-
PDF
- Split View
-
Views
-
Cite
Cite
Rebecca J Mroczek, R Kelly Dawe, Distribution of Retroelements in Centromeres and Neocentromeres of Maize, Genetics, Volume 165, Issue 2, 1 October 2003, Pages 809–819, https://doi.org/10.1093/genetics/165.2.809
Close - Share Icon Share
Abstract
Fluorescent in situ hybridization was used to examine the distribution of six abundant long terminal repeat (LTR) retroelements, Opie, Huck, Cinful-1, Prem-2/Ji, Grande, and Tekay/Prem-1 on maize pachytene chromosomes. Retroelement staining in euchromatin was remarkably uniform, even when we included the structurally polymorphic abnormal chromosome 10 (Ab10) in our analysis. This uniformity made it possible to use euchromatin as a control for quantitative staining intensity measurements in other regions of the genome. The data show that knobs, known to function as facultative neocentromeres when Ab10 is present, tend to exclude retroelements. A notable exception is Cinful-1, which accumulates in TR-1 knob arrays. Staining for each of the six retroelements was also substantially reduced in centromeric satellite arrays to an average of 30% of the staining in euchromatin. This contrasted with two previously described centromere-specific retrotransposable (CR) elements that were readily detected in centromeres. We suggest that retroelements are relatively rare in centromeres because they interrupt the long satellite arrays thought to be required for efficient centromere function. CR elements may have evolved mutualistic relationships with their plant hosts: they are known to interact with the kinetochore protein CENH3 and appear to accumulate in clusters, leaving long satellite arrays intact.
TRANSPOSABLE elements are divided into two major groups: class I elements, which transpose via an RNA intermediate, and class II elements, which transpose through DNA replication. Class I transposable elements include LINEs, SINEs, and long terminal repeat (LTR) retroelements. The latter occupy a significant portion of large plant genomes like that of maize, where intergenic regions are composed primarily of nested LTR retrotransposons (SanMiguel et al. 1996; SanMiguel and Bennetzen 1998). Roughly 50% of the maize, rye, barley, and wheat genomes are thought to be composed of LTR retroelements (SanMiguel et al. 1996; Pearce et al. 1997; SanMiguel and Bennetzen 1998; Vincent et al. 1999; Meyers et al. 2001; Wicker et al. 2001). The same variety of transposable elements appears to exist in smaller-genome species such as rice and Arabidopsis, but fewer representatives of each class are present. Only ∼14% of the Arabidopsis genome is composed of transposable elements (Arabidopsis Genome Initiative 2000). The fact that genome size varies greatly while gene number varies little (the C-value paradox; Cavalier-Smith 1978; Pagel and Johnstone 1992) can be largely attributed to extraordinary variation in the number of retroelements (Kumar and Bennetzen 1999; Tikhonov et al. 1999; Wicker et al. 2001; Bennetzen 2002).
LTR retroelements are separated into two major groups on the basis of the organization of the domains within their Pol genes: in the Ty1/copia-like group the integrase (INT) domain is located upstream of the reverse transcriptase (RT) domain, while in the Ty3/gypsy-like group the INT domain is located downstream of the RT domain (Xiong and Eickbush 1990). The two groups can be further subdivided into families on the basis of the similarity of their LTR sequences, which evolve faster than the internal coding regions (SanMiguel and Bennetzen 1998). The distribution of gypsy- and copia-like LTR retroelements has been analyzed in a number of large genome plants via fluorescent in situ hybridization (FISH; Pearce et al. 1996a,b; Brandes et al. 1997; Kumar et al. 1997; Friesen et al. 2001). The patterns of localization suggest that retroelements have insertional preferences. Ty1/copia retroelements are found throughout the euchromatin of Vicia faba (Pearce et al. 1996a) but are generally concentrated in the subtelomeric heterochromatin of Allium cepa (Kumar et al. 1997). There is also evidence from Allium and gymnosperms that individual retroelement families have discernibly different patterns of chromosomal localization (Pich and Schubert 1998; Friesen et al. 2001).
Although most retroelements are distributed nonrandomly throughout chromosomes, the most obvious discontinuities occur with respect to tandem repeat arrays. Brandes and co-workers (1997) examined a variety of organisms (A. cepa, Beta vulgaris, Brassica campestris, Brassica oleracea, Pennisetum glaucum, Pinus elliottii, Selaginella apoda, V. faba, and V. narbonensis) and demonstrated that the Ty1/copia group is dispersed throughout the euchromatic regions, but absent from regions where specialized tandem repeats are expected to lie, such as centromeres, telomeres, heterochromatin, and the nucleolus organizing region (NOR). FISH analyses in maize using portions of the Opie and Prem-2/Ji retroelements showed diffuse patterns of staining with reduced accumulation at the centromeres and NOR (Edwards et al. 1996; Ananiev et al. 1998a; Miller et al. 1998). In marked contrast are the gypsy-like centromeric retrotransposable (CR) elements of cereal grains, which accumulate specifically in centromeric satellites (Jiang et al. 1996; Presting et al. 1998; Hudakova et al. 2001; Zhong et al. 2002; Nagaki et al. 2003a). Recent data indicate that the maize CR elements interact with the kinetochore protein centromeric histone H3 (CENH3; Zhong et al. 2002), suggesting that they participate in centromere function.
