groups. Both also lack two genes located between Ptp69D and KLC, CG32112 and CG10984, at least one of which is essential (our unpublished observations). The other two deficiencies derive from 11F3, an insertion into the 5′ untranslated region of Ptp69D:Ptp69D1, which lacks all but the first 115 bp of the 5′ untranslated region (UTR), the translational start site, and sequences that encode most of the extracellular domain; and Ptp69D4, which lacks the first 115 bp of the 5′ UTR, the entire promoter, and the nearby small nuclear ribonucleoprotein (snRNP-69D) gene. Ptp69D1 and Ptp69D4 are also mutant for mirror (mirr), a nearby transcription factor gene. Embryos homozygous for each of the deletion alleles fail to express DPTP69D, as assayed using monoclonal antibodies to the extracellular domain. These four alleles were used to screen for and analyze our panel of 18 EMS-generated Ptp69D alleles.
Isolation of new Ptp69D alleles: As described in materials and methods, male progeny of mutagenized males were crossed individually to Ptp69D1, Ptp69D4, or Df(3L)8ex34 virgin females. Candidates that failed to complement these three deletions could bear mutations in Ptp69D, KLC, snRNP69D, mirr, or other essential genes uncovered by Df(3L)8ex34. To distinguish among these possibilities, all candidates were crossed to the other three deletion alleles. Using these tests, six alleles of KLC, two alleles of mirr, and 11 other lethal mutations defining six complementation groups uncovered by Df(3L)8ex34 (data not shown) were isolated in addition to 18 potential Ptp69D alleles. All 18 of these candidates map to the genetic interval between hairy and thread, phenotypic markers that flank the Ptp69D locus. A Ptp-69D transgene was able to rescue animals homozygous for 17 of the candidates, confirming their identity as Ptp69D alleles. The remaining allele, Ptp69D16, bears a closely linked unselected lethal mutation (see Temperature-sensitive Ptp69D alleles).
Strength of Ptp69D deletion alleles: To examine phenotypic differences among the original deletion alleles, flies heterozygous for each of the 18 EMS alleles were crossed with flies heterozygous for each of the four deletion alleles at 25° and 18° and the fate of the cross progeny was examined. As can be seen in Tables 1 and 2, the progeny of crosses with Ptp69D4 and Df(3L)8ex25 generally have a greater probability of developing to the pupal and adult stages than do offspring of crosses with Df(3L)8ex34 or Ptp69D1. Summing the data from all 25° crosses reveals that Ptp69D4 and Df(3L)8ex25 trans-heterozygous animals are twice as likely to develop into healthy adults and 1.5 times as likely to eclose as are Df(3L)8ex34 and Ptp69D1 trans-heterozygotes (data not shown). Df(3L)8ex34 and Ptp69D1 display little or no difference in viability in trans to any of the new alleles. We conclude that the four deletion alleles vary in strength, with Df(3L)8ex34 = Ptp69D1 > Df(3L)8ex25 > Ptp69D4 in order of decreasing strength.
Strength of Ptp69D EMS alleles: To determine the strength of the new alleles and to identify temperature-sensitive alleles, we examined the fate of animals homozygous for each allele at 18° and 25° (Table 3). One striking result of these experiments is that Ptp69D7, Ptp-69D11, and Ptp69D16 homozygotes rarely, if ever, form pupae. The observation that animals hemizygous for Ptp69D7 or Ptp69D11 can form pupae (Tables 1 and 2),
Homozygous viability
| . | 25° . | 18° . | ||||
|---|---|---|---|---|---|---|
| Allele . | A . | E . | P . | A . | E . | P . |
| Ptp69D7 | 0 | 0 | 0 | 0 | 0 | 0 |
| Ptp69D16 | 0 | 0 | 0 | ND | ND | 0 |
| Ptp69D11 | 0 | 0 | 3 | 0 | 0 | 11 |
| Ptp69D15 | 0 | 1 | 4 | 6 | 7 | 11 |
| Ptp69D8 | 0 | 2 | 8 | 0 | 2 | 5 |
| Ptp69D14 | 1 | 3 | 8 | 0 | 0 | 12 |
| Ptp69D21 | 1 | 3 | 11 | 1 | 7 | 35 |
| Ptp69D17 | 1 | 3 | 16 | 0 | 20 | 39 |
| Ptp69D18 | 0 | 0 | 23 | 82 | 85 | 107 |
| Ptp69D6 | 0 | 0 | 23 | 0 | 0 | 17 |
| Ptp69D13 | 1 | 8 | 19 | 1 | 53 | 80 |
| Ptp69D22 | 0 | 2 | 29 | 8 | 16 | 102 |
| Ptp69D12 | 0 | 1 | 34 | 59 | 87 | 90 |
| Ptp69D19 | 3 | 12 | 27 | 6 | 27 | 59 |
| Ptp69D9 | 3 | 19 | 35 | 0 | 9 | 28 |
| Ptp69D5 | 44 | 62 | 67 | 36 | 43 | 69 |
| Ptp69D10 | 77 | 80 | 83 | 85 | 93 | 88 |
| Ptp69D20 | 96 | 98 | 99 | 101 | 102 | 96 |
| Ptp69Dy98 | ND | ND | ND | ND | ND | ND |
| Ptp69Dy100 | 0 | 5 | 26 | ND | ND | ND |
| Ptp69Dy125 | ND | ND | ND | ND | ND | ND |
| . | 25° . | 18° . | ||||
|---|---|---|---|---|---|---|
| Allele . | A . | E . | P . | A . | E . | P . |
| Ptp69D7 | 0 | 0 | 0 | 0 | 0 | 0 |
| Ptp69D16 | 0 | 0 | 0 | ND | ND | 0 |
| Ptp69D11 | 0 | 0 | 3 | 0 | 0 | 11 |
| Ptp69D15 | 0 | 1 | 4 | 6 | 7 | 11 |
| Ptp69D8 | 0 | 2 | 8 | 0 | 2 | 5 |
| Ptp69D14 | 1 | 3 | 8 | 0 | 0 | 12 |
| Ptp69D21 | 1 | 3 | 11 | 1 | 7 | 35 |
| Ptp69D17 | 1 | 3 | 16 | 0 | 20 | 39 |
| Ptp69D18 | 0 | 0 | 23 | 82 | 85 | 107 |
| Ptp69D6 | 0 | 0 | 23 | 0 | 0 | 17 |
| Ptp69D13 | 1 | 8 | 19 | 1 | 53 | 80 |
| Ptp69D22 | 0 | 2 | 29 | 8 | 16 | 102 |
| Ptp69D12 | 0 | 1 | 34 | 59 | 87 | 90 |
| Ptp69D19 | 3 | 12 | 27 | 6 | 27 | 59 |
| Ptp69D9 | 3 | 19 | 35 | 0 | 9 | 28 |
| Ptp69D5 | 44 | 62 | 67 | 36 | 43 | 69 |
| Ptp69D10 | 77 | 80 | 83 | 85 | 93 | 88 |
| Ptp69D20 | 96 | 98 | 99 | 101 | 102 | 96 |
| Ptp69Dy98 | ND | ND | ND | ND | ND | ND |
| Ptp69Dy100 | 0 | 5 | 26 | ND | ND | ND |
| Ptp69Dy125 | ND | ND | ND | ND | ND | ND |
The adult survival (A), eclosion (E), and pupation (P) rates (in percent) of homozygous progeny from self-crosses of Ptp69D alleles (left column) are shown. Alleles are ranked from the strongest to weakest at 25°, starting at the top. More than 200 progeny were counted for each cross. ND, not done.
