-
PDF
- Split View
-
Views
-
Cite
Cite
Niedharsan Pooranachandran, Jarema J Malicki, Unexpected Roles for Ciliary Kinesins and Intraflagellar Transport Proteins, Genetics, Volume 203, Issue 2, 1 June 2016, Pages 771–785, https://doi.org/10.1534/genetics.115.180943
Close - Share Icon Share
Abstract
Transport of proteins in the ciliary shaft is driven by microtubule-dependent motors, kinesins. Prior studies suggested that the heterotrimeric ciliary kinesin may be dispensable for certain aspects of transport in specialized cilia of vertebrate photoreceptor cells. To test this possibility further, we analyzed the mutant phenotype of the zebrafish kif3a gene, which encodes the common motor subunit of heterotrimeric ciliary kinesins. Cilia are absent in all organs examined, leading to the conclusion that kif3a is indispensable for ciliogenesis in all cells, including photoreceptors. Unexpectedly, kif3a function precedes ciliogenesis as ciliary basal bodies are mispositioned in mutant photoreceptors. This phenotype is much less pronounced in intraflagellar transport (IFT) mutants and reveals that kif3a has a much broader role than previously assumed. Despite the severity of their basal body phenotype, kif3a mutant photoreceptors survive longer compared to those in IFT mutants, which display much weaker basal body mispositioning. This effect is absent in kif3a;IFT double mutants, indicating that IFT proteins have ciliary transport-independent roles, which add to the severity of their photoreceptor phenotype. kif3a is dispensable for basal body docking in otic vesicle sensory epithelia and, surprisingly, short cilia form in mechanosensory cristae even in the absence of kif3a. In contrast to Kif3a, the functions of the Kif3c-related protein, encoded by the kif3c-like (kif3cl) gene, and the homodimeric ciliary kinesin, kif17, are dispensable for photoreceptor morphogenesis. These studies demonstrate unexpected new roles for both ciliary heterotrimeric kinesins and IFT particle genes and clarify the function of kif17, the homodimeric ciliary kinesin gene.
CILIA are multifunctional organelles that form on the surface of cells. Almost without exceptions, their structure is supported by nine microtubule doublets that extend from the mother centriole, also known as the ciliary basal body (Satir and Christensen 2006; Carvalho-Santos et al. 2011; Ishikawa and Marshall 2011). Most motile cilia contain an additional pair of microtubules in the center of the ciliary shaft and function by generating fluid flow. Their motility is essential for a number of processes, including left–right asymmetry formation, mucus clearance in the respirator system, and the movement of sperm cells (Satir and Christensen 2006; Lindemann and Lesich 2010; Hirokawa et al. 2012). Nonmotile cilia, on the other hand, function by housing components of signaling pathways involved in processes as diverse as morphogenesis, cell growth and proliferation, neuromodulation, mechanosensation, olfaction, vision, and metabolism (Kennedy and Malicki 2009; Goetz and Anderson 2010; Louvi and Grove 2011; Basten and Giles 2013).
The formation and function of the cilium require intraflagellar transport (IFT), a crucial mechanism for delivering both structural and functional components into the cilium (Cole et al. 1998; Scholey and Anderson 2006). The bidirectional movement of IFT along the axoneme requires microtubule-dependent motors, kinesins, and dyneins for anterograde and retrograde transport, respectively. Members of the kinesin 2 family are the main motors for anterograde IFT (Kozminski et al. 1995; Nonaka et al. 1998; Sarpal et al. 2003; Snow et al. 2004; Evans et al. 2006; Zhao et al. 2012). Two motors belong to this family: the homodimeric kinesin, which functions as a dimer of a single subunit, and the heterotrimeric one, which contains two divergent motor subunits and an ancillary KAP subunit (recently reviewed in Scholey 2013).
In vertebrates, the heterotrimeric kinesin 2 is thought to exist in two forms, Kif3A/Kif3B/KAP3 and Kif3A/Kif3C/KAP3, which differ by one motor subunit (Muresan et al. 1998; Yang and Goldstein 1998; Marszalek et al. 1999; Zhao et al. 2012). Mouse mutations in kif3a or kif3b cause embryonic lethality, whereas kif3c mutants are viable and fertile, suggesting that this subunit is of secondary importance (Nonaka et al. 1998; Marszalek et al. 1999; Yang et al. 2001). In zebrafish, kif3c and kif3b function redundantly in certain tissues including the photoreceptor cell layer in the retina and auditory cristae (Zhao et al. 2012). The homodimeric kinesin, Kif17, on the other hand, appears to carry out more specialized functions, as its loss affects morphology only in olfactory cilia (Zhao et al. 2012).
Although ciliary kinesins have been studied for many years now, their functions are not fully understood. The role of photoreceptor ciliary kinesins in particular appeared difficult to interpret (Avasthi et al. 2009). Photoreceptor cells are unusual in that they differentiate cilia of exceptionally large size and complex morphology (Rodieck 1973; Steinberg et al. 1980; Kennedy and Malicki 2009). Using the zebrafish as a model organism, we show that kif3a is required for cilia formation in nearly all cells, including photoreceptors. Unexpectedly, however, a subset of mechanosensory hair cells differentiates short cilia even in the absence of kif3a gene function. Similar to wild-type cilia, these rudimentary kif3a−/− cilia are supported by microtubule cytoskeleton. Moreover, kif3a function extends beyond IFT and is also necessary for basal body anchoring in photoreceptors but not in hair cells. Although basal bodies dock improperly, transition zone appears to form, at least in part, in kif3a−/− mutant photoreceptors. IFT genes may also be necessary for basal body positioning, although to a lesser degree. In contrast to the basal body phenotype, photoreceptor degeneration is markedly stronger in IFT particle mutants, compared to kif3a mutant animals. Double mutants of kif3a and IFT particle genes display a similar degree of photoreceptor degeneration as IFT single mutants but a similar extent of basal body displacement to kif3a single mutants. These observations demonstrate that kif3a and IFT particle genes display both common and independent functions. In contrast to kif3a, the kif3c-related gene kif3cl and the homodimeric ciliary kinesin, kif17, are not necessary for ciliogenesis and do not contribute to the morphogenesis of photoreceptor cilia. These studies shed new light on kinesin function in vertebrate ciliogenesis.
Materials and Methods
Zebrafish strains and maintenance
kif3asa1617 and kif17sa18340 mutant alleles were obtained from the Sanger Institute. Several kif3cl mutant alleles, kif3clsh348, kif3clsh349, kif3clsh350 were generated using TALEN nucleases as described previously (Yao Zu et al. 2013). All other alleles used in this work were reported previously (Doerre and Malicki 2002; Tsujikawa and Malicki 2004; Omori et al. 2008; Zhao et al. 2012). Zebrafish were maintained in accordance with the UK Home Office regulations and UK Animals (Scientific Procedures) Act 1986.
Morpholino knockdown experiments
Morpholino knockdown experiments were performed as described previously (Tsujikawa and Malicki 2004; Omori et al. 2008). The following morpholinos were used: GTCCAGCTTATTGCTCGGCATTATC for kif3a and CCTCTTACCTCAGT TACAATTTATA as the control morpholino.
Immunohistochemistry and microscopy
Sectioning and immunohistochemistry were performed using standard protocols (Malicki et al. 2011). Transverse cryosections, 15 µm thick, through the retina or the ear were used for staining with the following antibodies: anti-acetylated α-tubulin (1:500; Sigma), zpr3 (1:500; Zebrafish International Resource Center), zpr1 (1:500; Zebrafish International Resource Center), anti-Ift88 (1:200; gift from Brian Perkins), anti-γ-tubulin (1:200; Sigma), anti-α-tubulin (1:1000; GeneTex), anti-BBS4 (1:400; gift from Nicholas Katsanis), anti-CC2D2A (1:20; gift from Ruxandra Bachmann-Gagescu), and anti-Kif17 (1:1000; Abcam). Alexa Fluor 488- and 647-conjugated secondary antibodies were used for all staining procedures. Images of cryosections were collected using an Olympus FV1000 confocal microscope or the PerkinElmer spinning disk system, using a ×20, ×40, or ×60 lens. Images of immunostained whole embryos were collected using the Olympus FV1000 confocal microscope with either a ×40 or ×60 water dipping lens.