Here we describe and quantify the accumulation patterns of a variety of maize retroelement families from both the Ty3/Gypsy and Ty1/Copia groups, including two different types of maize CR elements. We find that both the Gypsy and Copia groups are found throughout the euchromatic portions of the genome. In contrast, all but one of the retroelements analyzed are under-represented in knobs, which are known to function as facultative centromeres. Additionally, all of the retroelements outside of the CR clade are largely excluded from centromeric satellite arrays. The data suggest that centromeric satellite arrays are under selection for their function in chromosome movement, much like genic regions (SanMiguel et al. 1996).
MATERIALS AND METHODS
Maize stocks: The standard maize inbred lines W23 and KYS were used for the bulk of the cytological analysis. The strain containing abnormal chromosome 10 (Ab10) was originally obtained from Marcus Rhoades and subsequently backcrossed into the W23 background seven times.
Phylogenetic analysis: Nucleotide sequences were retrieved from GenBank for the following maize retroelements: Grande 1-4 (GenBank X97604), Huck (AF391808-1), Tekay (AF050455), Fourf (AF050436), Mare 5 (AB033252.1), Reina (U69258), Rle (AF057037), Cinful-1 (AF049110), Cinful-2 (AF049111), Prem-2 (U41000), Opie-2 (U68408), Cent-A (AF078917), CRM (AY129008), CRR (AC022352), Cereba (AY040832), and two Arabidopsis thaliana retroelement sequences (AAD11616 and BAB40826). The reverse transcriptase regions were identified following the guidelines set forth by Xiong and Eickbush (1990). A progressive alignment of the RT regions was prepared using the Pileup option in GCG's Seqlab with a BLOSUM 30 transition matrix (GCG 1982–2000; Feng and Doolittle 1987; Henikoff and Henikoff 1992). The alignments were adjusted manually using previously published RT alignments as visual templates (Xiong and Eickbush 1988; Bowen and McDonald 2001). Both parsimony heuristics and neighbor joining (NJ) were used to generate trees from our alignment using PAUP* (Swofford 1999). The NJ trees were derived using uncorrected pairwise distances (Swofford 1999), and bootstrap values were determined using PAUP*.
Probe preparation: Primers specific to LTRs were designed from the sequences noted above with the exceptions of Huck (AF050438) and Grande (AF050437). The lengths of the amplified products and the primers used for the amplification are: Cinful-1, 562 bp (F-5′-CGCCGAAGGTCTTCTAGGAA-3′, R-5′-GGAGACTCGTTCTCAAGTGCTA-3′); Grande, 350 bp (F-5′-ATGCGAGGATAAGTCGGCGAAG-3′, R-5′-GGTGTTTTTAG GAGTAGGACGGTG-3′); Huck, 673 bp (F-5′-TCCACTGACC GACCTGACAA-3′, R-5′-GGTTTTGGCACCCTGTTCAT-3′); Opie-2, 526 bp (F-5′-CAAACACAAGTGCTTAAAT-3′, R-5′-GTC CGGTGCCCGATTTGT-3′); Prem-2/Ji, 573 bp (F-5′-ACATTT GGTGGTTGGGGCTA-3′, R-5′-GGGTGAATAGGGCGAAAC TGAA-3′); Tekay, 537 bp (F-5′-ATTTGTGCGACCGCTCAA-3′, R-5′-AGGAGTCCAGGCTGCTCTTA-3′); Cent-A, 1234 bp (F-5′-CATAACCCGCACAGATATGAC-3′, R-5′-ATAAACCCAACG GGTAGAAGGG-3′; and CRM, 513 bp (F-5′-TCGTCAACT CAACCATCAGGTGAT-3′, R-5′-GCAAGTAGCGAGAGCTAA ACTTGA-3′). The PCR fragments were each cloned into the TOPO cloning vector (Invitrogen, Carlsbad, CA) and verified by sequencing.
In situ hybridization: Anthers from maize inbred lines were fixed as previously described (Hiatt et al. 2002). Probes were amplified from plasmids and labeled with FITC using a random primer labeling kit (Prime-It Fluor fluorescence labeling kit, Stratagene, La Jolla, CA). The CentC and TR-1 repeats were detected using rhodamine-labeled oligonucleotides specific to each sequence (Hiatt et al. 2002; Zhong et al. 2002). Chromosome straightening and in situ hybridization were carried out as previously described (Dawe et al. 1994; Zhong et al. 2002). All data were collected and analyzed using a DeltaVision 3D light microscope workstation and associated software. Intensity data were collected by first selecting 9 × 9, two-dimensional “pixel boxes.” From these boxes we recorded the total intensity values (amount of light detected by the CCD camera in that region) from the DNA 4′,6-diamidino-2-phenylindole (DAPI) and retroelement (FITC) channels. Background, euchromatin, centromere, and knob readings were taken in each channel for each cell. Background measurements were averaged from 5 different pixel boxes and euchromatin measurements were averaged from 10 different pixel boxes. For knobs and centromeres the number of pixel boxes for which data were collected was limited by the number of centromeres or knobs that were unobstructed by chromosome arms; between 4 and 7 pixel boxes were taken for each structure in each cell. The pixel box data from each channel were averaged, and appropriate background levels were subtracted for each cell. The overall intensity values varied from cell to cell, as is typical for FISH experiments. We therefore calculated within-cell ratios of centromere/euchromatin or knob/euchromatin staining intensities for both channels and compared them. By taking advantage of an internal control (euchromatin staining), we effectively normalized the data, making it possible to average the information from different cells.