Homozygous viability
| . | 25° . | 18° . | ||||
|---|---|---|---|---|---|---|
| Allele . | A . | E . | P . | A . | E . | P . |
| Ptp69D7 | 0 | 0 | 0 | 0 | 0 | 0 |
| Ptp69D16 | 0 | 0 | 0 | ND | ND | 0 |
| Ptp69D11 | 0 | 0 | 3 | 0 | 0 | 11 |
| Ptp69D15 | 0 | 1 | 4 | 6 | 7 | 11 |
| Ptp69D8 | 0 | 2 | 8 | 0 | 2 | 5 |
| Ptp69D14 | 1 | 3 | 8 | 0 | 0 | 12 |
| Ptp69D21 | 1 | 3 | 11 | 1 | 7 | 35 |
| Ptp69D17 | 1 | 3 | 16 | 0 | 20 | 39 |
| Ptp69D18 | 0 | 0 | 23 | 82 | 85 | 107 |
| Ptp69D6 | 0 | 0 | 23 | 0 | 0 | 17 |
| Ptp69D13 | 1 | 8 | 19 | 1 | 53 | 80 |
| Ptp69D22 | 0 | 2 | 29 | 8 | 16 | 102 |
| Ptp69D12 | 0 | 1 | 34 | 59 | 87 | 90 |
| Ptp69D19 | 3 | 12 | 27 | 6 | 27 | 59 |
| Ptp69D9 | 3 | 19 | 35 | 0 | 9 | 28 |
| Ptp69D5 | 44 | 62 | 67 | 36 | 43 | 69 |
| Ptp69D10 | 77 | 80 | 83 | 85 | 93 | 88 |
| Ptp69D20 | 96 | 98 | 99 | 101 | 102 | 96 |
| Ptp69Dy98 | ND | ND | ND | ND | ND | ND |
| Ptp69Dy100 | 0 | 5 | 26 | ND | ND | ND |
| Ptp69Dy125 | ND | ND | ND | ND | ND | ND |
| . | 25° . | 18° . | ||||
|---|---|---|---|---|---|---|
| Allele . | A . | E . | P . | A . | E . | P . |
| Ptp69D7 | 0 | 0 | 0 | 0 | 0 | 0 |
| Ptp69D16 | 0 | 0 | 0 | ND | ND | 0 |
| Ptp69D11 | 0 | 0 | 3 | 0 | 0 | 11 |
| Ptp69D15 | 0 | 1 | 4 | 6 | 7 | 11 |
| Ptp69D8 | 0 | 2 | 8 | 0 | 2 | 5 |
| Ptp69D14 | 1 | 3 | 8 | 0 | 0 | 12 |
| Ptp69D21 | 1 | 3 | 11 | 1 | 7 | 35 |
| Ptp69D17 | 1 | 3 | 16 | 0 | 20 | 39 |
| Ptp69D18 | 0 | 0 | 23 | 82 | 85 | 107 |
| Ptp69D6 | 0 | 0 | 23 | 0 | 0 | 17 |
| Ptp69D13 | 1 | 8 | 19 | 1 | 53 | 80 |
| Ptp69D22 | 0 | 2 | 29 | 8 | 16 | 102 |
| Ptp69D12 | 0 | 1 | 34 | 59 | 87 | 90 |
| Ptp69D19 | 3 | 12 | 27 | 6 | 27 | 59 |
| Ptp69D9 | 3 | 19 | 35 | 0 | 9 | 28 |
| Ptp69D5 | 44 | 62 | 67 | 36 | 43 | 69 |
| Ptp69D10 | 77 | 80 | 83 | 85 | 93 | 88 |
| Ptp69D20 | 96 | 98 | 99 | 101 | 102 | 96 |
| Ptp69Dy98 | ND | ND | ND | ND | ND | ND |
| Ptp69Dy100 | 0 | 5 | 26 | ND | ND | ND |
| Ptp69Dy125 | ND | ND | ND | ND | ND | ND |
The adult survival (A), eclosion (E), and pupation (P) rates (in percent) of homozygous progeny from self-crosses of Ptp69D alleles (left column) are shown. Alleles are ranked from the strongest to weakest at 25°, starting at the top. More than 200 progeny were counted for each cross. ND, not done.
albeit infrequently, indicates that their viability phenotype is significantly worse than the null phenotype and suggests that these may be neomorphic alleles. Alternatively, it is possible that Ptp69D7 and Ptp69D11 harbor closely linked unselected mutations that decrease the viability of homozygotes. Both alleles have been backcrossed and can be rescued by a Ptp69D genomic transgene, however, supporting the conclusion that Ptp69D7 and Ptp69D11 are unusually strong alleles. Ptp69D16 is discussed below (see Temperature-sensitive Ptp69D alleles). At the other end of the spectrum, Ptp69D10 and Ptp69D20 are so weak that homozygotes can survive and propagate at either 18° or 25°, and Ptp69D5 is a weak, semiviable allele. Thus these 18 new mutations define an allelic series ranging in strength from subliminal to worse than null.
Ptp69D null phenotype: To identify when animals mutant for Ptp69D die, we balanced the new alleles and deletion alleles with TM6B, which confers a dominant Tubby pupal phenotype, allowing the easy identification of trans-heterozygous (non-Tubby) pupae and subsequent evaluation of their fate. On one hand, in crosses of Ptp69D10 or Ptp69D20 with any of the deficiencies >50% of trans-heterozygous progeny survive to adulthood at 25° (Table 1). By contrast, <20% of animals hemizygous or homozygous for the strongest of the new alleles form pupae, and ≤5% eclose. These few adults eclose 1–3 days after their heterozygous siblings, are feeble, and become trapped in the food like ancient mammals in the La Brea tar pits (tar pit phenotype). These observations suggest that alleles conferring these phenotypes, including Ptp69D1, Ptp69D7, Ptp69D8, Ptp69D11, Ptp69D16, and Ptp69D21, as well as Ptp69DY98 and Ptp69DY100, described in Newsome et al. (2000), represent complete loss of Ptp69D function at 25°. Note that Ptp69D1 and Ptp69DY100 are also presumed to be null alleles by molecular criteria (see Figure 1). In all cases, crosses of new EMS alleles with either strong deficiency produce fewer-than-expected trans-heterozygous pupae. Moreover, in all cases, a lower proportion of trans-heterozygous pupae hatch than do balanced siblings. Taken together, these observations indicate that most animals lacking DPTP69D die as larvae, a minority survive to the pupal stage, and rare individuals develop into feeble adults.
18° null alleles: On the basis of the 18° phenotypes of the molecular null alleles Ptp69D1 and Ptp69DY100, the Ptp69D null phenotype is temperature independent (Table 2). Six of the eight alleles identified as putative null alleles at 25°, Ptp69D1, Ptp69D7, Ptp69D8, Ptp69D11, Ptp69D16, and Ptp69DY100, also appear to be null at 18° (Tables 2 and 3).
Temperature-sensitive Ptp69D alleles: The viability phenotype of Ptp69D10, Ptp69D12, and Ptp69D18 is especially temperature sensitive. Although Ptp69D10 adults survive at both temperatures, Ptp69D12 and Ptp69D18 adults are viable at 18°, but do not eclose or suffer the tar pit phenotype at 25°. Ptp69D17, Ptp69D19, Ptp69D21, and Ptp69D22 are also temperature sensitive; twice as many animals homozygous for these alleles survive to the pupal stage at 18° than at 25° (Table 3). These results indicate that Ptp69D12 and Ptp69D18 are ideal alleles for temperature-shift experiments as adults bearing these alleles are quite healthy at 18° but die at 25° (see below).
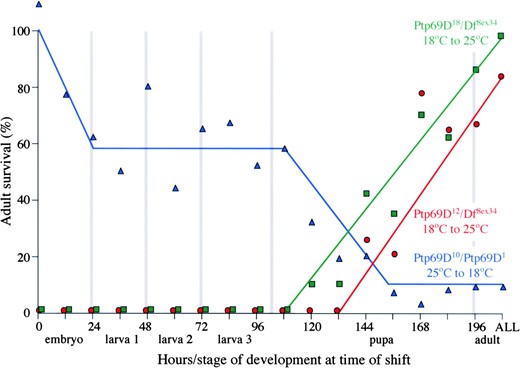
—Temperature dependence of adult health. The adult fates of Ptp69D10/Ptp69D1 animals raised initially at 25° and then shifted to 18° and Ptp69D12/Df(3L)8ex34 and Ptp69D18/Df(3L)8ex34 animals grown initially at 18° and then shifted to 25° are shown in blue, red, and green, respectively. The developmental stage and hours of development (25° equivalent) at the time of temperature shift are indicated along the x-axis. The rate of adult survival is indicated on the y-axis.
One allele, Ptp69D16, gives anomalous complementation results. In trans to Ptp69D1 and Ptp69D4, Ptp69D16 behaves like a temperature-sensitive allele, with 25° pupation rates of 13 and 27%, respectively, and 18° pupation rates of 28 and 52%, respectively. In contrast, Ptp69D16 behaves like an extremely strong allele in trans to Df(3L)8ex34 and Df(3L)8ex25, having a pupation rate of <1% at all temperatures. Similarly, Ptp69D16 homozygotes die before the pupal stage. One interpretation of these results is that Ptp69D16, Df(3L)8ex34, and Df(3L)8ex25 share a common second-site lethal mutation, presumably in KLC, CG32112, or CG10984. This interpretation is supported by the observation that a Ptp69D transgene rescues Ptp69D16/Ptp69D1 but not Ptp69D16/Df(3L)8ex34 (data not shown).