Stochastic optical reconstruction microscopy (STORM) images were collected using the N-STORM system installed on a Nikon Eclipse Ti-E microscope, equipped with an Apo TIRF ×100 1.49 oil lens and processed using the Nikon Elements software package.
Opsin trafficking assay
To evaluate opsin trafficking into photoreceptor outer segments, a heat-shock promoter-driven GFP-opsinCT44 construct was injected into one-cell-stage embryos (Zhao and Malicki 2011; Zhao et al. 2012). Larvae were subsequently heat shocked at 37° for 30 min at 3 days postfertilization (dpf), fixed at 4, 9, and 24 hr post-heat shock, and cryosectioned using standard protocols. Sections were counterstained with DAPI and imaged using the Olympus FV1000 confocal microscope.
Transmission electron microscopy
For electron microscopy, embryos were fixed in Karnovsky’s fixative and processed as described previously (Doerre and Malicki 2002). Images were collected on a FEI Tecnai G2 microscope using a Gatan Orius SC1000B camera and Gatan Digital Micrograph software. The distance between basal bodies and the retinal pigment epithelium (RPE) was measured by first drawing a line in Adobe Photoshop along the apical surface of the RPE. Subsequently, FIJI software was used to measure the distance from the docked end of the basal body to the apical surface of the RPE. In the case of basal bodies that were not obviously docked, the centriole closer to the RPE layer was used to measure the distance to the RPE. To avoid bias in basal body position measurements due to sectioning angle, we only imaged sections through the center of the retina that were perpendicular to the body axis. In addition, we evaluated the ratio between the long and short axes of photoreceptor nuclei and found no significant differences between mutant and wild-type samples.
Quantification and image analysis
To evaluate photoreceptor survival, the number of photoreceptor cells was counted within a defined region of the photoreceptor cell layer. This region was delineated by using a 30° angle on either side of a straight line drawn from the center of the lens to the midpoint of the photoreceptor cell layer. Photoreceptor cells that fell within this area were counted.
Statistical analysis
Statistical significance was calculated using a two-tailed t-test or two-tailed Mann–Whitney U-test. P-values of <0.05 were deemed significant.
Western blotting
Fifty whole embryos at 5 dpf were homogenized in a lysis buffer. The lysate was diluted in 4× SDS buffer and boiled at 95° for 5 min. Proteins were resolved by SDS–PAGE, transferred to nitrocellulose membrane (Amersham GE Healthcare), and incubated with antibodies. The primary antibodies used were mouse anti-α-tubulin (1:5000; Genetex) and rabbit anti-kif17 (1:1000; Abcam). Goat anti-mouse DyLight 800 4× PEG conjugate (1:5000; ThermoFisher), and goat anti-rabbit Alexa Fluor 680 conjugate (1:5000; ThermoFisher) were used as secondary antibodies. Proteins were visualized with the Odyssey SA Infrared Imaging System (LI-COR Biosciences) and signal intensity was measured using the Image Studio Lite program (LI-COR Biosciences).
Reagents and data availability
All animal strains and reagents will be distributed through international stock centers or directly by the Malicki laboratory. The authors state that all data necessary for confirming the conclusions presented in the article are represented fully within the article.
Results
kif3a is required for ciliogenesis
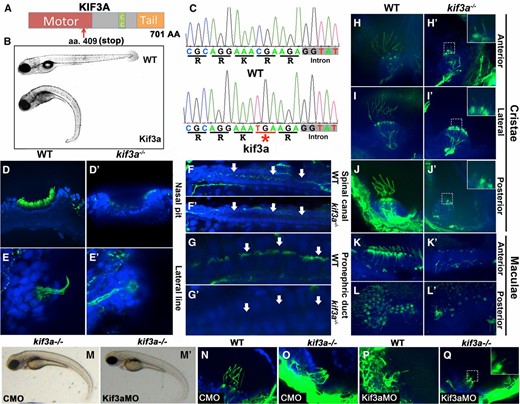
Mutations in kif3a kinesin cause early embryonic lethality in the mouse (Marszalek et al. 1999). Consequently, the analysis of kif3a function at later stages of development requires the use of conditional mutant alleles (Marszalek et al. 2000; Jimeno et al. 2006a; Avasthi et al. 2009). This is sometimes challenging due to the lack of appropriate tissue-specific promoters. In zebrafish, the presence of maternal contribution may allow one in some cases to analyze the function of housekeeping genes, such as the ones that encode cytoplasmic motor components, for example, beyond early morphogenetic events that take place during the first 2 days of development (Doerre and Malicki 2001; Jing and Malicki 2009). To determine kif3a function in vertebrate development, we analyzed its mutant phenotype in zebrafish using a truncating mutant allele, kif3asa1617, which contains a nonsense codon at position 409 of the open reading frame. This mutation eliminates the C-terminal portion of the motor domain as well as both the stalk and the tail domains resulting, in all likelihood, in a null phenotype (Figure 1A). While heterozygotes have wild-type appearance, homozygous mutants display phenotypes typical of ciliogenesis defects: curved body axis by 2.5 dpf and kidney cyst by 5 dpf (Figure 1B). We did not find left–right asymmetry defects, however. Consistent with these phenotypes, the loss of kif3a function results in the absence of cilia in the nasal pit (Figure 1, D and D′), the lateral line (Figure 1, E and E′), the spinal canal (Figure 1, F and F′), and the kidney (Figure 1, G and G′). Furthermore, cilia were largely absent in both cristae (Figure 1, H–J′) and maculae (Figure 1, K–L′) in the ear. We did, nonetheless, observe short cilia stumps in auditory cristae (Figure 1, H′, I′, and J′, insets). In zebrafish, wild-type cristae differentiate particularly long cilia of ∼20–25 μm at 5 dpf. In the mutant, these cilia average ∼2.5 μm and ∼3 μm at 3 and 5 dpf, respectively, and reach up to 5.4 μm in length. Although much shorter than wild-type cilia, they are nonetheless longer than many primary cilia in animal models or in tissue culture cells (Kramer-Zucker et al. 2005; Wilson et al. 2012; Lu et al. 2015a). The presence of cilia in kif3asa1617−/− mutant cristae suggests that another motor contributes to ciliogenesis in this tissue.
Cilia phenotype in kif3a mutants. (A) Schematic of the Kif3a protein domain structure. CC, coiled coil domain. Red arrow indicates approximate site of the stop codon in the sa1617 allele. (B) Phenotypes of wild-type (top) and kif3a−/− (bottom) zebrafish larvae at 5 dpf. (C) Sequences of wild-type and mutant kif3a−/− protein. The mutation (underlined in red and indicated with an asterisk) is in the motor domain and results in a premature stop codon. (D–L′) Whole mount immunostaining of wild-type and kif3a−/− mutant embryos at 5 dpf in several tissues as indicated. Cilia are labeled with antiacetylated tubulin antibody in green and cell nuclei are counterstained with DAPI in blue. (D and D′) Nasal pit, (E and E′) lateral line, (F and F′) spinal canal, (G and G′) kidney, and (H–L′) sensory patches of the ear. Arrows indicate kidney and spinal canal lumen. Insets in H′, I′, and J′ and Q show stubs of cilia in cristae. (M′) Morpholino knockdown of kif3a in kif3a−/− mutants does not exacerbate the body curvature at 3 dpf compared to (M) control morpholino knockdown. (N–Q) Whole mount immunostaining of lateral cristae with antiacetylated α-tubulin antibody (in green) at 3 dpf. (N) The phenotype of control morpholino (CMO)-treated WT embryos, (O) CMO-treated kif3a−/− embryos, (P) kif3aMO-treated WT embryos and (Q) Kif3aMO-treated kif3a−/− embryos.
To assess whether the kif3asa1617−/− mutant allele is null, we performed morpholino knockdown experiments (Figure 1, M–Q). The external body curvature of kif3a−/− embryos was not exacerbated by kif3a morpholino (kif3aMO) knockdown in the kif3asa1617−/− mutant background, compared to control morpholino (CMO) knockdown (Figure 1, M and M′). Similarly, the ciliary stubs observed in lateral cristae of kif3a−/− embryos remained even after kif3aMO knockdown (Figure 1, N–Q). These data provide further evidence that kif3asa1617−/− mutants are amorphic. The above observations support the conclusion that the Kif3a motor subunit is necessary for cilia differentiation in most tissues, including photoreceptors, although additional ciliary transport mechanism may exist in auditory cristae.