RESULTS
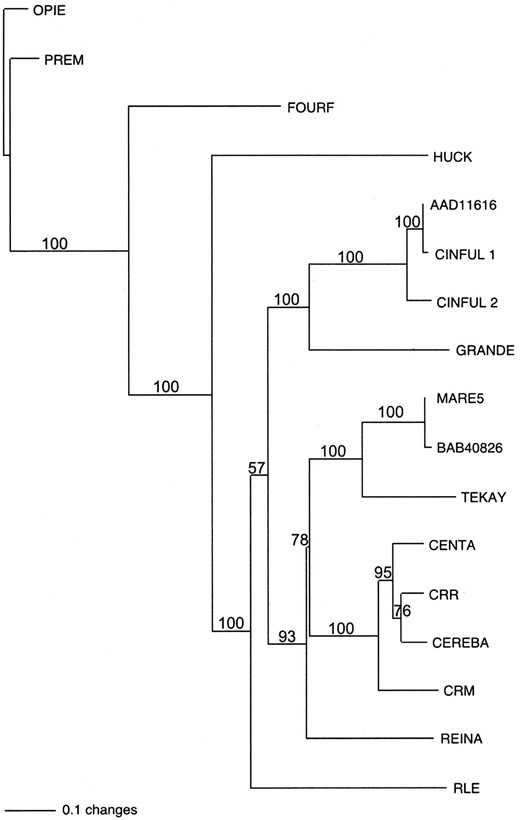
Phylogeny of the retroelement families analyzed: We identified the RT regions of the retroelements examined in this report and compared their sequences to several other retroelements phylogenetically. Neighbor-joining and parsimony analysis of the retroelement RT regions produced identical trees. Figure 1 is a neighbor-joining tree, which shows that the retroelements can be divided into two major clades representing the gypsy- and copia-like groups of the LTR retroelements, a finding consistent with other phylogenetic analyses (Xiong and Eickbush 1990; Malik and Eickbush 1999; Bowen and McDonald 2001). The known CR elements group together to form a monophyletic clade within the gypsy-like group (Langdon et al. 2000; Nagaki et al. 2003a). In maize there are two different CR elements, CRM and Cent-A, which have closely related internal regions (67% nucleotide similarity) but different LTRs (Ananiev et al. 1998a; Zhong et al. 2002). Among the known maize retroelements, Tekay/Prem-1 and Reina are the most closely related to the CR elements. However, only Tekay/ Prem-1 is abundant enough (SanMiguel and Bennetzen 1998; Meyers et al. 2001) to be readily detected by FISH.
—Neighbor-joining tree of the RT amino acid sequences from a variety of plant retroelements. AAD11616 and BAB40826 represent RT sequences from Arabidopsis. CRM and Cent-A are CR elements from maize. CRR and Cereba are CR elements from rice and barley, respectively. All other retroelements are from maize. Numbers at the nodes represent the bootstrap values obtained using PAUP*.
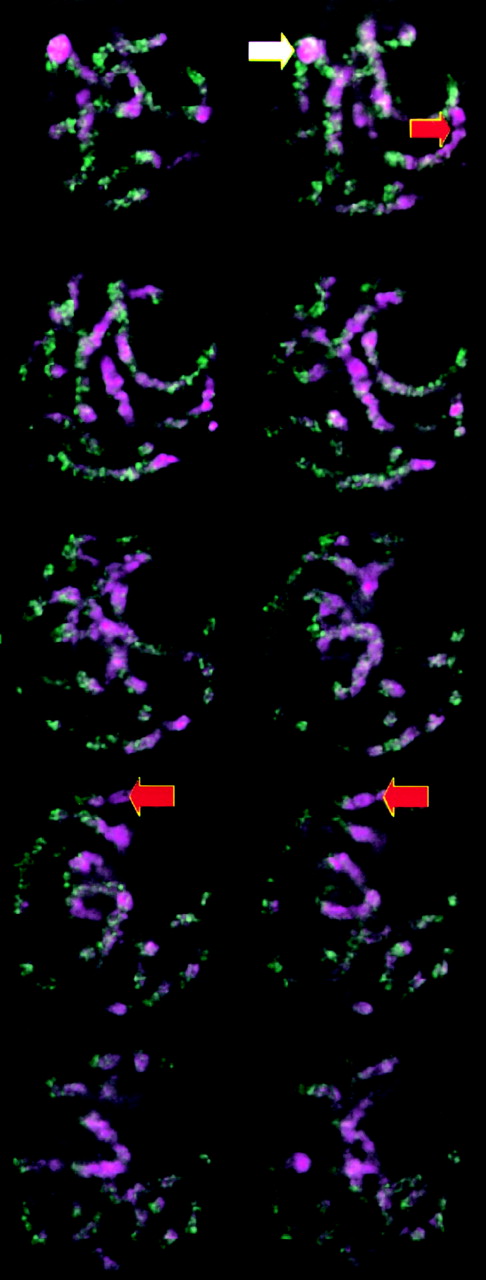
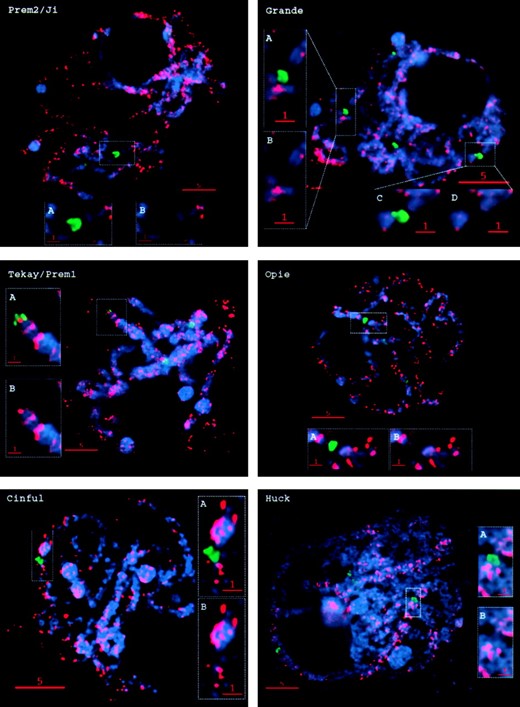
The abundant maize retroelement families show genome-wide distributions: We chose six of the most abundant maize retrotransposon families for in situ hybridization analysis: Huck, Opie, Grande, Prem-2/Ji, Cinful-1, and Tekay/Prem-1 (SanMiguel et al. 1996; SanMiguel and Bennetzen 1998; Meyers et al. 2001). Probes were generated to portions of the LTRs that are unique to individual families. These LTR fragments were then fluorescently labeled and hybridized to maize pachytene chromosome preparations, and the results were analyzed using deconvolution 3D light microscopy. Each of the retroelement families outside of the CR clade showed roughly uniform euchromatic staining patterns throughout the genome (Figures 2 and 3), with subtle differences among families. Opie is evenly distributed in a pattern of neat dots along the chromosomes (Figure 3), Huck has a patchy distribution (Figure 3), and Prem-2/Ji has a genome-wide distribution but on some chromosomes is underrepresented in pericentromeric heterochromatin (Figure 2, arrows; Figure 3).