Adult health requires DPTP69D until the pupal stage: To determine the developmental requirement for DPTP69D function, the fate of animals bearing temperature-sensitive alleles raised at permissive or nonpermissive temperatures was examined. Figure 2 shows the effect of temperature on the health of adults. When raised entirely at 25°, <20% of Ptp69D12 and Ptp69D18 hemizygotes eclose, and those that do are feeble, fall into the food, and die. By contrast, >75% of Ptp69D12 and Ptp69D18 hemizygotes survive to become healthy, free-walking adults when raised at 18°. Ptp69D10/Ptp69D1 animals are similarly temperature sensitive with 20% developing into healthy adults at 25° as compared with 100% at 18°. Temperature-shift experiments involving any of these genotypes demonstrate a pupal requirement for DPTP69D function. Specifically, to survive after eclosion, Ptp69D12 and Ptp69D18 hemizygotes had to be mid-stage pupae prior to the 25° shift, and Ptp69D10/ Ptp69D1 animals had to be shifted to 18° as early pupae. Even late-stage pupae that are hemizygous for Ptp69D12 or Ptp69D18 suffer from shifting, indicating an ongoing requirement for DPTP69D function throughout the pupal stage.
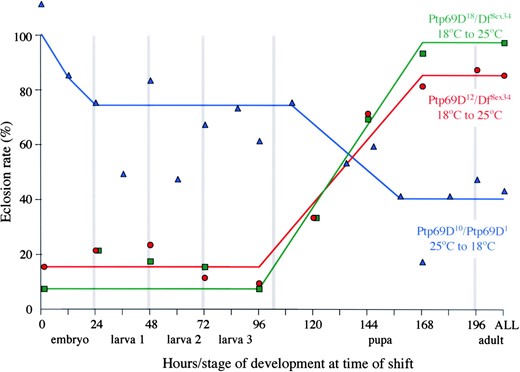
—Temperature dependence of eclosion. The eclosion rates for Ptp69D10/Ptp69D1 animals raised initially at 25° and then shifted to 18° and Ptp69D12/Df(3L)8ex34 and Ptp69D18/Df(3L)8ex34 animals grown initially at 18° and then shifted to 25° are shown in blue, red, and green, respectively. The developmental stage and hours of development (25° equivalent) at the time of temperature shift are indicated along the x-axis. The eclosion rate of mutants is indicated on the y-axis.
Eclosion requires DPTP69D during embryogenesis: Nonpermissive-to-permissive shifts involving Ptp69D10/ Ptp69D1 animals also reveal an embryonic requirement for DPTP69D function, as animals reared for as little as 12 hr at 25° are less likely to survive as adults than animals reared entirely at 18°. The pupal and embryonic requirements for DPTP69D function are also reflected in eclosion rates (Figure 3). The rate of eclosion drops from 100% at 18° to 75% when Ptp69D10/Ptp69D1 animals are reared at 25° for their first 24 hr. Eclosion drops further to 40% if Ptp69D10/Ptp69D1 animals are raised until the early pupal stage at the high temperature. Similarly, the eclosion rate of Ptp69D12 and Ptp69D18 hemizygotes drops from >80% when raised at 18° to <20% when shifted to the nonpermissive temperature prior to the onset of pupation. Unlike adult health, the eclosion rates of Ptp69D12 and Ptp69D18 hemizygotes are unaffected by late pupal shifts to the nonpermissive temperature. Thus we conclude that adult health requires DPTP69D function throughout the pupal stage whereas eclosion requires DPTP69D function during embryonic and early to mid-pupal stages.
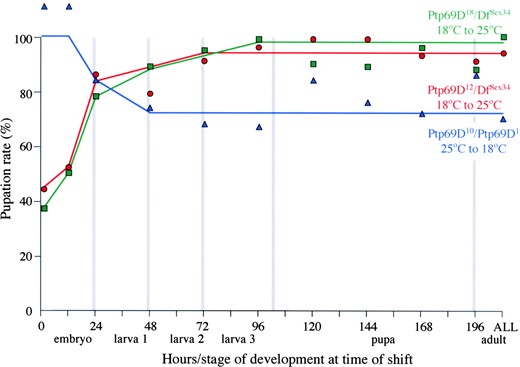
Larval survival requires DPTP69D during late embryogenesis: In addition to its requirement for eclosion, the underrepresentation of hemizygous pupae among the progeny of flies bearing Ptp69D12 or Ptp69D18 crossed to Df(3L)8ex34 indicates a function for DPTP69D prior to the pupal stage. To determine the developmental requirement for DPTP69D function for larval survival, Ptp69D10/Ptp69D1, Ptp69D12/Df(3L)8ex34, and Ptp69D18/Df (3L)8ex34 pupal counts were correlated with temperature as shown in Figure 4. As can be seen, 100% of Ptp69D10/Ptp69D1 animals develop into pupae when maintained entirely at the permissive temperature. A similar result was obtained for animals shifted to the permissive temperature after 12 hr. Strikingly, development at the nonpermissive temperature for as little as 24 hr caused a 25% decrease in the expected number of pupae. Complementary temperature-shift experiments confirm this embryonic period of sensitivity, as survival to the pupal stage increases from 40 to 80% when Ptp69D12 and Ptp-69D18 hemizygotes are shifted to the nonpermissive temperature after embryogenesis. These results reveal that larval survival requires DPTP69D function during late embryonic development.
Protein expression in Ptp69D mutants: To examine the effects of the EMS alleles on DPTP69D expression, mutant embryos were examined with anti-DPTP69D monoclonal antibodies. These experiments revealed that the alleles fell into three categories of DPTP69D expression. While Ptp69D7, Ptp69D10, Ptp69D20, and Ptp-69D21 embryos express wild-type levels of DPTP69D, DPT-P69D levels are reduced in Ptp69D5, Ptp69D13, Ptp69D17, and Ptp69D19 embryos and undetectable in embryos homozygous for the remaining nine alleles (Table 4). In no case did we detect changes in DPTP69D localization.
—Temperature dependence of larval survival. The pupation rates of Ptp69D10/Ptp69D1 animals raised initially at 25° and then shifted to 18° and Ptp69D12/Df(3L)8ex34 and Ptp69D18/Df(3L)8ex34 animals grown initially at 18° and then shifted to 25° are shown in blue, red, and green, respectively. The developmental stage and hours of development (25° equivalent) at the time of temperature shift are indicated along the x-axis. The percentage of mutants that formed pupae is indicated on the y-axis.
Ptp69D alleles cause motor axon defects: The role of DPTP69D in axon guidance has been inferred from the examination of the neuromuscular system of Ptp69D1/ Df(3L)8ex25 trans-heterozygous embryos (Desai et al. 1996). Although Ptp69D1 appears to represent a null allele, our results suggest that Df(3L)8ex25 may not completely abrogate DPTP69D function, as Ptp69D1/Df(3L)8ex25 animals develop to the pupal and adult stages more frequently than do Ptp69D1/Df(3L)8ex34 animals (Tables 1 and 2). Indeed, several of the new alleles confer stronger motor axon phenotypes. For example, embryos homozygous for Ptp69D7, Ptp69D10, Ptp69D21, and Ptp69D22 display motor axon defects that are both more severe and of higher penetrance than those of Ptp69D1/Df(3L)8ex25 embryos
Protein expression by mutant alleles
| . | Staining by monoclonal . | ||
|---|---|---|---|
| Allele . | 3F11 . | 3F12 . | 5A6 . |
| Ptp69D5 | Reduced | None | None |
| Ptp69D6 | None | None | None |
| Ptp69D7 | Wild type | ||
| Ptp69D8 | None | None | None |
| Ptp69D9 | None | ||
| Ptp69D10 | Wild type | ||
| Ptp69D11 | None | None | None |
| Ptp69D12 | Nonea | None | |
| Ptp69D13 | Reduced | None | None |
| Ptp69D14 | None | ||
| Ptp69D15 | None | None | |
| Ptp69D16 | None | ||
| Ptp69D17 | Reduced | None | |
| Ptp69D18 | Nonea | ||
| Ptp69D19 | Reduced | None | |
| Ptp69D20 | Wild type | ||
| Ptp69D21 | Wild type | Wild type | |
| Ptp69D22 | None | None | None |
| Ptp69Dy98 | None | ||
| Ptp69Dy100 | None | ||
| Reduced | |||
| Ptp69Dy125 | |||
| . | Staining by monoclonal . | ||
|---|---|---|---|
| Allele . | 3F11 . | 3F12 . | 5A6 . |
| Ptp69D5 | Reduced | None | None |
| Ptp69D6 | None | None | None |
| Ptp69D7 | Wild type | ||
| Ptp69D8 | None | None | None |
| Ptp69D9 | None | ||
| Ptp69D10 | Wild type | ||
| Ptp69D11 | None | None | None |
| Ptp69D12 | Nonea | None | |
| Ptp69D13 | Reduced | None | None |
| Ptp69D14 | None | ||
| Ptp69D15 | None | None | |
| Ptp69D16 | None | ||
| Ptp69D17 | Reduced | None | |
| Ptp69D18 | Nonea | ||
| Ptp69D19 | Reduced | None | |
| Ptp69D20 | Wild type | ||
| Ptp69D21 | Wild type | Wild type | |
| Ptp69D22 | None | None | None |
| Ptp69Dy98 | None | ||
| Ptp69Dy100 | None | ||
| Reduced | |||
| Ptp69Dy125 | |||
α-DPTP69D monoclonal antibody staining of embryos from balanced stocks of Ptp69D alleles is shown. If one-quarter of the embryos failed to stain, then staining was listed as “none.” If all embryos stained equally, then staining was listed as “wild type.” If one-quarter of the embryos displayed reduced staining relative to their siblings, the staining was noted as “reduced.”