Photoreceptor phenotype of kif3a−/− mutants
Vertebrate photoreceptor cells feature unusually bulky cilia. Their membranes form hundreds of folds containing the visual pigment, opsin, and other component of the phototransduction apparatus (Rodieck 1973; Kennedy and Malicki 2009). It is estimated that the photoreceptor cilium of some species contains 109 of opsin molecules (Pugh and Lamb 2000), and opsin appears to be transported into the cilium at the rate of 100–1000 molecules per second (Malicki and Besharse 2012). Given this unusually intense stream of ciliary traffic, photoreceptor cilia may require additional motors. Genetic analysis in the mouse suggested, in fact, that this is the case: conditional mouse kif3a−/− mutants differentiate abnormal but nonetheless fairly bulky cilia in both rods and cones (Avasthi et al. 2009). These experiments suggested that protein transport sufficient for outer segment formation is present in cilia even in the absence of the kif3a motor subunit. To test this possibility further, we investigated the photoreceptor phenotype of kif3a−/− mutant zebrafish. Staining with antiacetylated tubulin antibody revealed that photoreceptor cilia are absent in kif3a−/− mutant homozygotes at 3 dpf (Figure 2, A and A′).
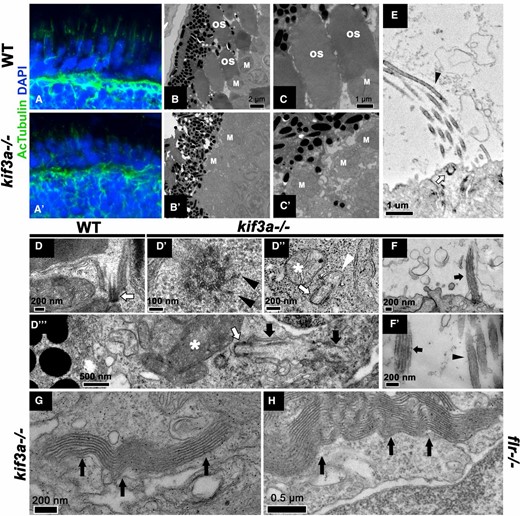
Photoreceptor phenotype in kif3a mutants. (A and A′) Cryosections through the wild-type (A) and kif3a−/− mutant (A′) retina immunostained for acetylated tubulin in green and counterstained with DAPI in blue. Cilia are absent in (A′) kif3a−/− embryos at 3 dpf. (B–C′) Transmission electron microscopy of photoreceptors in wild-type (B and C) and kif3a−/− (B′ and C′) retinae at 5 dpf. Outer segments are missing in the mutant. Bars, 2 µm (B and B′) and 1 µm (C and C′). OS, outer segment; M, mitochondrial clusters. (D–D′′′) Transmission electron microscopy of the basal body in wild-type (D) and kif3a−/− (D′–D′′′) embryos at 5 dpf. (D′) Transverse cross-section through kif3a−/− mutant basal body. Transition fibers (arrowheads) differentiate on at least some basal bodies. Bar, 100 nm. (D′′) An example of a basal body (arrow) attached to a preciliary vesicle (arrowhead). Bar, 200 nm. (D′′′) An example of a membrane-docked basal body (arrow). Note that although it forms axonemal microtubules, it does not assemble outer segment membrane folds. Bar, 0.5 µm. White arrows, basal body; black arrows, ciliary microtubules; asterisks, mitochondrial clusters. (E) Transmission electron microscopy of stereocilia in the lateral crista of kif3a−/− embryos at 3 dpf. White arrow indicates docked basal body; black arrowhead indicates stereocilia. Bar, 1 um. (F) Transmission electron microscopy of cilia in the lateral crista of kif3a−/− embryos at (F) 5 dpf and (F′) 3 dpf. Black arrows indicate cilia and black arrowheads stereocilia. (G and H) Ectopic membranes are found in kif3a−/− (G) and fleer−/− (H) photoreceptors at 5 dpf. In G and H, apical is left; black arrows, ectopic membrane folds.
To investigate this phenotype further, we performed ultrastructural analysis. kif3a−/− mutant photoreceptors differentiate fairly robust inner segments as evidenced by the presence of apical clusters of mitochondria (Figure 2, B′–C′). The cilia (outer segments) of the same cells are, however, completely absent. These results indicate that similar to IFT mutants, kif3a−/− animals do not differentiate apical photoreceptor outer segments (Doerre and Malicki 2002; Pazour et al. 2002; Tsujikawa and Malicki 2004). Unexpectedly, a closer inspection of basal bodies in kif3a−/− mutant photoreceptors at 5 dpf revealed that they frequently (∼60%) do not localize to the apical surface of inner segments (Figure 2, D–D′′′; Table 1), even though they appear to dock to intracellular membranes (Figure 2D′′). At 3 dpf, many kif3a−/− basal bodies (∼20%) are found deep in the inner segment cytoplasm surrounded by mitochondria (Figure 3, G and G′; Table 1). Although in one case we observed a photoreceptor cell that differentiated a rudimentary cilium, the basal body of this cilium localized basally to the mitochondrial cluster of the inner segment and the cilium itself extended away from the RPE and the apical terminus of the cell (Figure 2D′′′). This further illustrates the role of kif3a in the positioning of basal bodies. Although mislocalized, kif3a−/− basal bodies appear to form normal transition fibers and on occasion are found associated with a vesicle (Figure 2, D′ and D′′). Such docking of basal bodies to a cytoplasmic vesicle has been described at early stages of ciliogenesis in several cell types but not, to our knowledge, in photoreceptors (Sorokin 1962, 1968; Lu et al. 2015b). Thus kif3a does not appear to function in basal body docking to the membrane, but is necessary for basal body localization in the apical cytoplasm of photoreceptors.
Localization and membrane docking of basal bodies
| Genotype . | Basal body localization . | n . | Docked (%) . | n . | |||
|---|---|---|---|---|---|---|---|
| Apical (%) . | Basal (%) . | Central (%) . | Lateral (%) . | ||||
| kif3a−/− | 9 (33) | 7 (26) | 6 (22) | 5 (19) | 27 | 10 (59) | 17 |
| elipsa−/− | 16 (67) | 2 (8) | 2 (8) | 4 (17) | 24 | 9 (60) | 15 |
| Fleer−/− | 16 (70) | 3 (13) | 4 (17) | 0 (0) | 23 | 14 (61) | 23 |
| kif3a−/−; eli−/− | 5 (38) | 6 (23) | 9 (35) | 6 (23) | 26 | 4 (40) | 10 |
| Kif3a−/− 5 dpf | 9 (39) | 5 (22) | 2 (9) | 7 (30) | 23 | 10 (50) | 20 |
| Genotype . | Basal body localization . | n . | Docked (%) . | n . | |||
|---|---|---|---|---|---|---|---|
| Apical (%) . | Basal (%) . | Central (%) . | Lateral (%) . | ||||
| kif3a−/− | 9 (33) | 7 (26) | 6 (22) | 5 (19) | 27 | 10 (59) | 17 |
| elipsa−/− | 16 (67) | 2 (8) | 2 (8) | 4 (17) | 24 | 9 (60) | 15 |
| Fleer−/− | 16 (70) | 3 (13) | 4 (17) | 0 (0) | 23 | 14 (61) | 23 |
| kif3a−/−; eli−/− | 5 (38) | 6 (23) | 9 (35) | 6 (23) | 26 | 4 (40) | 10 |
| Kif3a−/− 5 dpf | 9 (39) | 5 (22) | 2 (9) | 7 (30) | 23 | 10 (50) | 20 |
Localization of basal bodies in photoreceptor cells relative to mitochondrial clusters for the following genotypes: kif3a−/−, elipsa−/−, fleer−/−, kif3a−/−;elipsa−/− at 3 dpf and for kif3a−/− at 5 dpf. Docking of basal bodies to the membrane, including membranes of intracellular vesicles, was also evaluated. Only sections that included the long axis of either the mother or the daughter centriole were considered in counts of docked basal bodies. If all centrosomes dock to the membrane, this analysis is expected to show 50% of centrioles as attached to the membrane.