Retroelement abundance in euchromatic regions is relatively uniform but varies in knobs: Visual inspection of the images in Figures 2 and 3 indicated a remarkable uniformity in the distribution of retroelements along chromosome arms, suggesting that euchromatin staining might serve as a suitable internal control for the staining in centromeric regions. To test this idea, we compared the euchromatin on Ab10 to other chromosome arms. Ab10 is an alternative version of the normal chromosome 10 (N10) that is present in ∼10% of teosinte (the ancestor of maize) and ∼2% of known maize strains (Kato 1976). The terminal portion of the long arm of Ab10 is responsible for the phenomena of neocentromere activity and meiotic drive. This region contains few essential genes (Hiatt and Dawe 2003) and an ∼14 map unit inversion (Rhoades and Dempsey 1985). In addition, there are three small knobs containing arrays of a 350-bp TR-1 knob repeat and a large knob composed primarily of a 180-bp repeat (Peacock et al. 1981; Rhoades and Dempsey 1985; Ananiev et al. 1998b; Hiatt et al. 2002). The striking structural polymorphism between Ab10 and N10, most notably the inverted region, is thought to be responsible for the fact that recombination between the distal portion of Ab10 and N10 rarely occurs (Kikudome 1959; Rhoades and Dempsey 1985). The unusual structural features and evolutionary history of Ab10 suggested that it might have an unusual distribution or abundance of retroelements. We assayed the localization patterns of a sample of retroelements (Huck, Prem-2/Ji, and Cinful-1) on the distal portion of the Ab10 chromosome. As can be seen in Figure 4, retroelement staining in the euchromatic portion of the distal region of Ab10 was nearly identical to the intensity and pattern of staining observed for the rest of the genome. To quantify this observation we took advantage of the fact that chromatin (DAPI) and retroelement (FITC) staining are measured and stored as separate images during data collection and that deconvolution microscopy is quantitative. Staining intensity readings were taken for both chromatin and retroelements and compared to the staining intensities found in other euchromatic regions. We found no significant difference between Ab10 and the rest of the genome (t-test, P < 0.01). Although the DAPI intensity in Ab10 euchromatin was relatively low compared to the genome as a whole (mean = 0.73, SD ±0.53), retro-element intensities were similarly reduced (mean = 0.77, SD ±0.21). These data support the view that retroelements are spread evenly and uniformly throughout maize euchromatin.
—Genome-wide distribution of the Prem-2/Ji retroelement family in maize. Images are single optical sections taken from a maize pachytene meiocyte, separated by 4 μm. DNA is represented in magenta and the Prem-2/Ji retroelement family is represented in green. Absence of Prem-2/Ji staining at centromeres is indicated with red arrowheads. Prem-2/Ji staining at a knob is shown with a white arrowhead.
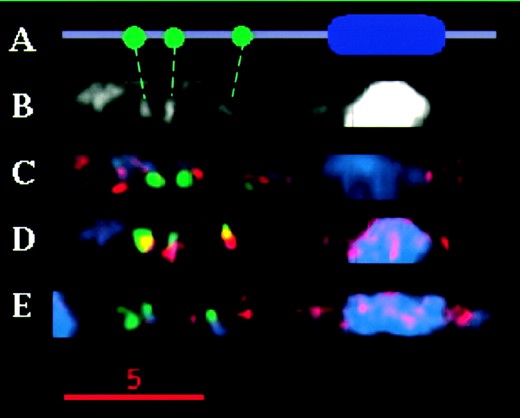
Our analysis of Ab10 also revealed that euchromatin and knobs stain differently for retroelements. For instance, Huck is nearly absent from the large knob of Ab10 (Figure 4C), and Cinful-1 is highly abundant within the TR-1-containing chromomeres (indicated by staining that appears yellow in Figure 4D). As described below, these observations were pursued in more detail by analyzing a variety of other knobs in the genome.