No staining at 25° but wild-type staining at 18°.
Protein expression by mutant alleles
| . | Staining by monoclonal . | ||
|---|---|---|---|
| Allele . | 3F11 . | 3F12 . | 5A6 . |
| Ptp69D5 | Reduced | None | None |
| Ptp69D6 | None | None | None |
| Ptp69D7 | Wild type | ||
| Ptp69D8 | None | None | None |
| Ptp69D9 | None | ||
| Ptp69D10 | Wild type | ||
| Ptp69D11 | None | None | None |
| Ptp69D12 | Nonea | None | |
| Ptp69D13 | Reduced | None | None |
| Ptp69D14 | None | ||
| Ptp69D15 | None | None | |
| Ptp69D16 | None | ||
| Ptp69D17 | Reduced | None | |
| Ptp69D18 | Nonea | ||
| Ptp69D19 | Reduced | None | |
| Ptp69D20 | Wild type | ||
| Ptp69D21 | Wild type | Wild type | |
| Ptp69D22 | None | None | None |
| Ptp69Dy98 | None | ||
| Ptp69Dy100 | None | ||
| Reduced | |||
| Ptp69Dy125 | |||
| . | Staining by monoclonal . | ||
|---|---|---|---|
| Allele . | 3F11 . | 3F12 . | 5A6 . |
| Ptp69D5 | Reduced | None | None |
| Ptp69D6 | None | None | None |
| Ptp69D7 | Wild type | ||
| Ptp69D8 | None | None | None |
| Ptp69D9 | None | ||
| Ptp69D10 | Wild type | ||
| Ptp69D11 | None | None | None |
| Ptp69D12 | Nonea | None | |
| Ptp69D13 | Reduced | None | None |
| Ptp69D14 | None | ||
| Ptp69D15 | None | None | |
| Ptp69D16 | None | ||
| Ptp69D17 | Reduced | None | |
| Ptp69D18 | Nonea | ||
| Ptp69D19 | Reduced | None | |
| Ptp69D20 | Wild type | ||
| Ptp69D21 | Wild type | Wild type | |
| Ptp69D22 | None | None | None |
| Ptp69Dy98 | None | ||
| Ptp69Dy100 | None | ||
| Reduced | |||
| Ptp69Dy125 | |||
α-DPTP69D monoclonal antibody staining of embryos from balanced stocks of Ptp69D alleles is shown. If one-quarter of the embryos failed to stain, then staining was listed as “none.” If all embryos stained equally, then staining was listed as “wild type.” If one-quarter of the embryos displayed reduced staining relative to their siblings, the staining was noted as “reduced.”
No staining at 25° but wild-type staining at 18°.
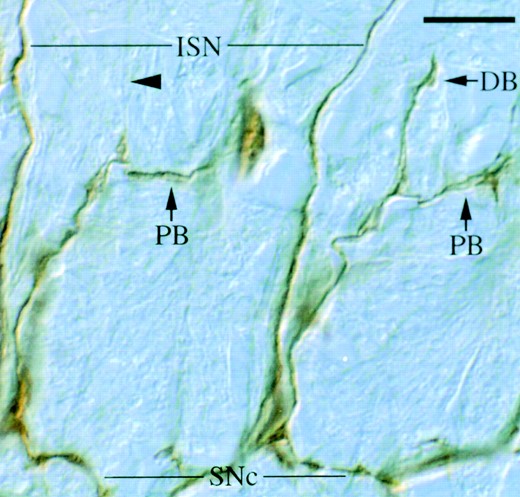
(Table 5). In all cases ISNb nerve outgrowth is most sensitive to the loss of DPTP69D. Embryos homozygous for strong alleles (excluding Ptp69D7; see DPTP69D and DLAR allow ISNb muscle innervation) display defects in 10–33% of ISNb nerves, including premature termination, ectopic or reduced defasciculation, and defects in target recognition (Figure 5). Other motor nerves display less penetrating defects in mutant embryos: 5–22% of SNas are defective, most lacking a dorsal or posterior branch; 3–14% of ISNs are defective, often defasciculating abnormally; and 0–12% of SNcs are defective. See Figures 5,6,7,8 for representative defects. These observations indicate that DPTP69D functions to facilitate motor axon guidance.
Ptp69D motor axon defects are partially penetrant: The relatively normal appearance of most nerves in embryos presumed to lack zygotic DPTP69D indicates that ISNb axons are not completely dependent upon DPTP69D function. One caveat to this conclusion is that DPTP69D, presumably of maternal origin, can be detected in early embryos (Fitzpatrick et al. 1995). However, embryos lacking both maternal and zygotic sources of DPTP69D appear generally similar to those lacking only embryonic DPTP69D (C. Desai, unpublished observations). These results support the idea that molecules such as the other neural RPTPs partially compensate for the loss of DPTP69D.
DPTP69D and DLAR allow ISNb muscle innervation: Bypass defects arise when some or all ISNb axons fail to enter the target ventro-lateral muscle (VLM) field, but instead grow dorsally within or next to the ISN nerve (Figure 5). As can be seen in Table 6, these defects occur at a rate of ∼15% in Ptp69D null embryos and 30% in Dlar mutants (Krueger et al. 1996; Desai et al. 1997a). Strikingly, one of our new alleles, Ptp69D7, confers the bypass phenotype at rates approaching those seen in Dlar mutants (Table 6). Previous results demonstrated that the Dlar bypass phenotype requires a third RPTP, DPTP99A (Desai et al. 1997a). The ISNb bypass phenotype observed in Ptp69D embryos is likewise dependent upon DPTP99A (Table 6). Note that we removed only one copy of Ptp99A in these experiments because Ptp69D Ptp99A double-mutant embryos display severe ISNb defects that can preempt the bypass phenotype (Desai et al. 1996, 1997a). These results suggest that DPTP69D and DLAR function together to allow ISNb axons to innervate the VLM by counteracting DPTP99A at this choice point.