| Genotype . | Basal body localization . | n . | Docked (%) . | n . | |||
|---|---|---|---|---|---|---|---|
| Apical (%) . | Basal (%) . | Central (%) . | Lateral (%) . | ||||
| kif3a−/− | 9 (33) | 7 (26) | 6 (22) | 5 (19) | 27 | 10 (59) | 17 |
| elipsa−/− | 16 (67) | 2 (8) | 2 (8) | 4 (17) | 24 | 9 (60) | 15 |
| Fleer−/− | 16 (70) | 3 (13) | 4 (17) | 0 (0) | 23 | 14 (61) | 23 |
| kif3a−/−; eli−/− | 5 (38) | 6 (23) | 9 (35) | 6 (23) | 26 | 4 (40) | 10 |
| Kif3a−/− 5 dpf | 9 (39) | 5 (22) | 2 (9) | 7 (30) | 23 | 10 (50) | 20 |
| Genotype . | Basal body localization . | n . | Docked (%) . | n . | |||
|---|---|---|---|---|---|---|---|
| Apical (%) . | Basal (%) . | Central (%) . | Lateral (%) . | ||||
| kif3a−/− | 9 (33) | 7 (26) | 6 (22) | 5 (19) | 27 | 10 (59) | 17 |
| elipsa−/− | 16 (67) | 2 (8) | 2 (8) | 4 (17) | 24 | 9 (60) | 15 |
| Fleer−/− | 16 (70) | 3 (13) | 4 (17) | 0 (0) | 23 | 14 (61) | 23 |
| kif3a−/−; eli−/− | 5 (38) | 6 (23) | 9 (35) | 6 (23) | 26 | 4 (40) | 10 |
| Kif3a−/− 5 dpf | 9 (39) | 5 (22) | 2 (9) | 7 (30) | 23 | 10 (50) | 20 |
Localization of basal bodies in photoreceptor cells relative to mitochondrial clusters for the following genotypes: kif3a−/−, elipsa−/−, fleer−/−, kif3a−/−;elipsa−/− at 3 dpf and for kif3a−/− at 5 dpf. Docking of basal bodies to the membrane, including membranes of intracellular vesicles, was also evaluated. Only sections that included the long axis of either the mother or the daughter centriole were considered in counts of docked basal bodies. If all centrosomes dock to the membrane, this analysis is expected to show 50% of centrioles as attached to the membrane.
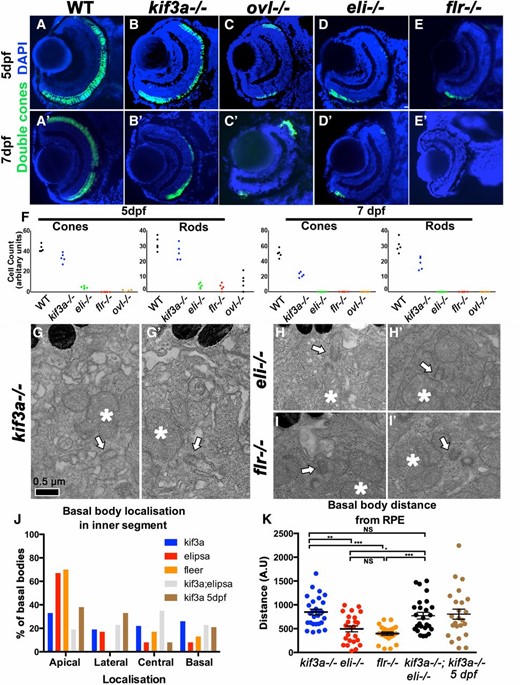
Photoreceptor survival and basal body phenotype in kif3a−/− and IFT mutants. (A–E′) Transverse cryosections through wild-type, kif3a−/−, oval−/−, elipsa−/−, and fleer−/− retinae as indicated, stained for double cones. Some loss of cones is observed in kif3a−/− (B) mutant compared to the wild type (A) at 5 dpf. A much more pronounced photoreceptor loss is seen in IFT mutants (C–E). Similar phenotypes are observed at 7 dpf (A′–E′). (F) Graphs illustrating the survival rates of cones and rods in wild type, kif3a−/−, oval−/−, elipsa−/−, and fleer−/− at 5 dpf and 7 dpf, as indicated. (G–I′) Transmission electron microscopy of basal bodies in kif3a−/− (G and G′), elipsa−/− (H and H′), and fleer−/− (I and I′) at 3 dpf. Bar, 0.5 µm. (J) Graph illustrating the percentage of basal bodies that localize to the apical, basal, lateral, or central (i.e., in the middle of the mitochondrial cluster) regions of the inner segment, in kif3a−/−, elipsa−/−, fleer−/−, and kif3a−/−;elipsa−/− double mutant at 3 dpf, and in kif3a−/− at 5 dpf as indicated. (K) Graph illustrating the distance between basal bodies and the RPE in kif3a−/−, elipsa−/−, fleer−/−, and kif3a−/−;elipsa−/− double mutants at 3 dpf, and in kif3a−/− at 5 dpf. The mean and standard error bars are indicated. For each experimental group, ≥23 basal bodies from three to four retinae were evaluated. *P < 0.05; **P < 0.01; ***P ≤ 0.001; NS, not significant, based on Mann–Whitney U-test. In G′–I′, apical is up. Asterisks, mitochondrial clusters; white arrows, basal bodies.
In contrast to photoreceptor cells, kif3a is not required to dock basal bodies in mechanosensory hair cells and, as described above, short cilia differentiate in mechanosensory cristae of kif3a−/− animals (Figure 1, H–J′). To further assess ciliogenesis in kif3a−/− mutants, we investigated the ultrastructure of cristae and maculae (Figure 2, E–F′). In nearly all cases examined (36/37), we found that basal bodies of these tissues are anchored to the apical surface (Figure 2E, arrow). Similar to wild-type cilia, they are supported by microtubules both at 3 and 5 dpf (Figure 2, F and F′).
It is noteworthy that multilayered membrane structures reminiscent of outer segments are observed ectopically in the retinae of kif3a−/− mutants at both 3 (5 of 26 sections analyzed) and 5 (13 of 27 sections analyzed) dpf (Figure 2, G and H). Such structures were previously found in the zebrafish oval/ift88 mutant (Tsujikawa and Malicki 2004), and we have now also observed them in fleer/ift70 mutants (Figure 2H). In both fleer−/− and kif3a−/− mutants, these ectopic membrane arrays differentiate on the lateral surface of photoreceptors and most of them do not contact RPE cells. In some cases (n = 5), it was clear that basal bodies are positioned away from these ectopic membrane arrays in the same cell, and in a few instances when mutant basal bodies extended short axoneme, these were not associated with membrane folds that characterize the outer segment (Figure 2D′′′). These observations reveal that the assembly of outer segment membrane folds does not require kif3a and at least some IFT genes and that these structures can form in a cilia-independent manner.
kif3a and IFT particle genes in photoreceptor survival
Rapid degeneration of both cones and rods is observed in IFT mutants, including oval, elipsa, and fleer (Doerre and Malicki 2002; Tsujikawa and Malicki 2004). Because both the heterotrimeric kinesin 2 and IFT particle components are parts of the same intraflagellar transport protein complex, it is expected that loss of the motor should result in photoreceptor degeneration similar to that observed in IFT particle mutants. This, however, is not the case. Photoreceptor degeneration is slower for both cones and rods in kif3a−/− embryos, compared to elipsa, oval, and fleer, which are mutants of IFT particle components (Figure 3, A–F).