Retroelement families are variably interspersed in maize knob satellite DNA: The TR-1 and 180-bp repeats present on Ab10 also occur at 22 other knob loci in differing proportions (Kato 1976; Ananiev et al. 1998c; Buckler et al. 1999; Hiatt et al. 2002). Many other classes of repeats may also be present in maize knobs. To obtain a more general perspective on knobs without a bias toward particular repeats, we scored retroelement staining in knobs as identified by their characteristic ball-shaped heterochromatic structure (Figure 5; see also Figures 2 and 3). We found that all families except Huck showed some staining in nearly every knob observed (Figure 5). The retroelement families Cinful-1, Grande, Tekay/Prem-1, Opie, and Prem-2/Ji all showed staining in the majority of knobs examined (84.8, 86.7, 87.1, 94.9, and 96.3%, respectively). In contrast, the Huck retroelement family was nearly absent in most knobs with some staining in only 20% of the knobs examined (Table 1). Retroelements from the Huck family apparently avoid or are selectively removed from knobs even though they occupy ∼10% of the maize genome (SanMiguel and Bennetzen 1998; Meyers et al. 2001). Our observations are consistent with the findings of Ananiev and co-workers (1998b), who found no copies of the Huck retroelement family in any of 23 cloned knob segments.
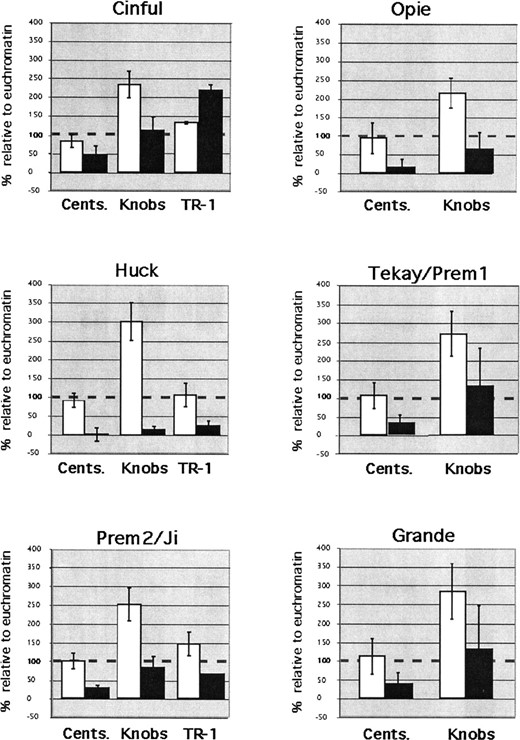
Probes for retroelements outside of the Huck and CR families appeared to stain euchromatin and knobs at about the same intensity (Figure 5). Since knobs stain with DAPI very brightly, this observation implied a relatively low abundance of retroelements within the knobs. Intensity measurements confirmed the interpretation: we found that knobs were 2.6 times brighter in the DAPI channel than an average segment of euchromatin, but that retroelement staining within knobs was only 1.1 times brighter (Figure 6). In the small percentage of knobs that showed Huck staining, retroelement staining intensities were only 15% of the levels in euchromatin. These data indicate that although most retroelement families are present in the satellite repeats of knobs, they are present at a reduced frequency when compared to euchromatin.
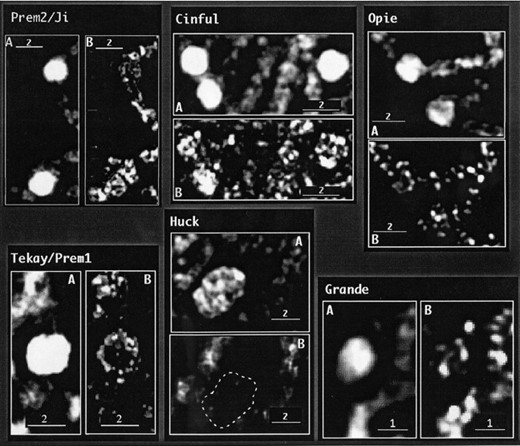
—Staining patterns of maize retroelement families throughout the genome with respect to the centromeres. Each image shows a single optical section of a cell at pachytene. DNA is shown in blue, CentC in green, and retroelement LTR staining in red. (A and C) Enlargements (2×) of the boxed areas showing retroelement staining around the centromere. (B and D) Same images as A and C, except with the CentC staining removed.
Data from the Ab10 chromosome indicated that Cinful-1 retroelements accumulate in TR-1 knobs (Figure 4D). To determine if Cinful-1 accumulation was limited to Ab10 knobs, we took intensity readings from the TR-1 knobs of Ab10 as well as three other TR-1-containing knobs. Overall, Cinful-1 staining was 2.2 times higher in TR-1 knobs than in euchromatin (Figure 6), and there was no significant difference between the TR-1 knobs on Ab10 and those elsewhere in the genome (t-test, P < 0.01). In contrast, the staining intensities for Huck and Prem-2/Ji in TR-1 knobs were similar to those in non-TR-1-containing knobs. The fact that Cinful-1 is substantially overrepresented in TR-1 arrays provides evidence that the reduced staining we detect for all the other retroelements is not a consequence of the heterochromatic nature of knobs.
Abundant retroelement families are largely absent from centromeres: Maize centromeres contain tandem arrays of the 156-bp Cent-C satellite repeat, interspersed with the CR elements Cent-A and CRM (Zhong et al. 2002; Nagaki et al. 2003a). Noncentromere-specific retroelements are also present to some extent, although the overall frequency of these elements in centromeres is not known (Ananiev et al. 1998a; Nagaki et al. 2003a) and will be difficult to determine by sequence analysis, given the inherent limitations associated with cloning and contiging long repeat arrays (Song et al. 2001; Henikoff 2002). In an effort to further examine and quantify the accumulation patterns of retroelements in centromeres, we labeled the major retroelement LTRs and Cent-C with different fluorescent dyes and analyzed the results on DAPI-stained maize pachytene chromosomes.