Molecular defects in Ptp69D alleles: To determine the molecular basis for reduced or absent function of the Ptp69D gene, selected alleles were sequenced. For the most part, alleles directing the expression of immunologically detectable DPTP69D were chosen for sequence analysis. The results are illustrated in Figure 1B and listed in Table 7. A shared molecular defect may be responsible for the similar phenotypes exhibited by two temperature-sensitive alleles, Ptp69D12 and Ptp69D18. Both alleles bear a missense mutation, glycine to aspartate, in the extracellular region of the protein, C-terminal to the cleavage site 47 residues from the start of the trans-membrane segment. Two of our EMS alleles, Ptp69D20 and Ptp69D21, as well as Ptp69DY125 (Newsome et al. 2000), have missense mutations in the active site of the D1 phosphatase domain. The mutation in Ptp69D21 changes the invariant catalytic cysteine residue to tyrosine (Figure 1B) and almost certainly abrogates the activity of this domain (Table 8). In Ptp69D20 and Ptp-69DY125, conserved active-site glycines are mutated to alanine and arginine, respectively (Figure 1B). It will be interesting to determine the effects of these changes on phosphatase activity and then to correlate activity with the ability to support axon guidance. The two alleles that cause the most severe axon guidance defects have strikingly different mutations. Ptp69D7 has a small three-amino-acid deletion in the D1 phosphatase domain that removes a conserved aspartate residue. In other phosphatases, this residue is required to complete catalysis; mutants in which this residue is changed to alanine bind but do not release their substrates. By
Motor axon defects
| . | ISNb . | ISN . | SNa . | SNc . | |||||
|---|---|---|---|---|---|---|---|---|---|
| Allele . | % all . | % sev . | N . | % all . | N . | % all . | N . | % all . | N . |
| Ptp69D1/Df(3L)8ex34 | 26 | 17 | 262 | 10 | 260 | 11 | 220 | 4 | 258 |
| Ptp69D4a | 18 | 6 | 238 | 6 | 238 | 9 | 200 | 5 | 239 |
| Ptp69D7 | 57 | 44 | 513 | 27 | 447 | 24 | 430 | 26 | 509 |
| Ptp69D10 | 33 | 28 | 420 | 14 | 399 | 22 | 350 | 4 | 415 |
| Ptp69D22 | 28 | 19 | 251 | 10 | 250 | 16 | 210 | 5 | 249 |
| Ptp69D8 | 30 | 16 | 227 | 9 | 235 | 14 | 197 | 12 | 208 |
| Pp69D15 | 36 | 15 | 129 | 6 | 127 | 13 | 109 | 6 | 125 |
| Ptp69D21 | 32 | 24 | 284 | 5 | 284 | 5 | 239 | 3 | 280 |
| Ptp69D11 | 32 | 13 | 233 | 7 | 236 | 8 | 199 | 1 | 229 |
| Ptp69D18 | 15 | 10 | 165 | 9 | 166 | 17 | 140 | 8 | 165 |
| Ptp69D12 | 18 | 15 | 223 | 9 | 230 | 7 | 200 | 6 | 217 |
| Ptp69D5 | 22 | 14 | 235 | 6 | 232 | 11 | 200 | 3 | 231 |
| Ptp69D19 | 18 | 11 | 153 | 7 | 154 | 10 | 130 | 3 | 152 |
| Ptp69D6 | 23 | 10 | 221 | 5 | 210 | 6 | 188 | 5 | 195 |
| Ptp69D14 | 20 | 10 | 234 | 3 | 238 | 7 | 197 | 5 | 220 |
| Ptp69D17 | 19 | 5 | 147 | 5 | 155 | 6 | 126 | 4 | 141 |
| Ptp69D9 | 15 | 8 | 160 | 6 | 158 | 6 | 132 | 0 | 158 |
| Ptp69D20 | 11 | 8 | 240 | 4 | 233 | 5 | 200 | 3 | 231 |
| Ptp69D13 | 10 | 5 | 229 | 3 | 237 | 5 | 200 | 1 | 222 |
| . | ISNb . | ISN . | SNa . | SNc . | |||||
|---|---|---|---|---|---|---|---|---|---|
| Allele . | % all . | % sev . | N . | % all . | N . | % all . | N . | % all . | N . |
| Ptp69D1/Df(3L)8ex34 | 26 | 17 | 262 | 10 | 260 | 11 | 220 | 4 | 258 |
| Ptp69D4a | 18 | 6 | 238 | 6 | 238 | 9 | 200 | 5 | 239 |
| Ptp69D7 | 57 | 44 | 513 | 27 | 447 | 24 | 430 | 26 | 509 |
| Ptp69D10 | 33 | 28 | 420 | 14 | 399 | 22 | 350 | 4 | 415 |
| Ptp69D22 | 28 | 19 | 251 | 10 | 250 | 16 | 210 | 5 | 249 |
| Ptp69D8 | 30 | 16 | 227 | 9 | 235 | 14 | 197 | 12 | 208 |
| Pp69D15 | 36 | 15 | 129 | 6 | 127 | 13 | 109 | 6 | 125 |
| Ptp69D21 | 32 | 24 | 284 | 5 | 284 | 5 | 239 | 3 | 280 |
| Ptp69D11 | 32 | 13 | 233 | 7 | 236 | 8 | 199 | 1 | 229 |
| Ptp69D18 | 15 | 10 | 165 | 9 | 166 | 17 | 140 | 8 | 165 |
| Ptp69D12 | 18 | 15 | 223 | 9 | 230 | 7 | 200 | 6 | 217 |
| Ptp69D5 | 22 | 14 | 235 | 6 | 232 | 11 | 200 | 3 | 231 |
| Ptp69D19 | 18 | 11 | 153 | 7 | 154 | 10 | 130 | 3 | 152 |
| Ptp69D6 | 23 | 10 | 221 | 5 | 210 | 6 | 188 | 5 | 195 |
| Ptp69D14 | 20 | 10 | 234 | 3 | 238 | 7 | 197 | 5 | 220 |
| Ptp69D17 | 19 | 5 | 147 | 5 | 155 | 6 | 126 | 4 | 141 |
| Ptp69D9 | 15 | 8 | 160 | 6 | 158 | 6 | 132 | 0 | 158 |
| Ptp69D20 | 11 | 8 | 240 | 4 | 233 | 5 | 200 | 3 | 231 |
| Ptp69D13 | 10 | 5 | 229 | 3 | 237 | 5 | 200 | 1 | 222 |
All, the percentages of SNb, ISN, SNa, and SNc defective in embryos homozygous for Ptp69D alleles; sev, the percentage of SNb nerves that displayed severe defects, including the bypass, partial bypass, premature termination proximal to muscle 13, aberrant defasciculation, and detour phenotypes; N, the total number of nerves counted.
Data are from w; Ptp69D4 P[snRNP-1.5]/TM6B × w; Ptp69D4.
Motor axon defects
| . | ISNb . | ISN . | SNa . | SNc . | |||||
|---|---|---|---|---|---|---|---|---|---|
| Allele . | % all . | % sev . | N . | % all . | N . | % all . | N . | % all . | N . |
| Ptp69D1/Df(3L)8ex34 | 26 | 17 | 262 | 10 | 260 | 11 | 220 | 4 | 258 |
| Ptp69D4a | 18 | 6 | 238 | 6 | 238 | 9 | 200 | 5 | 239 |
| Ptp69D7 | 57 | 44 | 513 | 27 | 447 | 24 | 430 | 26 | 509 |
| Ptp69D10 | 33 | 28 | 420 | 14 | 399 | 22 | 350 | 4 | 415 |
| Ptp69D22 | 28 | 19 | 251 | 10 | 250 | 16 | 210 | 5 | 249 |
| Ptp69D8 | 30 | 16 | 227 | 9 | 235 | 14 | 197 | 12 | 208 |
| Pp69D15 | 36 | 15 | 129 | 6 | 127 | 13 | 109 | 6 | 125 |
| Ptp69D21 | 32 | 24 | 284 | 5 | 284 | 5 | 239 | 3 | 280 |
| Ptp69D11 | 32 | 13 | 233 | 7 | 236 | 8 | 199 | 1 | 229 |
| Ptp69D18 | 15 | 10 | 165 | 9 | 166 | 17 | 140 | 8 | 165 |
| Ptp69D12 | 18 | 15 | 223 | 9 | 230 | 7 | 200 | 6 | 217 |
| Ptp69D5 | 22 | 14 | 235 | 6 | 232 | 11 | 200 | 3 | 231 |
| Ptp69D19 | 18 | 11 | 153 | 7 | 154 | 10 | 130 | 3 | 152 |
| Ptp69D6 | 23 | 10 | 221 | 5 | 210 | 6 | 188 | 5 | 195 |
| Ptp69D14 | 20 | 10 | 234 | 3 | 238 | 7 | 197 | 5 | 220 |
| Ptp69D17 | 19 | 5 | 147 | 5 | 155 | 6 | 126 | 4 | 141 |
| Ptp69D9 | 15 | 8 | 160 | 6 | 158 | 6 | 132 | 0 | 158 |
| Ptp69D20 | 11 | 8 | 240 | 4 | 233 | 5 | 200 | 3 | 231 |
| Ptp69D13 | 10 | 5 | 229 | 3 | 237 | 5 | 200 | 1 | 222 |
| . | ISNb . | ISN . | SNa . | SNc . | |||||
|---|---|---|---|---|---|---|---|---|---|
| Allele . | % all . | % sev . | N . | % all . | N . | % all . | N . | % all . | N . |
| Ptp69D1/Df(3L)8ex34 | 26 | 17 | 262 | 10 | 260 | 11 | 220 | 4 | 258 |
| Ptp69D4a | 18 | 6 | 238 | 6 | 238 | 9 | 200 | 5 | 239 |
| Ptp69D7 | 57 | 44 | 513 | 27 | 447 | 24 | 430 | 26 | 509 |
| Ptp69D10 | 33 | 28 | 420 | 14 | 399 | 22 | 350 | 4 | 415 |
| Ptp69D22 | 28 | 19 | 251 | 10 | 250 | 16 | 210 | 5 | 249 |
| Ptp69D8 | 30 | 16 | 227 | 9 | 235 | 14 | 197 | 12 | 208 |
| Pp69D15 | 36 | 15 | 129 | 6 | 127 | 13 | 109 | 6 | 125 |
| Ptp69D21 | 32 | 24 | 284 | 5 | 284 | 5 | 239 | 3 | 280 |
| Ptp69D11 | 32 | 13 | 233 | 7 | 236 | 8 | 199 | 1 | 229 |
| Ptp69D18 | 15 | 10 | 165 | 9 | 166 | 17 | 140 | 8 | 165 |
| Ptp69D12 | 18 | 15 | 223 | 9 | 230 | 7 | 200 | 6 | 217 |
| Ptp69D5 | 22 | 14 | 235 | 6 | 232 | 11 | 200 | 3 | 231 |
| Ptp69D19 | 18 | 11 | 153 | 7 | 154 | 10 | 130 | 3 | 152 |
| Ptp69D6 | 23 | 10 | 221 | 5 | 210 | 6 | 188 | 5 | 195 |
| Ptp69D14 | 20 | 10 | 234 | 3 | 238 | 7 | 197 | 5 | 220 |
| Ptp69D17 | 19 | 5 | 147 | 5 | 155 | 6 | 126 | 4 | 141 |
| Ptp69D9 | 15 | 8 | 160 | 6 | 158 | 6 | 132 | 0 | 158 |
| Ptp69D20 | 11 | 8 | 240 | 4 | 233 | 5 | 200 | 3 | 231 |
| Ptp69D13 | 10 | 5 | 229 | 3 | 237 | 5 | 200 | 1 | 222 |
All, the percentages of SNb, ISN, SNa, and SNc defective in embryos homozygous for Ptp69D alleles; sev, the percentage of SNb nerves that displayed severe defects, including the bypass, partial bypass, premature termination proximal to muscle 13, aberrant defasciculation, and detour phenotypes; N, the total number of nerves counted.