To test whether the severity of photoreceptor degeneration in kif3a−/− and IFT mutants correlate with basal body mispositioning, we compared kif3a−/−, elipsa−/−, and fleer−/− phenotypes using transmission electron microscopy. As the majority of photoreceptors have degenerated in IFT mutants by 5 dpf, we examined photoreceptor basal bodies at 3 dpf. In contrast to kif3a−/−, in mutants of two IFT genes, elipsa−/− (ift54) and fleer−/− (ift70), the majority of basal bodies were docked to the apical surface of the inner segment or localized close to the RPE (Figure 3, H–K). Detailed analysis of basal body localization revealed that kif3a−/− basal bodies localize with approximately equal frequencies: apical 33%, basal 26%, or lateral 19% relative to the inner segment mitochondrial cluster at 3 dpf (Table 1). Equally frequently, they are also found in the middle of mitochondria 22% (Figure 3J, Table 1). In contrast to that, 67% (16/24) and 70% (16/23) of basal bodies localize apically in elipsa−/− and fleer−/−, respectively (Figure 3J, Table 1). Measurements of the distance from basal bodies to the RPE revealed similarly pronounced differences between mutant phenotypes of kif3a and IFT genes (Figure 3K). These data show that kif3a functions in the apicobasal polarity of photoreceptors by positioning the basal body to the apical terminus of the cell. IFT proteins may also contribute to this process, although their role is less pronounced. Moreover, as basal body mislocalization is considerably more severe in kif3a−/− photoreceptors compared to elipsa−/− and fleer−/− mutant cells, which degenerate faster, it is not a nonspecific consequence of photoreceptor degeneration.
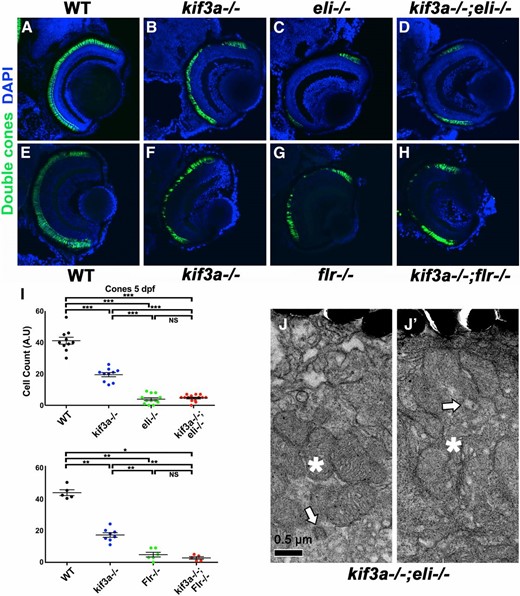
The difference between kif3a−/− and IFT mutant phenotypes is open to several interpretations. We considered a possibility that Kif3a protein accumulation in the cytoplasm of photoreceptors mutant for IFT genes is toxic to the cell. Alternatively, IFT proteins may have intraflagellar transport-independent functions in the cell. To distinguish between these possibilities, we constructed double mutants of kif3a and IFT genes. kif3a−/−;elipsa−/− or kif3a−/−;fleer−/− double mutants display photoreceptor degeneration rates similar to those in single IFT mutants (Figure 4, A–I). As cilia and ciliary outer segments are missing both in kif3a and in IFT gene mutants, which implies that intraflagellar transport is absent, these observations indicate that IFT proteins have cilia and intraflagellar transport-independent function in photoreceptors.
Double mutant phenotypes of kif3a and IFT genes. (A–H) Cryosections of wild-type, kif3a−/−, elipsa−/−, fleer−/−, and kif3a−/−;elipsa−/− and kif3a−/−;fleer−/− double mutants as indicated. Sections were immunostained to visualize cones and counterstained with DAPI to visualize nuclei. No significant differences are found in photoreceptor survival between elipsa−/− single mutants and kif3a−/−;elipsa−/− double mutants. Similarly, fleer−/− single mutants and kif3a−/−;fleer−/− double mutants display very similar photoreceptor degeneration rates. (I) Graphs illustrating cone survival rates in single and double mutants at 5 dpf as indicated. The mean and standard error bars are indicated. All experimental groups include data from five or more animals. *P ≤ 0.05; **P ≤ 0.01; ***P ≤ 0.001; NS, not significant, based on Mann–Whitney U-test. (J and J′) Transmission electron microscopy of basal bodies in kif3a−/−;elipsa−/− double mutant embryos at 3 dpf. Bar, 0.5 µm. Apical is up. Arrows, basal body; asterisks, mitochondrial clusters.
As discussed above, kif3a is necessary for basal body localization to the apical surface in photoreceptor cells. This phenotype of kif3a−/− may be mediated by an ectopic function of IFT proteins in the basal body area. To test this possibility, we investigated basal body localization in kif3a−/−;elipsa−/− double mutants and found that kif3a is necessary for basal body localization even in the absence of elipsa−/− (Figure 3, J and K, Figure 4, J and J′; Table 1). These results show that kif3a basal body positioning defect is not caused by an ectopic activity of the IFT particle.
In many cells, the centrosome acts as a microtubule organizing center. Centrosome defects in kif3a−/− mouse fibroblasts lead to microtubule disorganization (Kodani et al. 2013). As centrosomes and basal bodies are closely related, kif3a may also function in organizing photoreceptor microtubules. To investigate this possibility, we used super-resolution microscopy on kif3a−/− retinal cryosections stained with anti-α-tubulin antibodies to visualize microtubules. We did not detect, however, any obvious abnormalities in microtubule arrangement of kif3a−/− mutant photoreceptor cells (Figure 5, A–B′). Similar to kif3a−/−, BBS4 loss of function results in microtubule disorganization (Kim et al. 2004). BBS4 is mislocalized in photoreceptors of zebrafish mikre oko/dynactin p150 mutants (Tsujikawa et al. 2007), and kif3a is required to localize p150 to subdistal appendages of centrioles (Kodani et al. 2013). These observations suggest that kif3a may be required to localize BBS4 in photoreceptor cells. This was not, however, the case as we observed no differences in BBS4 localization between WT and kif3a−/− photoreceptors (Figure 5, D and D′).
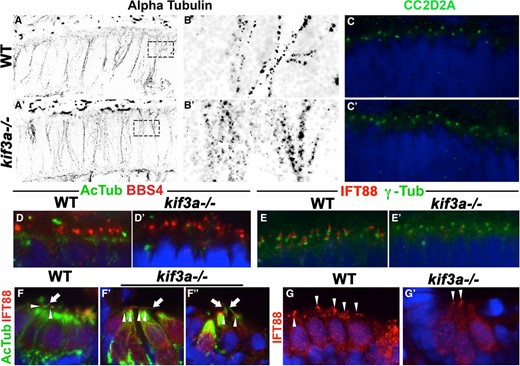
(A and A′) STORM imaging of cryosections through WT and kif3a−/− mutant retinae, immunostained with anti-α-tubulin antibody to visualize microtubules at 3 dpf. No obvious differences between WT and mutants are found. (B and B′) Enlargements of areas indicated by dashed boxes in A and A′. (C–C′) Transverse cryosections through the retina in WT and kif3a−/− mutants immunostained for CC2D2A (in green) at 3 dpf. No obvious differences between WT and mutants are observed. (D and D′) Transverse cryosections of WT and kif3a−/− mutant retinae immunostained for BBS4 (in red) and acetylated-α-tubulin (in green) at 3 dpf. No obvious differences are found between WT and mutants. (E and E′) Transverse cryosections of WT and kif3a−/− mutant retinae immunostained for IFT88 (in red) and γ-tubulin (in green) at 3 dpf. IFT88 localizes apical to γ-tubulin in WT, but is strongly reduced in kif3a−/− mutants. (F–F′′) Transverse cryosections of lateral cristae stained for IFT88 (in red) and acetylated-α-tubulin (in green) at 5 dpf. Ift88 signal persists in mutant cristae. (G and G′) Transverse cryosections of the lateral crista stained for IFT88 (in red) at 5 dpf. Ift88 localization to the apical surface of hair cells is not affected in kif3a−/− mutants. In (C–G′), sections are counterstained with DAPI (in blue) to visualize nuclei.