—Staining patterns of retroelements in the polymorphic portion of Ab10. (A) Graphical representation of the Ab10 chromosome showing the small TR-1 knobs in green, the large 180-bp repeat knob in blue, and the euchromatic regions in purple. (B) Black and white image of a computationally straightened Ab10 chromosome showing only DNA (DAPI) staining; chromomeres are indicated with connecting lines to the graphical representation in A. (C) Huck staining on a computationally straightened Ab10 pachytene chromosome. TR-1 staining is in green and retroelement staining is in red. (D) Straightened chromosome from a cell stained with Cinful-1 in red and TR-1 in green. Intense Cinful staining at the TR-1 knobs makes them appear yellow, due to the overlap of the red and green colors. (E) Straightened chromosome showing Prem-2/Ji in red and TR-1 in green.
We found that each of the retroelement families, Huck, Opie, Grande, Prem-2/Ji, Cinful-1, and Tekay/Prem-1, are poorly represented at centromeres (Figures 3 and 6 and Table 1). The percentage of centromeres with detectable staining for these retroelements ranged from 3.4 to 37%, depending on the family (Table 1). Similarly, while DAPI-staining intensities were roughly equivalent in centromeres and chromosome arms, non-CR retroelement staining in centromeres averaged only 30% of the levels found in euchromatin (Figure 6).
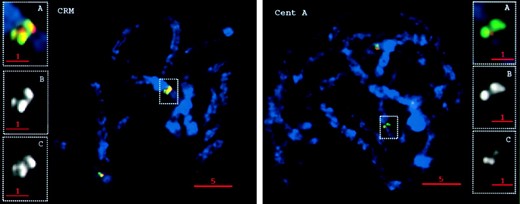
In marked contrast to other maize retroelement families, in situ hybridization with the CR elements Cent-A (Ananiev et al. 1998a) and CRM (Zhong et al. 2002; Nagaki et al. 2003a) revealed strict centromeric localization on all 10 maize chromosomes (Figure 7, Table 1). We often observed overlap of the CentC and CR signals (Figure 7, CRM), and in many cases the two signals were clearly separate (Figure 7, Cent-A). These data are consistent with fiber-FISH data from rice, which suggest that CR elements tend to insert into large clusters distinct from the regions composed mainly of satellite repeats (Cheng et al. 2002).
DISCUSSION
In this study we provide a perspective on the distribution of the most abundant retroelements in maize with particular emphasis on the centromeres and neocentromeres. Previous reports have used general reverse transcriptase probes to examine overall distribution patterns of retroelements (Pearce et al. 1996a,b, 1997; Brandes et al. 1997; Kumar et al. 1997) or have localized specific families without a special emphasis on centromeres (Pich and Schubert 1998; Friesen et al. 2001). Retroelement distribution is also being analyzed in the sequenced plant genomes of Arabidopsis and rice (Arabidopsis Genome Initiative 2000; Feng et al. 2002). However, sequence data are generally unreliable in regions containing long satellite arrays (Henikoff 2002). Even the size of most eukaryotic centromeres is still under debate (Haupt et al. 2001; Hosouchi et al. 2002). The FISH strategy employed here has a lower resolution than DNA sequencing, but can provide a general and quantitative perspective on the frequency of retroelements in large centromeres. By focusing on a single organism and using retroelement family-specific probes and quantitative light microscopy, we have been able to draw new conclusions and hypotheses about the forces driving retrotransposon accumulation in large-genome species.
Chromosomal localization with respect to evolutionary history of the elements: The families examined represent both the gypsy- and copia-like groups of LTR retroelements. Huck, Cinful-1, Tekay/Prem-1, Grande, Cent-A, and CRM belong to the gypsy-like group and Prem-2/Ji and Opie fall into the copia-like group. In our analysis of eight retroelement families from maize we saw no obvious correlation between the type of retroelement (gypsy- or copia-like) and chromosomal localization patterns. The rapid evolution of localization patterns is particularly evident with regard to the CR elements and the closely related Tekay/Prem-1 retroelement family. Although Tekay/Prem-1 shares a more recent common ancestor with the CR elements than do the other families examined here, it is no more likely to be found in or near the centromere than are more distantly related families. As can be seen in Table 1 and Figure 6, the Cinful-1 and Opie families are just as likely to show centromeric staining as the Tekay/Prem-1 family is. The available data suggest that the strict centromeric localization pattern and apparent function of CR elements in recruiting centromeric histone are recently evolved features primarily limited to the cereal grains.