Data are from w; Ptp69D4 P[snRNP-1.5]/TM6B × w; Ptp69D4.
contrast, the change in Ptp69D10 converts a valine between the immunoglobulin domains to a glutamate. Finally, embryos homozygous for Ptp69D9 and Ptp69D17 express low or undetectable levels of DPTP69D. Both alleles have changes in the trans-membrane region, suggesting that the C-terminal cleavage product may stabilize the DPTP69D extracellular portion and/or anchor it to the cell surface.
DISCUSSION
The 18 new chemically induced mutations in Ptp69D form an allelic series. A matrix of crosses between these 18 alleles and 4 existing alleles derived from the imprecise excision of nearby P elements reveals differences not only among the new alleles, but also among the deletion alleles. Among the deletion alleles, Ptp69D4 appears weakest but Df(3L)8ex25 also appears to retain residual DPTP69D activity. Specifically, animals bearing the new alleles in trans to Ptp69D4 or Df(3L)8ex25 develop to later stages more frequently than do Ptp69D1 and Df(3L)8ex34 trans-heterozygotes. These results lead to two conclusions. First, levels of DPTP69D undetectable by immunohistohemistry are functionally relevant. Second, the Df(3L)8ex25 allele apparently directs the expression of an N-terminal fragment that lacks phosphatase domains but retains residual biological activity. These observations also indicate that previous descriptions of the Ptp69D mutant phenotype, based upon Ptp69D1/ Df(3L)8ex25 embryos, did not describe the effects of complete loss of zygotic DPTP69D expression.
—ISNb defects typical of Ptp69D mutants. Four adjacent ISNb nerves from a Ptp69D20 embryo are shown. The center ISNbs display wild-type morphology. The large arrows indicate clefts between muscles 12, 13, 7, and 6, sites at which the ISNb normally synapses. The right ISNb prematurely terminates at the proximal edge of muscle 13 and so lacks the distalmost synapse between muscles 13 and 12 (large arrowhead). The left ISNb displays a partial bypass defect in which axons that normally would innervate muscle 13 instead rejoin the ISN nerve, indicated by small arrows (out of focus in the other hemisegments). The two branches of the split ISNb are indicated by small arrowheads. Bar, ∼10 μm.
—SNa defects typical of Ptp69D mutants. Two adjacent SNa nerves from a Ptp69D19 embryo are shown. The right SNa bifurcates normally at the distal edge of muscle fiber 12 (out of focus), forming a dorsal branch (DB) and a posterior branch (PB). In older embryos, the dorsal branch bifurcates (Figure 8). The left SNa lacks the dorsal branch (arrowhead), but forms the posterior branch normally. The ISN and SNc nerves are indicated. Bar, ∼10 μm.
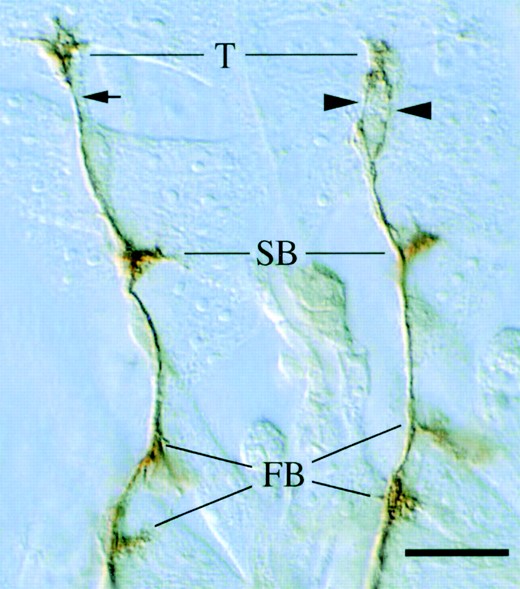
—ISN defects typical of Ptp69D mutants. Two adjacent ISN nerves from a Ptp69D12 embryo are shown. The left ISN displays normal morphology, defasciculating only to form synapses at the terminal arbor (T), second branch point (SB), and first branch points (FB). The right ISN synapses normally but also defasciculates between the second branch point and the terminal arbor (arrowheads; compare with small arrow). Bar, ∼10 μm.
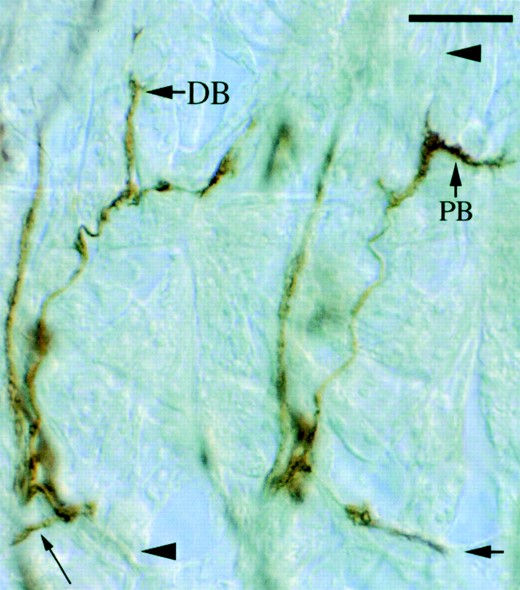
—SNc defects typical of Ptp69D mutants. Two adjacent SNc nerves from a Ptp69D22 embryo are shown. The right SNc displays normal morphology, growing posterio-medially (large arrow). The left SNa branches normally but then turns anterio-medially (small arrow). The bifurcation of the SNa dorsal branch is indicated (DB). Note that the right SNa fails to form a dorsal branch (arrowhead) and that the posterior branch (PB) is thick, suggesting the presence of ectopic dorsal branch axons. Bar, ∼10 μm.
Ptp69D7 may be a neomorphic allele: The severe axon and viability defects caused by Ptp69D7 could be due to closely linked unselected modifier mutations or to neomorphic activity of the mutant DPTP69D encoded by this allele. Because Ptp69D transgenes rescue the viability of Ptp69D7 and because Ptp69D7 directs the expression of wild-type levels of DPTP69D, we favor the second hypothesis. The molecular defect in the Ptp69D7 allele supports this hypothesis: lacking a conserved aspartate residue required for catalysis, Ptp69D7 might bind and sequester substrate proteins. Previous studies show that the cytoplasmic domains of DLAR and DPT-P69D can substitute for one another in retinal and motor axon guidance, implying that they can utilize the same substrates (Desai et al. 1997a; Maurel-Zaffran et al. 2001). Thus the defects observed in Ptp69D7 embryos could be due to loss of DPTP69D coupled with impaired signaling through DLAR and/or other neural RPTPs.