As kif3a−/− basal bodies dock to the membrane (Table 1), we asked whether transition zone assembly proceeds normally in kif3a−/− mutants. To address this, we determined the localization of a transition zone protein, CC2D2A/MKS6, which genetically localizes downstream of several other transition zone components (Williams et al. 2011). Surprisingly, CC2D2A puncta are still present in the same quantity (on average 15 per an arbitrary unit of length for both WT and mutant, n ≥ 9), suggesting that the transition zone forms at least partially in kif3a−/− mutants (Figure 5, C and C′). This finding is consistent with EM data showing that basal bodies differentiate transition fibers and attach to vesicles in the cytoplasm (Figure 2, D′ and D′′). kif3a is, however, required for IFT particle assembly at photoreceptor basal bodies as Ift88 protein is absent from basal bodies of kif3asa1617−/− mutants visualized with antibodies to γ-tubulin (Figure 5, E and E′). These results show that kif3a acts after the initial steps of transition zone assembly but prior to basal body translocation to the apical membrane and IFT particle assembly.
The presence of cilia in kif3a−/− mutants suggested the possibility that an kif3a-independent transport mechanism operates in cilia of cristae. To test whether this mechanism involves intraflagellar transport, we stained kif3asa1617−/− mutant maculae and cristae with antibodies to IFT88 (Figure 5, F–G′, and not shown). In kif3a−/− photoreceptors and in ear maculae, IFT88 is absent by 3 dpf (Figure 5, E and E′, and not shown). In contrast to that, IFT88 puncta are present at the cilia base in cristae of kif3a−/− mutants even at 5 dpf (Figure 5, F–G′). Thus, surprisingly, unlike in other tissues, kif3a is not required to localize IFT proteins in cristae cilia, suggesting that IFT may be functioning in this tissue with the aid of another motor.
kif3c-like is not required for ciliogenesis
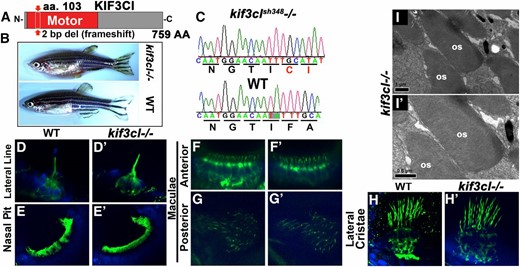
We previously showed that the zebrafish genome contains two kif3c-related genes, which we named kif3c and kif3c-like (kif3cl) (Zhao et al. 2012). The function of kif3cl was previously studied using antisense oligonucleotides. These studies did not reveal phenotypic abnormalities. To determine kif3cl function later in development and in adult individuals, we mutated it using TALEN nucleases. Three alleles were generated: kif3clsh348−/−, kif3clsh349−/−, and kif3clsh350−/−, which contain 2-, 11-, and 25-bp deletions, respectively, within the motor domain and result in frameshifts. Mutant homozygotes for all three alleles have wild-type appearance. The kif3clsh348−/− allele was used in subsequent studies (Figure 6, A and C). kif3clsh348−/− mutant homozygotes survive to adulthood, do not display phenotypic abnormalities, and are fertile (Figure 6B). The analysis of cilia in kif3clsh348−/− via antiacetylated tubulin staining did not reveal abnormalities in the lateral line, the nasal pit, or in the sensory patches of the ear (Figure 6, D–H′). Similarly, kif3clsh348−/− staining with antibodies to rod and cone photoreceptors at 5 dpf did not reveal defects (data not shown). Ultrastructural analysis of photoreceptors also showed that kif3clsh348−/− photoreceptor outer segments are normal at 5 dpf (Figure 6, I and I′). To test for more subtle defects in photoreceptor cilia, we applied opsin transport assay as described by us previously (Zhao and Malicki 2011; Zhao et al. 2012). In this assay, a GFP–opsin–CT44 fusion protein is briefly expressed from the heat-shock promoter and the rate of its translocation from the cell body to the cilium is evaluated. Also this test does not reveal abnormalities in kif3clsh348−/− mutant photoreceptors (data not shown).
kif3cl mutant phenotype. (A) Schematic of Kif3cl protein domain structure. The sh348 mutant allele contains a 2-bp deletion (red arrow) in the motor domain that results in a frameshift. (B) Phenotype of wild-type (bottom) and kif3cl−/− homozygous mutant (top) adult zebrafish. (C) Sequences of kif3cl−/− mutant and the wild-type gene. The 2-bp deletion (shaded) and the resulting frameshift are indicated. (D–H′) Whole mount immunostaining of wild-type and kif3cl−/− zebrafish with antiacetylated tubulin antibody to visualize cilia at 5 dpf (in green). Tissue is counterstained with DAPI in blue. No obvious differences are found between wild-type and mutant cilia. (D and D′) Lateral line; (E and E′) nasal pit; (F–H′) sensory patches of the ear. (I) Transmission electron microscopy of outer segments in kif3cl−/− mutants at 5 dpf. Apical is up. OS, outer segment.
It has been previously shown that the rate of opsin transport increases in the presence of stress such as elevated temperature (Young 1967). To accommodate for such an increase in transport volume, additional kinesins may be required. Thus, the effect of kif3cl absence may become obvious under stress conditions. To test this, we maintained kif3cl−/− embryos at elevated temperatures (32° and 36°) for 2.5 days starting at 2.5 dpf and stained photoreceptors with antirod opsin antibodies at 5 dpf. This treatment did not seem to affect opsin trafficking, as antibody staining did not reveal opsin mislocalization to inner segments of photoreceptor cells (data not shown). To further assess kif3cl’s role in photoreceptor ciliogenesis, we generated double mutants with kif3b and investigated the outer segments of kif3b−/−;kif3cl−/− animals. Outer segment defects of these double mutants did not appear to be obviously more severe, compared to those in kif3b−/− single mutants (data not shown). The lack of phenotype in kif3cl−/− mutants may be due to compensation by kif3c, and kif3cl may only be upregulated in the absence of kif3c. This possibility will require further genetic testing.
kif17 does not contribute to photoreceptor morphogenesis
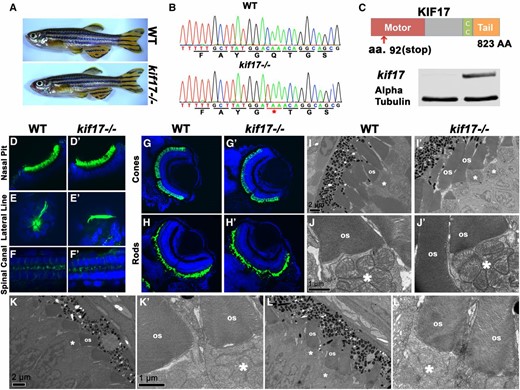
The gene encoding the homodimeric kinesin 2 in Caenorhabditis elegans, osm-3, is required for the formation of amphid cilia (Evans et al. 2006; Mukhopadhyay et al. 2007). The role of its vertebrate homolog, kif17, is, however, less clear. Morpholino knockdown of kif17 was reported to cause outer segment loss at 3 dpf (Insinna et al. 2008). In contrast to that, a study of a kif17−/− mutant allele, kif17sa0119, which contains a stop codon in the tail domain, did not reveal any photoreceptor defects and homozygous adults display normal opsin trafficking to the outer segment (Zhao et al. 2012). Although this is unlikely, it has been suggested that this allele may not entirely eliminate kif17 activity (Bader et al. 2012). To clarify kif17 function, we obtained another kif17 mutant allele, kif17sa18340, which contains a stop codon within the motor domain (Figure 7C). The resulting protein is truncated at the amino acid residue 92, effectively losing 90% of its sequence (Figure 7C). Western blot analysis confirmed the loss of Kif17 protein in kif17sa18340−/− animals, although a small amount of wild-type-length polypeptide (∼1%) remained detectable in the mutant upon blot overexposure (not shown). The adult kif17sa18340 homozygotes are viable and fertile and do not display any overt developmental defects (Figure 7A). Cilia are also morphologically normal in the lateral line, the nasal pit, and the spinal canal of kif17sa18340−/− homozygotes (Figure 7, D–F′). To investigate kif17sa18340 phenotype further, we immunostained kif17sa18340−/− mutants for rods or cones at 3 (data not shown) and 5 dpf (Figure 7, G–H′). This analysis did not reveal any abnormalities in either cone or rod photoreceptors either. To further investigate potential defects in mutant photoreceptors, we examined the ultrastructure of kif17sa18340−/− photoreceptor cells at 3 and 5 dpf (Figure 7, I–L′). Again, no obvious differences were found compared to wild-type siblings. We conclude that kif17 is not necessary for outer segment morphogenesis in vertebrate photoreceptors.