Accumulation of retroelements in Ab10 and the satellite repeats of maize knobs: The abnormal 10 chromosome of maize provides all knobs in the genome with the capacity to move as neocentromeres and preferentially segregate to progeny (Rhoades 1942). Within the terminal region of the long arm of Ab10 is an inversion of chromatin from normal 10, three small knobs composed primarily of TR-1 repeats, a large knob composed primarily of 180-bp repeats, and regions of apparently novel chromatin (Hiatt et al. 2002). Genetic data suggest that there are few essential genes in the latter half of this large structural polymorphism (Hiatt and Dawe 2003). Meiotic drive systems may spread to some extent without regard to organismal fitness and are known to accumulate deleterious mutations (Ardlie 1998). Similarly, retroelements have been shown to accumulate in Drosophila inversions, presumably because recombination events that would eliminate them are reduced there
Presence of retroelements at centromeres and knobs
| Retroelement . | Total centromeres counted . | % centromeres with detectable staining . | Total knobs counted . | % knobs with detectable staininga . |
|---|---|---|---|---|
| Opie | 19 | 26.3 | 39 | 94.9 |
| Prem-2/Ji | 58 | 3.4 | 80 | 96.3 |
| Cinful-1 | 27 | 37.0 | 46 | 84.8 |
| Grande | 13 | 15.4 | 15 | 86.7 |
| Tekay/Prem-1 | 31 | 22.6 | 31 | 87.1 |
| Huck | 30 | 6.7 | 40 | 20.0 |
| Cent-A | 32 | 100.0 | b | b |
| CRM | 60 | 100.0 | b | b |
| Retroelement . | Total centromeres counted . | % centromeres with detectable staining . | Total knobs counted . | % knobs with detectable staininga . |
|---|---|---|---|---|
| Opie | 19 | 26.3 | 39 | 94.9 |
| Prem-2/Ji | 58 | 3.4 | 80 | 96.3 |
| Cinful-1 | 27 | 37.0 | 46 | 84.8 |
| Grande | 13 | 15.4 | 15 | 86.7 |
| Tekay/Prem-1 | 31 | 22.6 | 31 | 87.1 |
| Huck | 30 | 6.7 | 40 | 20.0 |
| Cent-A | 32 | 100.0 | b | b |
| CRM | 60 | 100.0 | b | b |
Number of knobs observed for which retroelement staining was at least as intense as that observed throughout the euchromatin of the chromosomes.
Cent-A and CRM are not detected at knobs or euchromatin.
Presence of retroelements at centromeres and knobs
| Retroelement . | Total centromeres counted . | % centromeres with detectable staining . | Total knobs counted . | % knobs with detectable staininga . |
|---|---|---|---|---|
| Opie | 19 | 26.3 | 39 | 94.9 |
| Prem-2/Ji | 58 | 3.4 | 80 | 96.3 |
| Cinful-1 | 27 | 37.0 | 46 | 84.8 |
| Grande | 13 | 15.4 | 15 | 86.7 |
| Tekay/Prem-1 | 31 | 22.6 | 31 | 87.1 |
| Huck | 30 | 6.7 | 40 | 20.0 |
| Cent-A | 32 | 100.0 | b | b |
| CRM | 60 | 100.0 | b | b |
| Retroelement . | Total centromeres counted . | % centromeres with detectable staining . | Total knobs counted . | % knobs with detectable staininga . |
|---|---|---|---|---|
| Opie | 19 | 26.3 | 39 | 94.9 |
| Prem-2/Ji | 58 | 3.4 | 80 | 96.3 |
| Cinful-1 | 27 | 37.0 | 46 | 84.8 |
| Grande | 13 | 15.4 | 15 | 86.7 |
| Tekay/Prem-1 | 31 | 22.6 | 31 | 87.1 |
| Huck | 30 | 6.7 | 40 | 20.0 |
| Cent-A | 32 | 100.0 | b | b |
| CRM | 60 | 100.0 | b | b |
Number of knobs observed for which retroelement staining was at least as intense as that observed throughout the euchromatin of the chromosomes.
Cent-A and CRM are not detected at knobs or euchromatin.
(Sniegowski and Charlesworth 1994). Because of its association with meiotic drive, reduced recombination, and relative lack of genes, we considered whether the distal portion of Ab10 would be a favored spot for the accumulation of retroelements. However, outside of the Cinful-1 family, which accumulates in all TR-1 arrays, the retroelement families we examined appear to be no more abundant on this chromosome than they are throughout the rest of the genome (Figure 4). These data suggest that the reduced recombination and meiotic drive typical of the long arm of Ab10 have had little impact on retroelement distribution over the time span in which the drive system has existed (see Buckler et al. 1999).
—Patterns of retroelement staining at knobs. Enlargements of knobs are from the cells shown in Figure 3, although some are from different optical sections. (A) Image of DNA showing intensely staining heterochromatic knobs. (B) Same image showing only the FITC (retroelement) staining. Note that Huck staining at the knob is markedly reduced.
The remarkable uniformity of retroelement distribution in maize euchromatin gave us a useful internal control for the staining of retroelements in knobs and centromeres. The analysis indicates that retrotransposons are substantially underrepresented in knobs relative to euchromatin (Figures 4 and 6). One explanation for the underrepresentation of retroelements in knobs is that the structural organization of knob heterochromatin is such that FISH probes cannot gain access. However, the fact that we detected an abundance of Cinful-1 elements in TR-1 arrays strongly suggests that knob structure did not serve as a substantial barrier to FISH probes. In addition, we have corroborated a previous conclusion, on the basis of the analysis of cosmid clones, that Huck is underrepresented in knobs (Ananiev et al. 1998b). We believe that our data provide an accurate view of the prevalence of retroelements in knob repeat arrays.
—Staining intensities of (□) DNA and (▪) retroelements in centromeres, cytologically defined knobs, and TR-1 arrays. Staining intensities are expressed relative to euchromatin. Values below the dashed line indicate relatively low staining intensities; values above the line indicate high staining intensities. Vertical lines indicate the standard deviation.