ISNb bypass defect requires DPTP99A
| Genotype . | % complete bypass . | % partial bypass . | N . |
|---|---|---|---|
| w; dlar5.5/dlar13.2a | 19 | 12 | 260 |
| w; dlar5.5/dlar13.2; Ptp99A1/Df(3R)R3a | 0 | 1 | 214 |
| w; Ptp69D1/Df(3L)8ex34 | 6 | 9 | 262 |
| w; Ptp69D1 Ptp99Ab/Df(3L)8ex34 + | 0 | 2 | 215 |
| w; Ptp69D7 | 25 | 15 | 453 |
| w; Ptp69D7 Ptp99A1/Ptp69D7 + | 2 | 5 | 314 |
| w; Ptp69D10 | 9 | 15 | 420 |
| w; Ptp69D10 Ptp99A1/Ptp69D10 + | 2 | 6 | 300 |
| w; Ptp69D21 | 1 | 16 | 284 |
| w; Ptp69D22 | 9 | 10 | 251 |
| Genotype . | % complete bypass . | % partial bypass . | N . |
|---|---|---|---|
| w; dlar5.5/dlar13.2a | 19 | 12 | 260 |
| w; dlar5.5/dlar13.2; Ptp99A1/Df(3R)R3a | 0 | 1 | 214 |
| w; Ptp69D1/Df(3L)8ex34 | 6 | 9 | 262 |
| w; Ptp69D1 Ptp99Ab/Df(3L)8ex34 + | 0 | 2 | 215 |
| w; Ptp69D7 | 25 | 15 | 453 |
| w; Ptp69D7 Ptp99A1/Ptp69D7 + | 2 | 5 | 314 |
| w; Ptp69D10 | 9 | 15 | 420 |
| w; Ptp69D10 Ptp99A1/Ptp69D10 + | 2 | 6 | 300 |
| w; Ptp69D21 | 1 | 16 | 284 |
| w; Ptp69D22 | 9 | 10 | 251 |
The penetrance of the ISNb bypass defect for various genotypes is shown. Complete bypass, all ISNb axons extend beyond the VLM; partial bypass, some ISNb axons enter the VLM while some extend beyond it; N, number of hemisegments scored.
Reported in Desai et al. (1997a).
Data are from embryos heterozygous for Ptp99A1 or Df(3R)R3.
ISNb bypass defect requires DPTP99A
| Genotype . | % complete bypass . | % partial bypass . | N . |
|---|---|---|---|
| w; dlar5.5/dlar13.2a | 19 | 12 | 260 |
| w; dlar5.5/dlar13.2; Ptp99A1/Df(3R)R3a | 0 | 1 | 214 |
| w; Ptp69D1/Df(3L)8ex34 | 6 | 9 | 262 |
| w; Ptp69D1 Ptp99Ab/Df(3L)8ex34 + | 0 | 2 | 215 |
| w; Ptp69D7 | 25 | 15 | 453 |
| w; Ptp69D7 Ptp99A1/Ptp69D7 + | 2 | 5 | 314 |
| w; Ptp69D10 | 9 | 15 | 420 |
| w; Ptp69D10 Ptp99A1/Ptp69D10 + | 2 | 6 | 300 |
| w; Ptp69D21 | 1 | 16 | 284 |
| w; Ptp69D22 | 9 | 10 | 251 |
| Genotype . | % complete bypass . | % partial bypass . | N . |
|---|---|---|---|
| w; dlar5.5/dlar13.2a | 19 | 12 | 260 |
| w; dlar5.5/dlar13.2; Ptp99A1/Df(3R)R3a | 0 | 1 | 214 |
| w; Ptp69D1/Df(3L)8ex34 | 6 | 9 | 262 |
| w; Ptp69D1 Ptp99Ab/Df(3L)8ex34 + | 0 | 2 | 215 |
| w; Ptp69D7 | 25 | 15 | 453 |
| w; Ptp69D7 Ptp99A1/Ptp69D7 + | 2 | 5 | 314 |
| w; Ptp69D10 | 9 | 15 | 420 |
| w; Ptp69D10 Ptp99A1/Ptp69D10 + | 2 | 6 | 300 |
| w; Ptp69D21 | 1 | 16 | 284 |
| w; Ptp69D22 | 9 | 10 | 251 |
The penetrance of the ISNb bypass defect for various genotypes is shown. Complete bypass, all ISNb axons extend beyond the VLM; partial bypass, some ISNb axons enter the VLM while some extend beyond it; N, number of hemisegments scored.
Reported in Desai et al. (1997a).
Data are from embryos heterozygous for Ptp99A1 or Df(3R)R3.
The phenotypic and genetic similarities of the ISNb bypass defect in Dlar and Ptp69D7 mutants support this conclusion.
Temperature-sensitive alleles: Ptp69D12 and Ptp69D18 confer very similar phenotypes at all temperatures, consistent with their shared molecular defect. At 25°, <5% of Ptp69D12 or Ptp69D18 homozygotes survive to the adult stage, all of which fall into the food and die. By contrast, at 18° most Ptp69D12 and Ptp69D18 homozygotes develop into healthy flies. The high frequency of adult survival at this permissive temperature indicates that DPTP69D12 and DPTP69D18 attain close to wild-type activity at 18°.
The third strongly temperature-sensitive allele, Ptp69D10,
Molecular changes in selected alleles
| Allele . | Base . | Coding change . | Domain . |
|---|---|---|---|
| Ptp69D7 | Δ 9 bp | ΔD1065 F1066 M1067 | D1 |
| Ptp69D9 | G-A | G812 to D | Tm |
| Ptp69D10 | T-A | V134 to D | Ig1/Ig2 |
| Ptp69D12 | T-A | W171 to R | Ig2 |
| G-A | G757 to E | Fn/Tm | |
| G-T | R903 to L | D1 | |
| Ptp69D17 | W821 to stop | Tm | |
| Ptp69D18 | G-A | G757 to E | Fn/Tm |
| Ptp69D20 | G-A | G1102 to S | D1 active site |
| Ptp69D21 | G-A | C1097 to Y | D1 active site |
| Ptp69DY98 | C-T1 | Q931 to stopa | D1a |
| Ptp69DY100 | G-A1 | W54 to stopa | Ig1a |
| Ptp69DY125 | G-A1 | G1105 to Ra | D1 active sitea |
| Allele . | Base . | Coding change . | Domain . |
|---|---|---|---|
| Ptp69D7 | Δ 9 bp | ΔD1065 F1066 M1067 | D1 |
| Ptp69D9 | G-A | G812 to D | Tm |
| Ptp69D10 | T-A | V134 to D | Ig1/Ig2 |
| Ptp69D12 | T-A | W171 to R | Ig2 |
| G-A | G757 to E | Fn/Tm | |
| G-T | R903 to L | D1 | |
| Ptp69D17 | W821 to stop | Tm | |
| Ptp69D18 | G-A | G757 to E | Fn/Tm |
| Ptp69D20 | G-A | G1102 to S | D1 active site |
| Ptp69D21 | G-A | C1097 to Y | D1 active site |
| Ptp69DY98 | C-T1 | Q931 to stopa | D1a |
| Ptp69DY100 | G-A1 | W54 to stopa | Ig1a |
| Ptp69DY125 | G-A1 | G1105 to Ra | D1 active sitea |
The molecular changes in DNA sequence are listed for each allele in the second column, and the predicted change in amino acid sequence is noted in the third column. The protein domain in which the predicted change appears is listed in the fourth column.
Data for Ptp69DY98, Ptp69DY100, and Ptp69DY125 were reported in Newsome et al. (2000).