kif17 mutant phenotype. (A) Phenotypes of wild-type (top) and kif17−/− homozygous mutant (bottom) adult zebrafish. (B) Sequences of kif17−/− mutant and wild-type alleles. The mutation site is indicated with an asterisk and underlined in red. (C, top) Schematic of Kif17 protein domain structure. Red arrow indicates the approximate site of the stop codon at position 92. (C, bottom) Western blot for Kif17 in WT and kif17−/− embryos. α-Tubulin is included as a control. (D–F′) Whole mount immunostaining of wild-type and kif17−/− mutant zebrafish with antiacetylated tubulin antibody to visualize cilia at 5 dpf (in green). Tissue is counterstained with DAPI to visualize nuclei (in blue). No differences in cilia morphology are seen between mutant and wild-type animals. (D and D′) The nasal pit; (E and E′) the lateral line; (F and F′) the spinal canal. (G–H′) Cryosections through retinae of wild-type and kif17−/− mutant animals at 5 dpf immunostained for cones (G and G′) or rods (H and H′), and counterstained with DAPI to visualize nuclei. No significant differences are found in photoreceptor survival between wild type and kif17−/− mutants. (I–L′) Transmission electron microscopy showing outer segments in kif17−/− mutant and wild-type animals at 3 (I–J′) and 5 (K–L′) dpf. Bars, 1 µm for high magnification images and 2 µm for low magnification ones. Apical is up. OS, outer segment; asterisks indicate mitochondrial clusters.
Discussion
Our data show that a heterotrimeric kinesin 2 motor subunit, kif3a, is required for the formation of cilia in nearly all tissues examined, including photoreceptor cells. In contrast to that, the homodimeric kinesin 2 gene, kif17, does not play a role in vertebrate ciliogenesis in most tissues, including the photoreceptor cell layer. The kif3c-like gene, one of the two kif3c homologs in zebrafish, does not appear to be required for ciliogenesis either. The absence of photoreceptor outer segments is often associated with rapid photoreceptor degeneration. Unexpectedly, kif3a−/− photoreceptors degenerate at a slower pace, compared to IFT protein mutants. This is surprising as kif3a−/− and IFT mutants, such as oval−/−, elipsa−/−, and fleer−/− display complete absence of photoreceptor cilia (Doerre and Malicki 2002; Tsujikawa and Malicki 2004). Consequently, a similar level of photoreceptor degeneration is expected in heterotrimeric kinesin and IFT mutants. Contrary to this expectation, kinesin and IFT mutations have different impacts on photoreceptor survival. kif3a−/−;elipsa−/− and kif3a−/−;fleer−/− double mutants and IFT single mutants display similar rates of photoreceptor loss, indicating that IFT proteins display a cilia-independent function in photoreceptors. Equally unexpected is the observation that kif3a−/− functions in the polarity of photoreceptor inner segments and its absence causes mispositioning of photoreceptor basal bodies. Finally, stubs of cilia persist in the cristae of kif3a−/− mutants at 3 and 5 dpf, suggesting that partial ciliogenesis does not require kif3a in this tissue. Taken together, this analysis reveals new functions for both ciliary kinesins and genes encoding IFT particle components.
kif3a phenotypes
Several attempts to conditionally remove kif3a in mouse photoreceptor cells produced inconsistent results. Initially it was shown that opsin mislocalizes in mouse photoreceptor inner segments when kif3a is specifically deleted in photoreceptors using the IRBP-Cre or RHO-Cre drivers (Marszalek et al. 2000; Jimeno et al. 2006b). In contrast to that, another Cre driver, iCre75, which conditionally removes kif3a in mouse rods, had very little effect on rhodopsin localization (Avasthi et al. 2009). Regardless of the driver used, rod outer segments differentiated in mutant photoreceptors. When the RHO-Cre driver was used to eliminate kif3a function, rod outer segments formed normally and remained normal up to P10 (Jimeno et al. 2006a). Similarly, when the iCre75 driver was applied, rod-connecting cilia and outer segments remained well differentiated until at least P13 (Avasthi et al. 2009). Cones also differentiated fairly robust, although not entirely normal, outer segments when conditional knockdown of kif3a was generated using human red/green opsin promoter to drive cre (Avasthi et al. 2009). This led to the idea that kif3a may not be the only motor required to transport rhodopsin as well as other transmembrane proteins in rod photoreceptors and that rod degeneration is due to causes other than opsin mislocalization in this mutant line (Avasthi et al. 2009). In contrast to these reports, a study of opsin–GFP trafficking in kif3a-deficient photoreceptor cells demonstrated a role for kif3a in opsin transport (Trivedi et al. 2012).
We show that kif3a is essential for outer segment differentiation in both cones and rods of zebrafish. The difference between mouse and zebrafish data is most likely due to difficulties in conditionally removing kif3a in mouse rods at an appropriate stage before the initiation of outer segment differentiation, while at the same time avoiding lethality due to a loss of kif3a function during early embryogenesis (Marszalek et al. 1999). In the case of the iCre75 transgene, the deletion of kif3a is completed by P14 (Li et al. 2005; Avasthi et al. 2009), which is most likely too late to eliminate kif3a from mouse photoreceptors (Avasthi et al. 2009).
Interestingly, we also found that kif3a is required for basal body localization to the apical surface of inner segments. Although kif3a−/− mutant basal bodies extend transition fibers and are frequently found associated with vesicles or membrane invaginations and thus are capable of docking to membranes, they are clearly displaced from photoreceptor apical termini. This demonstrates that kif3a functions in localizing the basal body to the apical area of the photoreceptor inner segment cytoplasm. Double mutants of kif3a and elipsa display the same phenotype as kif3a single mutants, indicating that kif3a basal body mislocalization is not caused by an ectopic activity of the Elipsa protein, which may accumulate at the basal body when cilia are not formed. Basal body mislocalization is also observed in IFT mutants but is much weaker. This suggests that IFT proteins may act along kif3a to localize basal bodies, although their contribution is smaller.
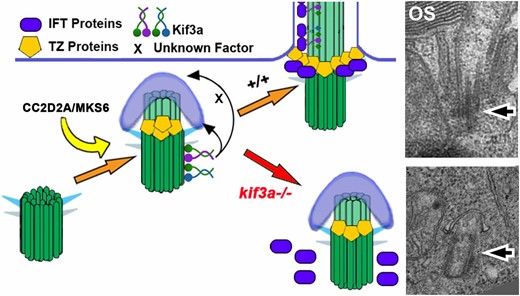
What is the mechanism of kif3a action in basal body positioning? Previously it has been reported that kif3a associates with the mother centriole and is necessary for the assembly of subdistal appendage proteins on the mother centriole (Kodani et al. 2013). Furthermore, the same study demonstrated that while microtubules of wild-type cells emerge from the centrosome in a radial pattern, microtubules of kif3a−/− mutant cells are disorganized. These findings suggest that basal body mislocalization in kif3a−/− photoreceptors could be due to the lack of microtubule organization necessary to anchor basal bodies to the apical surface. Kif3a could not only anchor the basal body to microtubules but also translocate the basal body along microtubules toward the apical surface. Although our super-resolution microscopy imaging did not reveal obvious defects in microtubule organization in kif3a−/− mutants, it remains possible that we were unable to detect a small population of subapically localized microtubules. To investigate the mechanism of basal body localization further, we tested several markers of cilia assembly, one of which was CC2D2A/MKS6, a transition zone protein that is thought to function in a docking mechanism at the base of the photoreceptor cilium for cilia-targeted vesicles (Bachmann-Gagescu et al. 2015). A number of elegant studies in C. elegans demonstrated a hierarchy in the assembly of transition zone proteins and found that MKS6 localizes to the transition zone downstream of several other proteins (Williams et al. 2011; Lambacher et al. 2015). Thus the presence of MKS6 in kif3a−/− mutants indicates that kif3a functions downstream or in parallel to the early steps in transition zone formation (summarized in Figure 8). It remains to be determined whether other transition zone proteins also localize correctly in kif3a−/− mutants.