The fact that retroelements are underrepresented in knobs indicates that long tracts of tandem repeats may be required for neocentromere function, as argued below for centromeres. Although there are clearly more retroelements in knobs than in centromeres (Figure 6), this may reflect the fact that neocentromeres are activated only when Ab10 is present, and as a result there are less stringent evolutionary restraints upon the content of knobs. Interestingly, TR-1 satellite arrays are considerably more neocentric (active on the spindle) than 180-bp arrays and are controlled by different transacting factors (Hiatt et al. 2002). Our data suggest that Cinful-1 does not actively interfere with TR-1-mediated neocentromere activity and leave open the possibility that Cinful-1 may contribute to knob motility, as has been suggested for the CR elements (Zhong et al. 2002).
—Staining patterns of the maize CR elements CRM and CentA. The DNA is shown in blue, the retroelement is shown in red, and the centromeric repeat CentC is shown in green. Large images are single optical sections from maize pachytene cells. (A) Enlargement (3×) of the boxed regions. (B) Same images as A except only CentC staining is shown. (C) Same images as A showing only the retroelement staining.
Abundance of maize retroelements in centromeres: An unusual feature of cereal centromeres is that they specifically accumulate a group of Ty3-gypsy-like retroelements known as CR elements. CR elements show remarkable sequence conservation and are found in the centromeres of rice, wheat, sorghum, barley, rye, and oats (Aragon-Alcaide et al. 1996; Jiang et al. 1996; Miller et al. 1998; Presting et al. 1998; Langdon et al. 2000). Zhong and co-workers have recently reported that maize CR elements interact with CENH3, indicating that they may have been coopted to perform a function in chromosome segregation (Zhong et al. 2002). The strict centromeric localization of CR elements is even more striking in light of data shown here that several other maize retroelement families are poorly represented within centromeres, at least as defined by colocalization with the CentC satellite repeat. CentC strongly interacts with the kinetochore protein CENH3, suggesting that it is a legitimate marker for the functional centromere (Zhong et al. 2002). Cent-A and CRM are abundant in centromeres (Figure 7), whereas retroelements that occur throughout the chromosome arms (Grande, Opie, Prem-2/Ji, Huck, Cinful-1, and Tekay/Prem-1) are significantly underrepresented in centromeres (Figures 2, 3, and 6, and Table 1). There is some family-to-family variation, but overall the non-CR retroelements are found threefold less frequently in centromeres than in euchromatin. The tandem repeats in maize centromeres apparently represent a niche of the genome that is favorable to CR elements and seemingly hostile to families of retroelements that proliferate throughout the intergenic, euchromatic niches of the genome.
Extended tandem repeat arrays are thought to be required for centromere function in animals (Grimes et al. 2002) and are characteristic of the centromeres in Arabidopsis and rice (Kumekawa et al. 2001; Cheng et al. 2002). Rampant insertion of retroelements in the centromere would interrupt the continuity of these long arrays. On the basis of our data, we hypothesize that such insertions adversely impact the formation of centromeric chromatin. Other functional domains of the chromosome, most notably those containing genes, also rarely contain the large insertions that retrotransposons generate (SanMiguel et al. 1996). Non-CR retrotransposons may be excluded from the centromere by the chromatin structure at the centromere or, alternatively, could be eliminated during the process of unequal crossing over that is thought to homogenize tandem repeat arrays (Eickbush 2002). The CR elements may circumvent elimination via recombination by continuously transposing back into the centromeric satellite arrays, as has been proposed for the maintenance of R1 and R2 elements in Drosophila rRNA gene arrays (Eickbush 2002; Perez-Gonzalez and Eickbush 2002). This interpretation is supported by sequencing data showing that most CR elements are recent insertions (Nagaki et al. 2003a). Further, fiber-FISH analyses (Cheng et al. 2002) and our own localization data (Figure 7) suggest that CRs transpose into specific domains of the centromere, leaving the tandem repeat arrays largely intact.
At least a subset of known retroelements seems to have evolved mutualistic relationships with their hosts. Two particularly well-studied examples in this category are the HeT-A and TART elements of Drosophila, which substitute for telomerase by continually transposing into chromosome ends (Pardue and Debaryshe 2000). Although the host normally responds to retroelements by actively repressing them, Drosophila may tolerate HeT-A and TART because it benefits from their presence. Similarly, CR elements accumulate in specific domains of the centromere where they interact with the key kinetochore protein CENH3 (Zhong et al. 2002). These data suggest that CR elements contribute to the specification and/or maintenance of the centromere/kinetochore complex. In the same manner that HeT-A and TART elements are limited to Drosophila species (Casacuberta and Pardue 2003), the CR elements and their association with centromeres may be limited to cereal grains. No Arabidopsis retroelements transpose specifically into the centromere core (Haupt et al. 2001; Kumekawa et al. 2001) and chromatin immunoprecipitation experiments have failed to reveal any association between retroelements and the Arabidopsis homolog of CENH3 (Nagaki et al. 2003b). The functional centromeric sequences in Arabidopsis, as in humans (Schueler et al. 2001; Grimes et al. 2002), appear to be long uninterrupted arrays of satellite repeats (Nagaki et al. 2003b).
Footnotes
Communicating editor: S. Henikoff
Acknowledgement
We thank Carolyn Lawrence for advice and help throughout the study, and Susan Wessler for suggesting that we analyze retroelement distribution on the Ab10 chromosome, as well as Evelyn Hiatt, Cedric Feschotte, and an anonymous reviewer for detailed comments on the manuscript. This work was supported by a grant from the National Science Foundation (9975827) to R.K.D. Additional support was provided to R.J.M. by a National Science Foundation interdisciplinary research training grant (BIR9220329).
LITERATURE CITED