Molecular changes in selected alleles
| Allele . | Base . | Coding change . | Domain . |
|---|---|---|---|
| Ptp69D7 | Δ 9 bp | ΔD1065 F1066 M1067 | D1 |
| Ptp69D9 | G-A | G812 to D | Tm |
| Ptp69D10 | T-A | V134 to D | Ig1/Ig2 |
| Ptp69D12 | T-A | W171 to R | Ig2 |
| G-A | G757 to E | Fn/Tm | |
| G-T | R903 to L | D1 | |
| Ptp69D17 | W821 to stop | Tm | |
| Ptp69D18 | G-A | G757 to E | Fn/Tm |
| Ptp69D20 | G-A | G1102 to S | D1 active site |
| Ptp69D21 | G-A | C1097 to Y | D1 active site |
| Ptp69DY98 | C-T1 | Q931 to stopa | D1a |
| Ptp69DY100 | G-A1 | W54 to stopa | Ig1a |
| Ptp69DY125 | G-A1 | G1105 to Ra | D1 active sitea |
| Allele . | Base . | Coding change . | Domain . |
|---|---|---|---|
| Ptp69D7 | Δ 9 bp | ΔD1065 F1066 M1067 | D1 |
| Ptp69D9 | G-A | G812 to D | Tm |
| Ptp69D10 | T-A | V134 to D | Ig1/Ig2 |
| Ptp69D12 | T-A | W171 to R | Ig2 |
| G-A | G757 to E | Fn/Tm | |
| G-T | R903 to L | D1 | |
| Ptp69D17 | W821 to stop | Tm | |
| Ptp69D18 | G-A | G757 to E | Fn/Tm |
| Ptp69D20 | G-A | G1102 to S | D1 active site |
| Ptp69D21 | G-A | C1097 to Y | D1 active site |
| Ptp69DY98 | C-T1 | Q931 to stopa | D1a |
| Ptp69DY100 | G-A1 | W54 to stopa | Ig1a |
| Ptp69DY125 | G-A1 | G1105 to Ra | D1 active sitea |
The molecular changes in DNA sequence are listed for each allele in the second column, and the predicted change in amino acid sequence is noted in the third column. The protein domain in which the predicted change appears is listed in the fourth column.
Data for Ptp69DY98, Ptp69DY100, and Ptp69DY125 were reported in Newsome et al. (2000).
confers a very weak viability defect. In fact, Ptp69D10 homozygotes survive even at 25°. The temperature-sensitive nature of Ptp69D10 is unmasked in hemizygotes; although two-thirds survive to the adult stage, only 11% remain free of the food at 25°, whereas adult survival is 100% at 18°. The observation that Ptp69D10 homozygotes survive while hemizygotes do not indicates that boosting the residual DPTP69D activity in Ptp69D10 hemizygotes by a factor of 2 or less is sufficient to support viability at 25°. Moreover, it is likely that reducing DPTP69D activity by a small amount could decrease the hemizygous eclosion and/or pupation rates. Therefore, mutations in genes whose products function in or regulate DPTP69D signal transduction are likely to dominantly modify the lethal phenotype of Ptp69D10 hemizygotes.
D1 inactive mutants retain residual activity: Ptp69D21 is weakly temperature sensitive as the pupation rate jumps from 6% at 25° to 32% at 18° for hemizygotes and from 11 to 35% for homozygotes. (Interestingly, the pupation rate of animals hemizyogous for another active site mutant, Ptp69DY125, is also weakly temperature sensitive, rising from 11% at 25° to 25% at 18°.) It is surprising that Ptp69D21 is temperature sensitive because the cysteine-to-tyrosine change in the D1 domain should eliminate catalytic activity at all temperatures. Several explanations could account for this temperature sensitivity. For example, the function of DPTP69D could be more critical at higher temperatures, although several other strong alleles, including Ptp69D7, Ptp69D8, Ptp69D9, and Ptp69D14, confer temperature-independent phenotypes. Alternatively, the introduction of a bulky tyrosine residue into the active site could disrupt the structure of the D1 domain at higher temperatures. Such disruption could destabilize the intracellular domain or inhibit noncatalytic aspects of DPTP69D function, such as docking. It is therefore of interest to determine if the loss of D1 activity is sufficient to mimic the viability defects
No intra-allelic complementation
| Genotype . | % pupation . | N . | % eclosion . | N . | % adulthood . | N . |
|---|---|---|---|---|---|---|
| w; Ptp69D10/Dr(3L)8ex34 | 101 | 773 | 61 | 646 | 27 | 554 |
| w; Ptp69D10/Ptp69D7 | 72 | 467 | 49 | 403 | 1 | 332 |
| w; Ptp69D10/Ptp69D21 | 61 | 441 | 39 | 331 | 1 | 282 |
| w; Ptp69D10/Ptp69DY125 | 91 | 535 | 65 | 437 | 29 | 375 |
| Genotype . | % pupation . | N . | % eclosion . | N . | % adulthood . | N . |
|---|---|---|---|---|---|---|
| w; Ptp69D10/Dr(3L)8ex34 | 101 | 773 | 61 | 646 | 27 | 554 |
| w; Ptp69D10/Ptp69D7 | 72 | 467 | 49 | 403 | 1 | 332 |
| w; Ptp69D10/Ptp69D21 | 61 | 441 | 39 | 331 | 1 | 282 |
| w; Ptp69D10/Ptp69DY125 | 91 | 535 | 65 | 437 | 29 | 375 |
Dependence of survival on allele type is shown. Viability defects conferred by alleles that encode extracellular defects are as severe in hemizyogtes as they are in trans to alleles that encode intracellular defects. N, number scored.
No intra-allelic complementation
| Genotype . | % pupation . | N . | % eclosion . | N . | % adulthood . | N . |
|---|---|---|---|---|---|---|
| w; Ptp69D10/Dr(3L)8ex34 | 101 | 773 | 61 | 646 | 27 | 554 |
| w; Ptp69D10/Ptp69D7 | 72 | 467 | 49 | 403 | 1 | 332 |
| w; Ptp69D10/Ptp69D21 | 61 | 441 | 39 | 331 | 1 | 282 |
| w; Ptp69D10/Ptp69DY125 | 91 | 535 | 65 | 437 | 29 | 375 |
| Genotype . | % pupation . | N . | % eclosion . | N . | % adulthood . | N . |
|---|---|---|---|---|---|---|
| w; Ptp69D10/Dr(3L)8ex34 | 101 | 773 | 61 | 646 | 27 | 554 |
| w; Ptp69D10/Ptp69D7 | 72 | 467 | 49 | 403 | 1 | 332 |
| w; Ptp69D10/Ptp69D21 | 61 | 441 | 39 | 331 | 1 | 282 |
| w; Ptp69D10/Ptp69DY125 | 91 | 535 | 65 | 437 | 29 | 375 |
Dependence of survival on allele type is shown. Viability defects conferred by alleles that encode extracellular defects are as severe in hemizyogtes as they are in trans to alleles that encode intracellular defects. N, number scored.
conferred by Ptp69D21 at 25°. Two other mutants, DPTP69D20 and DPTP69DY125, have active site substitutions predicted to decrease or eliminate phosphatase activity and should enable us to address this question. In any case, it appears that elimination of D1 phosphatase activity does not completely abolish DPTP69D function, at least at 18°.
Developmental requirement for DPTP69D function: DPTP69D is required during embryogenesis and throughout the pupal stage for full viability. The coincidence of the pupal and embryonic requirements for DPTP69D with axonogenesis suggests that axon defects underlie the morbidity of Ptp69D mutants. This conclusion is supported by the suppression of both viability and axon defects of Ptp69D mutants by selective expression of Ptp69D cDNA in the nervous system (Garrity et al. 1999). Motor axon defects found in Ptp69D mutants could account for their decreased fitness, as essential behaviors such as feeding and locomotion depend upon proper neuromuscular development. However, uncharacterized defects, such as CNS axon guidance errors, could also debilitate Ptp69D mutants. In particular, the requirement for function in late-stage pupae could indicate a role for DPTP69D in synaptogenesis.
Two primary goals motivated this work. First, we wanted to probe the structure of DPTP69D by analyzing mutants with reduced or otherwise altered function. Second, we wished to generate tools with which to investigate the DPTP69D signaling pathway. By characterizing our panel of new alleles, we have been able to define the developmental stages during which Ptp69D function is required. We have identified allelic combinations with a threshold of DPTP69D function, which are being used to screen for mutations that modify DPTP69D signaling. We have also discovered a putative “substrate trap” mutant with a severe axon guidance phenotype, which is being used to purify substrates of DPTP69D. Thus the alleles described in this article open the door to genetic and biochemical study of DPTP69D signal transduction.
Footnotes
Communicating editor: K. V. Anderson
Acknowledgement
We thank Anna Ingalls, Sharon Sams, Amir Lagstein, and Hong Wan for help with fly culture and mutagenesis; David Miller and Peter Kolodziej for critical reading of this manuscript and helpful comments; Kai Zinn for MAbs 3F11, 3F12, and 5A6; and Cory Goodman for MAb 1D4. J.P. was supported by a Training Program in Developmental Biology grant from the National Institutes of Health (NIH; 5T32HD0750205). This work was supported by a grant from the NIH (NSHD38141) to C.D. as well as a Basil O'Conner Starter Scholars Award (5-FY99-0111) from the March of Dimes Birth Defects Foundation.
LITERATURE CITED