Summary of Kif3a protein function in photoreceptor ciliogenesis. Kif3a functions at very early stages of ciliogenesis by mediating the anchoring of the basal body to the apical surface. This occurs after the formation, either partial or complete, of the transition zone, but before the assembly of IFT particles. Electron micrographs show examples of wild-type and mutant photoreceptor basal bodies at the same magnification. Arrows point to the basal body. OS, outer segment.
The function of kif3a in basal body positioning is, nonetheless, confined to a subset of cells, as basal bodies localize correctly in mechanosensory hair cells, for example. This would explain the correct apical surface anchoring of basal bodies in kif3a−/− mutant fibroblasts (Kodani et al. 2013). In addition to this, hair cells of cristae form cilia. Although substantially shorter than those in wild-type cristae, these cilia are nonetheless longer than primary cilia of many tissues: mouse brain ventricle cilia are ∼0.75 μm at embryonic day 12.5, the zebrafish spinal canal cilia ∼2.1 μm, and cilia of zebrafish somites are ∼1.8 μm at 15 hours postfertilization (Kramer-Zucker et al. 2005; Wilson et al. 2012; Lu et al. 2015a). Similarly in tissue culture, the cilia of NIH3T3 and IMCD3 measure ∼2 μm, and ∼1.5 μm, respectively (Lu et al. 2015a). To further investigate whether IFT trafficking is likely to take place in the ciliary stubs of kif3a−/− cristae, we examined the localization of Ift88. In contrast to kif3a−/− photoreceptors and hair cells of ear maculae, IFT88 localizes to cilia of kif3a−/− mutant cristae. This suggests that an unconventional IFT particle-dependent transport mechanism functions in the cilia of cristae. The cilia of cristae are unusual in that they are long and rigid (see for example Zhao et al. 2012). They may thus rely on additional kinesins for their formation. This is in agreement with our earlier findings that kif3b and kif3c function redundantly in the cilia of cristae but not maculae (Zhao et al. 2012). In summary, kif3a functions in photoreceptor basal body localization downstream or in parallel to the initial steps of transition zone formation. It also acts upstream of IFT particle assembly (Figure 8). It is, however, not required for partial ciliogenesis is specialized cilia of mechanosensory cristae.
kif17 function in photoreceptors
Studies in C. elegans have shown that the homodimeric kinesin 2, osm-3, plays a major role in ciliogenesis. In this model organism, the amphid channel cilia require kif17 homolog osm-3 for the formation of distal microtubule singlets, whereas, in the middle segment of the same cilia the heterotrimeric and homodimeric kinesins function redundantly (Snow et al. 2004; Evans et al. 2006). A recent study of phasmid cilia showed that kinesin II primarily functions in the import of IFT through the transition zone, whereas long distance transport is mainly driven by osm-3 (Prevo et al. 2015). A related functional relationship exists between homo- and heterotrimeric kinesins in amphid olfactory neuron wing C (AWC) cilia. There the heterotrimeric and homodimeric kinesin were thought to function redundantly to build the entire cilium (Snow et al. 2004; Evans et al. 2006). Although functional relationships between C. elegans ciliary kinesins are well characterized, the function of vertebrate osm-3 homolog, kif17, remained less clear. Morpholino knockdown of kif17 in zebrafish was reported to block photoreceptor outer segment morphogenesis, whereas kif17 zebrafish mutants differentiate normal outer segments (Insinna et al. 2008; Zhao et al. 2012). The kif17sa0119−/− allele contains a stop codon in the tail domain, and although this is unlikely, it has been suggested that this allele may retain some rudimentary function (Bader et al. 2012). To rule out this possibility, we obtained an alternative kif17 mutant allele that carries a stop codon in the motor domain, resulting in a truncation of the protein to approximately one-tenth of its full size. Although we detected a minute amount of the full-length Kif17 even in the kif17sa0119−/− mutant strain, this amount is much lower compared to that observed in the kif17 knockdown studies that reported outer segment loss (Insinna et al. 2008). Analysis of this allele corroborated previous findings that vertebrate kif17 does not have a major role in ciliogenesis, and does not contribute to photoreceptor outer segment differentiation. The discrepancies with previous findings may be attributed to technical shortcomings of the morpholino knockdown approach. This approach is increasingly deemed difficult to interpret based on observations that morphant phenotypes are frequently not reproduced in mutants (Kok et al. 2015).
Additional roles for IFT proteins
In photoreceptors, disruption of the IFT complex results in the mislocalization of opsin followed by rapid cell degeneration as shown by several zebrafish mutants for different IFT proteins (Doerre and Malicki 2002; Tsujikawa and Malicki 2004). As the heterotrimeric kinesin 2 is the main or perhaps the only motor that translocates the IFT particle in photoreceptors, disrupting the heterotrimeric kinesin should, in theory, result in a similar rate of photoreceptor degeneration as that seen in IFT mutants. Surprisingly, IFT mutants, oval/ift88, elipsa/ift54, and fleer/ift70, display faster photoreceptor degeneration rates, compared to kif3a−/− mutants. Double mutants of kif3a and IFT genes phenocopy IFT single mutants. The most likely explanation for this observation is that at least some IFT proteins have intraflagellar transport-independent roles in photoreceptors. An alternative but less probable scenario is that IFT protein accumulation in the cytoplasm of kif3a mutants improves photoreceptor survival. Such a function would also be cilia independent, however.
Consistent with nonciliary functions, IFT proteins have been found in nonciliary compartments of the photoreceptor cell (Sedmak and Wolfrum 2010). For example, IFT88 appears to localize to photoreceptor cell perikarya, whereas IFT20, IFT52, and IFT57 were detected in the synaptic terminals of photoreceptor cells (Sedmak and Wolfrum 2010). The roles of IFT proteins in these compartments remain, however, unclear. More recently, IFT proteins have been implicated in regulating autophagy, and, equally, autophagy regulates ciliogenesis by controlling the levels of ciliary or centriolar proteins (Pampliega et al. 2013; Tang et al. 2013). As autophagy in many retinal cells such as the lens, the cornea, and the RPE contributes to the maintenance of a healthy retina (Frost et al. 2014), it is plausible that a subset of IFT proteins functions separately from their canonical transport role to regulate autophagy in photoreceptors. A loss of this function would likely further deteriorate photoreceptor health and speed up degeneration. Such additional roles of IFT proteins are intriguing and require further studies.
Acknowledgments
The authors thank Tomer Avidor-Reiss and Freek van Eedan for comments on earlier versions of this manuscript. Chengtian Zhao and Ulla-Maria Jokipii helped with early analysis of kif3a and kif3cl function. Christopher Hill and Darren Robinson provided valuable assistance during electron microscopy and STORM imaging, respectively. Nico Katsanis and Ruxandra Bachmann provided the anti-BBS4 and anti-CC2D2A antibodies, respectively. Imaging work was performed at the University of Sheffield Wolfson Light Microscopy Facility, funded in part by Medical Research Council grant MR/K015753/1. This project was funded by National Institutes of Health/National Eye Institute R01 award EY018176 and MRC award MR/N000714.
Footnotes
Communicating editor: M. Halpern
Literature Cited
Jimeno, D., L. Feiner, C. Lillo, K. Teofilo, L. S. B. Goldstein et al., 2006a Analysis of kinesin-2 function in photoreceptor cells using synchronous Cre-loxP knockout of Kif3a with RHO-Cre. Invest. Ophthalmol. Vis. Sci. 47: 5039–5046.
Li, S., D. Chen, Y. Sauv, J. McCandless, Y.-J. Chen et al., 2005 Rhodopsin-iCre transgenic mouse line for Cre-mediated rod-specific gene targeting. Genesis 41: 73–80.
Pugh, E., and T. Lamb, 2000 Phototransduction in vertebrate rods and cones: pp. 183–255 in Handbook of Biological Physics. Elsevier Science, Amsterdam.
Scholey, J. M., 2013 Kinesin-2: a family of heterotrimeric and homodimeric motors with diverse intracellular transport functions. Annu. Rev. Cell Dev. Biol. 29: 443–469.